Back to Journals » OncoTargets and Therapy » Volume 7
Clinical significance of promoter region hypermethylation of microRNA-148a in gastrointestinal cancers
Authors Sun J, Song Y , Wang Z, Wang G, Gao P, Chen X, Gao Z, Xu H
Received 17 January 2014
Accepted for publication 8 April 2014
Published 28 May 2014 Volume 2014:7 Pages 853—863
DOI https://doi.org/10.2147/OTT.S60888
Checked for plagiarism Yes
Review by Single anonymous peer review
Peer reviewer comments 3
Jingxu Sun,1,* Yongxi Song,1,* Zhenning Wang,1 Guoli Wang,2 Peng Gao,1 Xiaowan Chen,1 Zhaohua Gao,1 Huimian Xu1
1Department of Surgical Oncology and General Surgery, First Hospital of China Medical University, Shenyang, People’s Republic of China; 2Department of Biochemistry and Molecular Biology, China Medical University, Shenyang, People’s Republic of China
*These authors contributed equally to this work
Background: MicroRNAs are associated with tumor genesis and progression in various carcinomas. MicroRNA-148a (miR-148a) was reported to have low expression in gastrointestinal cancers, and might be regulated by promoter region DNA methylation.
Methods: Bisulfite-modified sequencing was used to determine the promoter region DNA methylation status of human gastrointestinal cancer cell lines. Expression levels of miR-148a in cell lines treated with 5-aza-2′-deoxycytidine were determined by quantitative real-time polymerase chain reaction. Total DNA was extracted from the tissues of 64 patients with gastric cancer and 51 patients with colorectal cancer. Methylation status was determined by methylation-specific polymerase chain reaction. All statistical analyses were performed with SPSS 17.0 software.
Results: The promoter regions of genes in human gastrointestinal cancer cell lines were all hypermethylated, except for HT-29, and the expression of miR-148a tended to be higher than in controls after treatment with 5-aza-2′-deoxycytidine. The methylation-specific polymerase chain reaction results showed that 56.25% of gastric cancer tissues and 19.61% of colorectal cancer tissues were hypermethylated. A strong correlation was found between the expression of miR-148a and the methylation status of promoter regions (P<0.001, chi-square test and Pearson’s correlation). Furthermore, promoter region CpG site hypermethylation of miR-148a was correlated with increased tumor size (P=0.01) in gastric cancer after analyzing the correlation between methylation status and clinicopathologic characteristics.
Conclusion: The promoter region CpG sites were hypermethylated in gastrointestinal cancers. Promoter region hypermethylation status was associated with the expression of miR-148a and tumor invasiveness in gastric cancer, and may prove to be a new biomarker and method for treating gastric cancer.
Keywords: microRNAs, methylation, CpG site, clinicopathologic characteristics
Background
Gastrointestinal cancers, especially gastric and colorectal cancers (CRCs), are a major health concern worldwide. Gastric cancer is the fourth most common cancer and the second leading cause of cancer-related death in the world.1 CRC is the third most commonly diagnosed malignancy among humans and has become a growing health problem in China in recent years.2,3 Although treatment strategies have improved, the overall prognosis remains unsatisfactory. MicroRNAs (miRNAs) are a class of small noncoding RNAs that have been considered to be gene expression regulators,4 are involved in various cell functions, and may contribute to many different human cancers.5,6 Recently, an increasing number of studies have found that miRNAs may be potential biomarkers in the diagnosis and prognosis of gastrointestinal cancer.7–11
A general downregulation of miRNAs has been shown in human cancers,12 and miRNA-148a (miR-148a) is one of the most downregulated miRNAs that have been found in gastrointestinal, ovarian, and pancreatic cancer.13–15 In our previous study,13 we reported that miR-148a was downregulated in a large number of gastrointestinal cancer tissues. However, the mechanisms leading to its downregulation were unclear and may have been regulated by many factors, including deletions, mutations, transcription factors, and aberrant methylation. DNA hypermethylation of promoter CpG islands is an important process in DNA epigenetic alterations and is considered to be associated with silencing tumor suppressor genes and mediating tumorigenesis, invasiveness, and metastasis.16,17 Recently, Lehmann et al18 studied the correlation between methylation and miR-148a in breast cancer. In the same year, Lujambio et al19 reported that DNA methylation-associated silencing of tumor suppressor miRNAs contributes to the development of human cancer metastasis. Hanoun et al20 indicated that the silencing of miR-148a production by hypermethylation affected pancreatic carcinogenesis. Furthermore, Zhu et al21 reported that miR-148a was silenced by hypermethylation and interacted with DNA methyltransferase (DNMT)1 in gastric cancer. Takahashi et al22 found, in CRC, that the expression of miR-148a may be regulated by methylation status. Aberrant DNA hypermethylation has been expected to play an important role as a suitable progression biomarker for cancer progression and outcomes. Although there have been several reports about aberrant methylation of miR-148a, further evidence of the function of reliable prognostic biomarkers from large, well-defined sample studies in gastric cancers and CRCs is needed.18–22
The aim of this study was to identify the suppression of miR-148a, whose expression may be inactivated by DNA hypermethylation in gastric cancers and CRCs. In addition, we will discuss the associations between promoter region hypermethylation of miR-148a and clinicopathologic characteristics, which may be helpful for developing new prognostic biomarkers in gastric cancers and CRCs.
Materials and methods
Human tissue samples
Sixty-four pairs of human gastric cancer tissue samples and 51 pairs of human CRC tissue samples were obtained from patients that had undergone surgical resection at the First Hospital of China Medical University, Shenyang, People’s Republic of China and were diagnosed with gastric cancer and CRC based on histopathological evaluations. The matched non-tumor tissue was obtained from the section of the resected specimens that was farthest from the tumor tissue (>5 cm). The fresh samples were snap-frozen in liquid nitrogen and stored at −80°C. The samples used for histopathological evaluation were stained with hematoxylin–eosin. The histological grade of a tumor was assessed according to the World Health Organization criteria and was staged using the TNM staging of the Union for International Cancer Control (UICC)/American Joint Committee on Cancer (AJCC) system (2010). The present study was approved by the research ethics committee of China Medical University. Written informed consents were signed by all included patients.
Cell lines and culture conditions
Human gastric cancer cell lines (MGC-803, BGC-823, and SGC-7901) and CRC cell lines (HT-29 and HCT-116) were obtained from the Institute of Biochemistry and Cell Biology at the Chinese Academy of Sciences (Shanghai, People’s Republic of China). MGC-803, BGC-823, and SGC-7901 were cultured in RPMI 1640 medium (Thermo Fisher Scientific, Waltham, MA, USA) and HT-29 and HCT-116 were cultured in McCoy’s 5A medium (Thermo Fisher Scientific). The media were supplied with 10% fetal bovine serum and cultured at 37°C in 5% CO2. For inhibition of DNMT activity, we used 3 μmol/L (Sigma-Aldrich, St Louis, MO, USA) to treat cells for 72 hours. Treatment with dimethyl sulfoxide served as a control. Initial experiment and report showed that, under these conditions, cells could achieve maximal demethylation with minimal toxic effects.23
Extraction of total RNA and real-time polymerase chain reaction (PCR)
Total RNA was extracted from cell lines using RNAiso Plus (Takara Bio Inc., Shiga, Japan), according to the manufacturer’s protocol. The concentration was determined by ultraviolet-visible spectrophotometry (A260/A280<1.9). Then, total RNA was polyadenylated with adenosine triphosphate by Escherichia coli poly(A) polymerase at 37°C for 30 minutes, following the manufacturer’s protocol for a Poly(A) Tailing Kit (Thermo Fisher Scientific).24 The RNA extracted with poly(A) tail was used for a reverse transcriptase PCR to get cDNA and the cDNA was subsequently held at 4°C. Real-time PCR was performed using SYBR Premix Ex Taq II (Takara Bio Inc.) on a Rotor-Gene 6000 system (QIAGEN, Valencia, CA, USA). The reaction was incubated in a 36-well optical plate by 45 amplification cycles of 94°C for 5 seconds, 58°C for 20 seconds, and 72°C for 30 seconds.13 We used the following special primers for miR-148a: forward 5′-TCAGTGCACTACAGAACTTTGT-3′ and reverse 5′-GCTGTCAACGATACGCTACGT-3′. We also used U6 RNA as an endogenous reference for normalizing the expression levels of miR-148a. Results were normalized to the concentration of U6 RNA and calculated with the method of 2−ΔΔCT, where ΔCT was the difference in threshold cycle (CT) values.
Extraction of genomic DNA and bisulfite modification
Genomic DNA from the cultured cells and specimens was isolated using a Universal Genomic DNA Extraction Kit Ver3.0 (Takara Bio Inc.). The concentration and purity of DNA were controlled by ultraviolet-visible spectrophotometry (1.8<A260/A280<2.0). The genomic DNA was dissolved with ddH2O and stored at −80°C. The genomic DNA of each specimen was treated with bisulfite modification and the protocol was as per Herman et al.25 Genomic DNA (2 μg) was treated with sodium bisulfite (Sigma-Aldrich). In brief, genomic DNA was denatured in 3 mol/L NaOH for 15 minutes at 37°C. Cytosines were sulfonated in 3.6 mol/L sodium bisulfite and 1 mmol/L hydroquinone (Sigma-Aldrich) for 16 hours at 55°C. The modified DNA samples were desalted using a DNA clean-up system (Promega Corporation, Fitchburg, WI, USA). The modified DNA samples were dissolved with ddH2O and stored at −80°C.
DNA methylation bisulfite-modified sequencing
The University of California, Santa Cruz’s Genome Technology Center26 was used to search the sequence of miR-148a, pri-miR-148a, and pre-miR-148a. Results showed that most core promoters of miRNAs may be close to pre-miRNAs, and the possibility was >90% confined to 500 bp in the upstream region of the pre-miRNA.27 According to the sequence from the Genome Technology Center, we analyzed the distribution of CpG sites confined to 1,000 bp upstream and 500 bp downstream of miR-148a with Methyl Primer Express® software (v 1.0; Thermo Fisher Scientific). Referring to studies18,19,21 on miR-148a and the results from our analysis, we chose the sequence that was between 399 bp upstream and 130 bp downstream of mature miR-148a to sequence. The bisulfite-modified sequencing primers were designed with Methyl Primer Express software. We performed two sequencing runs of miR-148a with the first pair of primers (forward 5′-ATAAGTTAGAGTGAGTTAGTTGTTGGT-3′ and reverse 5′-TCCTACAAATATATTCCCCCAC-3′) and the second pair of primers (forward 5′-TGGGTATTTGTTTTTGTTGATTG-3′ and reverse 5′-ACTACACTTAAACCCCCTCAACC-3′). The whole process included DNA methylation bisulfite modification; PCR amplification, electrophoresis, and retrieval; PCR products connected to the pUC18-T vector and transformation; blue/white plaque selection; and extraction of plasmids, and the final process involved sequencing with the DNA of the plasmids. Each cell line had five clones from the blue/white plaque selection from sequencing. The whole process of sequencing was performed by Sangon Biotech (Shanghai, People’s Republic of China) with genomic DNA of the cell lines and the primers for sequencing provided by our group.
Methylation-specific PCR (MSP)
We designed the primers for MSP according to the results of bisulfite-modified sequencing: methylated forward 5′-GCGATTAGAATTTGTCGAATC-3′, reverse 5′-AAATACGACCGATAATAAAACG-3′; and unmethylated forward 5′-ATTGTGATTAGAATTTGTTGAATT-3′, reverse 5′-AAAATACAACCAATAATAAAACAAA-3′. The amplicon sizes were 114 bp and 109 bp for methylated and unmethylated productions, respectively. We used treated DNA in PCR amplification with TaKaRa Taq Hot Start Version (code number DR007A; Takara Bio Inc.). The PCRs were conducted under the following thermocycling conditions: 5 minutes at 94°C; 40 cycles of 30 seconds at 94°C; 30 seconds at 58°C; 30 seconds at 72°C; and a final incubation at 72°C for 10 minutes. Controls without DNA were used for each set of PCRs. Aliquots of 5 μL of the total 20 μL of the PCR mixture were loaded onto 3% agarose gels, stained with ethidium bromide, and directly visualized under ultraviolet illumination. MSP assays were repeated at least twice for each sample to determine the reproducibility of the results.
Statistical analysis
The expression level of miRNAs in tumors relative to controls was calculated using the method of 2−ΔΔCT, where ΔΔCT is the difference of the ΔCT value between the treatment tumor cells and the control:
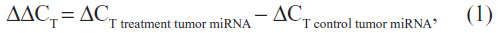
and ΔCT is the difference of the CT value between the target and endogenous reference:
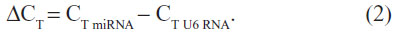
If the expression level of the treatment cells and the control cells was equal, ΔΔCT equaled 0 and 2−ΔΔCTequaled 1.28 Therefore, we could compare the expression level of miR-148a in gastric cancer and CRC cell lines by comparing the value of ΔCT treatment tumor miRNA and ΔCT control tumor miRNA. The results of MSP were expressed as hypermethylation status and hypomethylation status. All statistical analyses were performed using SPSS software (v 17.0; SPSS Inc., Chicago, IL, USA). The statistical significance between expression levels of miR-148a and the methylation status in gastric and colorectal tissues was calculated using a chi-square test (2×2 table Pearson’s analysis). Correlations between methylation status and clinicopathologic parameters were analyzed with a chi-square test and nonparametric test: Mann–Whitney U test between two groups and Kruskal–Wallis test for three or more groups. P<0.05 was considered significant.
Results
CpG island DNA methylation analysis of the miR-148a gene promoter region in the cell lines
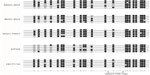
To identify the presence of CpG island methylation of miR-148a in gastric cancer and CRC cell lines, we undertook bisulfite-modified sequencing analyses of multiple clones in the cell lines. The CpG island is depicted and each single CpG site is illustrated by a vertical bar. Black and white circles represent hypermethylation and hypomethylation, respectively (Figure 1). We sequenced all 13 CpG sites in 399 bp upstream of the mature miR-148a. The results showed that the methylation level of the promoter regions in the gastric cancer cell line MGC-803 was 80.00% (53/65). In BGC-823, the methylation level was 78.46% (51/65), and in SGC-7901 it was 92.31% (60/65). In CRC cell line HT-29, the methylation level was 47.69% (31/65), and in HCT-116 it was 96.92% (63/65) (Figure 1). The CpG sites in 150 bp downstream of the mature miR-148a were all hypermethylated; in MGC-803, BGC-823, HT-29, and HCT-116, the methylation level was 97.5% (39/40), and the level in SGC-7901 was 95% (38/40).
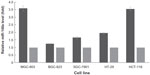
To further show that miR-148a was aberrantly inhibited by DNA hypermethylation, we treated gastric cancer and CRC cell lines with 5-aza-2′-deoxycytidine at a final concentration of 3 μmol/L and analyzed the expression of miR-148a after 72 hours. The values of 2−ΔΔCTwere 3.5801 in MGC-803, 1.2483 in BGC-823, 1.6586 in SGC-7901, 1.9588 in HT-29, and 3.5554 in HCT-116 cell lines. The values of 2−ΔΔCTwere all 1.00 in each of the control cell lines. The results showed that the expression of miR-148a was unregulated in all of the treated cell lines but not in the control cell lines, which indicated that hypermethylation might affect the expression of miR-148a in gastric cancer and CRC cell lines (Figure 2).
CpG island DNA methylation analysis of the miR-148a gene promoter region in tissues
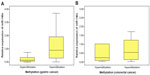
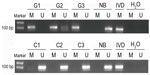
The methylation levels in gastric cancer and CRC tissues were analyzed by MSP. The tissues were treated with bisulfite and PCR was performed with special primers. The PCR results showed hypermethylation or hypomethylation according to the electrophoretic bands. In the 64 gastric cancer tissues, there were 36 samples that showed hypermethylation (56.25%) and 28 samples that showed hypomethylation (43.75%). In the 51 CRC tissues, there were ten hypermethylation samples (19.61%) and 41 hypomethylation samples (80.39%) (Figure 3). The MSP results were used to analyze correlation with the expression of miR-148a and clinicopathologic features in patients with gastric cancer and CRC.
The correlations between methylation and the expression of miR-148a in gastric cancer and CRC samples
The methylation levels were determined to be hypermethylated and hypomethylated, and the expression of miR-148a was determined to be hyperexpressed or hyporexpressed. A strong correlation between CpG island DNA methylation levels of the miR-148a gene promoter region and the expression of the miR-148a in gastric cancer tissues was demonstrated with Pearson’s regression analysis (P<0.01). The correlation coefficient was 0.622. The result showed that hypermethylation of the miR-148a gene promoter region was correlated with low expression of miR-148a in patients with gastric cancer. We also analyzed the correlations between methylation of miR-148a and the expression of miR-148a in CRC tissues, but the results showed that there was no statistical significance (P=0.23) (Table 1 and Figure 4).
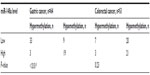  | Table 1 The correlations between methylation and the expression of microRNA-148a (miR-148a) in gastrointestinal cancer samples |
The correlation between methylation levels and clinicopathologic characteristics
We analyzed the correlations between the methylation status of miR-148a with clinicopathologic characteristics including sex, age, macroscopic type, pN stage, clinical stage, lymphatic vessel invasion, histologic grade, tumor location, and tumor size. We found that hypermethylation of the miR-148a gene promoter region was associated with increased tumor size (P=0.01) in gastric cancer samples. The correlation coefficient was −0.323 (Table 2). The results showed that gastric cancer patients with large tumor size may have had hypermethylation in the miR-148a gene promoter region. We also analyzed the correlations between methylation of miR-148a and the clinicopathologic characteristics of CRC tissues, but the results showed that there was no statistical significance (Table 3).
Discussion
Recently, the expression of miRNAs has been reported to be abnormal in most tumors and the function of miRNAs in tumor formation and development is critical. Our previous study analyzed 101 patients with gastric cancer and 101 patients with CRC as well as their matched non-tumor adjacent tissues, and we found that the expression of miR-148a was downregulated in gastric cancer and CRC.11 However, the cause of the changes remains unclear. The mechanisms that may regulate the expression of miRNAs include the following: DNA methylation and histone modification;19,23,29 regulation of transcription factors;30 regulation of splicing factors;31 gene mutation;32 single-nucleotide polymorphisms (SNPs);33 and downregulated expression of dicer, among others.34 The role of epigenetic modifications in the etiology of human disease has attracted the attention of researchers.35 As one of the most important types of epigenetic modifications, DNA methylation has frequently been reported as regulating gene transcription in the process of tumorigenesis.36,37 The epigenetic regulation of miRNA expression has been found in several kinds of cancers.38 When the promoter is in hypermethylation status, the process of transcription and the function of the tumor suppressor genes are inhibited. Furthermore, Lujambio et al19 reported that methylation-mediated silencing of miR-148a occurred in metastatic colonic cancer. In a study done by Zhu et al,21 the authors reported that the expression of miR-148a was inactivated by DNA hypermethylation in gastric cancer cell lines and 38 gastric cancer samples. The data from the current study suggest that hypermethylation of the promoter region may contribute to downregulation of the expression of miR-148a in gastric cancer and CRC patients.
In our previous research,13 we reported that the expression of miR-148a was downregulated in gastrointestinal cancer cell lines and tissues. Similar results were also indicated by Zhu et al21 in gastric cancer and Takahashi et al22 in CRC. In the present study, we used bisulfite sequencing to analyze the methylation status of gastric cancer and CRC cell lines, and, ultimately, downregulation of miR-148a was found. The results showed that the methylation levels of the promoter region were all more than 75% in gastric cell lines. This meant that, in gastric cell lines, the DNA in CpG islands of the promoter region were all hypermethylated. In CRC cell lines, the methylation levels were 46.03% (HT-29) and 96.88% (HCT-116). The sequencing results indicated that methylation status in different CRC cell lines may not be accordant with the results from gastric cancer cell lines. Furthermore, we confirmed that the expression of miR-148a was upregulated by treatment with the DNA methylation inhibitor 5-aza-2′-deoxycytidine in gastrointestinal cancer cell lines. The results indicated that DNA hypermethylation of the promoter region may be one of the key factors that causes silencing of miR-148a in gastric cancer and CRC.
Based on the results of bisulfite sequencing, we designed the primers for MSP with the sequences of the miR-148a promoter region. MSP is a simple, sensitive, and specific method for researching the methylation status of CpG-rich regions. Since it was first reported in 1996 by Herman et al25 for mapping DNA methylation patterns in CpG islands, many researchers have used this technology to analyze the methylation status in regions that affect the expression of microRNAs.19,39,40 In the present study, we chose 64 gastric cancer tissues and 51 CRC tissues from our previous study to analyze the promoter region methylation status of miR-148a with MSP. From the results of MSP, the hypermethylation rate in gastric cancer tissues was 56.25%, and in CRC tissues the hypermethylation rate was 19.61%. Then, we analyzed the correlation between the methylation levels and the expression of miR-148a in gastric cancer and CRC tissues. The results showed that, in gastric cancer, the downregulated expression of miR-148a had a significant correlation with CpG island DNA hypermethylation of the promoter region. Our study indicates that hypermethylation of the promoter region may be an effective factor for suppressing the expression of miR-148a in gastric cancer. A similar result in gastric cancer was reported by Zhu et al.21
The antitumor roles of miR-148a in gastric cancer have been reported in several studies. Zhu et al21 reported that downregulated expression of miR-148a in gastric cancer was associated with proliferation and invasion. Moreover, the results from Guo et al41 indicated that miR-148a downregulation might affect some antagonistic functions in gastric cancer cell proliferation during the process of tumorigenesis. In the present study, we analyzed the correlation between the methylation status of miR-148a and clinicopathologic characteristics in a large sample of gastric cancer. We found that methylation status of miR-148a was associated with the expression level of miR-148a and tumor size. In the same sample of gastric cancer, we reported that patients with lower expression of miR-148a tended to have larger tumor size in our previous study.13 These results show that hypermethylation of the promoter region might repress the expression of miR-148a and low expression of miR-148a might contribute to proliferation and invasion in gastric cancer.
However, we also analyzed the correlation between methylation status of the promoter region and expression of miR-148a in CRC tissues. There was no significant correlation between them. However, the expression of miR-148a may be affected by hypermethylation in metastatic colonic cancer,19 but our results do not provide evidence on the correlation between methylation status and expression of miR-148a in CRC tissues. It has been reported that the expression of miR-148a may be affected by epigenetic regulation in CRC22 and the results from our real-time PCR also indicated a trend that methylation status might affect the expression of miR-148a in CRC cell lines. We still cannot draw a conclusion about methylation status and the expression of miR-148a in CRC tissues based on our results. In the present study, we also analyzed the correlation between methylation status and clinicopathologic characteristics in CRC tissues. The results showed that there was no significant association between methylation and clinicopathologic characteristics. However, in our previous study,13 we found that low expression of miR-148a was associated with a more advanced pT stage. Therefore, we thought that there might be other factors that could affect the expression of miR-148a besides methylation in CRC, such as histone modification or SNP variation. Many other mechanisms should also be analyzed for a definitive conclusion in future studies.
It is interesting that several studies have reported the relationship between miR-148 and DNMTs, which are important for DNA methylation.42–45 In the study reported by Duursma et al, the authors indicated that the coding region of human genes could be targeted by miRNAs, and overexpression of miR-148 could downregulate the expression of DNMT3b1 among various cell lines.42 Similarly, Pan et al43 and Yan et al44 reported that miR-148a could repress the expression of DNMT1 in systemic lupus erythematosus and gastric cancer. In the study reported by Xu et al, the researchers found that DNMT1 was overexpressed in breast cancer, and the overexpression was responsible for hypermethylation of the miR-148a promoter region.45 Based on the results from our research and others, we supposed that the correlation between the methylation status of miR-148a promoter region and DNMTs might be an important process in gastrointestinal cancer. More studies on molecular mechanisms are needed in the future.
The present research was based on the results of our previous study13 on miR-148a in gastrointestinal cancers. We sequenced the CpG site and determined the methylation status of miR-148a. Then, we analyzed the correlation between the methylation status of miR-148a and clinicopathologic characteristics, which are associated with the proliferative and invasive behavior of tumors. The results showed that there was an aberrant hypermethylation status in the CpG islands of the promoter region of miR-148a in gastrointestinal cancers, and especially in gastric cancer. Our findings also indicated that CpG island hypermethylation-mediated silencing of miR-148a with tumor suppressor features contributed to gastrointestinal cancer. The methylation status could be used as a biomarker to predict tumor prognosis and biologic behavior in gastric cancer. Furthermore, the results might provide a molecular basis for treating cancer patients with drugs that have demethylating activity. We speculated that the CpG island methylation status might be one of the mechanisms of the downregulation of miR-148a in gastrointestinal cancer, and we will continue to research other possible causes of miR-148a expression regulation, such as SNP variations. Further investigation is needed to confirm our results.
Conclusion
This is the first study to use bisulfite-modified sequencing to determine the CpG site methylation status of miR-148a in gastrointestinal cancer cell lines. In the present study, we found that hypermethylation of the promoter region was associated with expression of miR-148a in gastric cancer. To our knowledge, we are also the first to find that the hypermethylation status was associated with large tumor sizes by analyzing clinicopathologic characteristics, which could be considered predictive of tumor prognosis and biologic behavior in gastric cancer tissues. Our study may provide a new biomarker and method for treating patients with gastric cancer, and large-scale and long-term follow-up studies about methylation status and miR-148a in CRC should be undertaken in the future.
Acknowledgments
We thank the Department of Surgical Oncology at the First Hospital of China Medical University for providing human gastric tissue samples. We also thank the College of China Medical University for technical assistance in experiments. This work was supported by the National Natural Science Foundation of China (81201888, 81372549, and 81172370), Specialized Research Fund for the Doctoral Program of Higher Education (20122104110009), and the Project of Science and Technology of Shenyang (F12-193-9-08).
Disclosure
The authors report no conflicts of interest in this work.
References
Ferlay J, Shin HR, Bray F, Forman D, Mathers C, Parkin DM. Estimates of worldwide burden of cancer in 2008: GLOBOCAN 2008. Int J Cancer. 2010;127(12):2893–2917. | |
Cunningham D, Atkin W, Lenz HJ, et al. Colorectal cancer. Lancet. 2010;375(9719):1030–1047. | |
Yang L, Parkin DM, Li L, Chen Y. Time trends in cancer mortality in China: 1987–1999. Int J Cancer. 2003;106(5):771–783. | |
Bartel DP. MicroRNAs: genomics, biogenesis, mechanism, and function. Cell. 2004;116(2):281–297. | |
Iorio MV, Croce CM. MicroRNAs in cancer: small molecules with a huge impact. J Clin Oncol. 2009;27(34):5848–5856. | |
Krützfeldt J, Poy MN, Stoffel M. Strategies to determine the biological function of microRNAs. Nat Genet. 2006;38 Suppl:S14–S19. | |
Esquela-Kerscher A, Slack FJ. Oncomirs – microRNAs with a role in cancer. Nat Rev Cancer. 2006;6(4):259–269. | |
Cho WC. OncomiRs: the discovery and progress of microRNAs in cancers. Mol Cancer. 2007;6:60. | |
Huang Z, Huang D, Ni S, Peng Z, Sheng W, Du X. Plasma microRNAs are promising novel biomarkers for early detection of colorectal cancer. Int J Cancer. 2010;127(1):118–126. | |
Chiang Y, Song Y, Wang Z, et al. Aberrant expression of miR-203 and its clinical significance in gastric and colorectal cancers. J Gastrointest Surg. 2011;15(1):63–70. | |
Song YX, Yue ZY, Wang ZN, et al. MicroRNA-148b is frequently down-regulated in gastric cancer and acts as a tumor suppressor by inhibiting cell proliferation. Mol Cancer. 2011;10:1. | |
Lu J, Getz G, Miska EA, et al. MicroRNA expression profiles classify human cancers. Nature. 2005;435(7043):834–838. | |
Chen Y, Song Y, Wang Z, et al. Altered expression of MiR-148a and MiR-152 in gastrointestinal cancers and its clinical significance. J Gastrointest Surg. 2010;14(7):1170–1179. | |
Zhou X, Zhao F, Wang ZN, et al. Altered expression of miR-152 and miR-148a in ovarian cancer is related to cell proliferation. Oncol Rep. 2012;27(2):447–454. | |
Delpu Y, Lulka H, Sicard F, et al. The rescue of miR-148a expression in pancreatic cancer: an inappropriate therapeutic tool. PLoS One. 2013;8(1):e55513. | |
Wolffe AP, Matzke MA. Epigenetics: regulation through repression. Science. 1999;286(5439):481–486. | |
Taby R, Issa JP. Cancer epigenetics. CA Cancer J Clin. 2010;60(6):376–392. | |
Lehmann U, Hasemeier B, Christgen M, et al. Epigenetic inactivation of microRNA gene hsa-mir-9-1 in human breast cancer. J Pathol. 2008;214(1):17–24. | |
Lujambio A, Calin GA, Villanueva A, et al. A microRNA DNA methylation signature for human cancer metastasis. Proc Natl Acad Sci U S A. 2008;105(36):13556–13561. | |
Hanoun N, Delpu Y, Suriawinata AA, et al. The silencing of microRNA 148a production by DNA hypermethylation is an early event in pancreatic carcinogenesis. Clin Chem. 2010;56(7):1107–1118. | |
Zhu A, Xia J, Zuo J, et al. MicroRNA-148a is silenced by hypermethylation and interacts with DNA methyltransferase 1 in gastric cancer. Med Oncol. 2012;29(4):2701–2709. | |
Takahashi M, Cuatrecasas M, Balaguer F, et al. The clinical significance of MiR-148a as a predictive biomarker in patients with advanced colorectal cancer. PLoS One. 2012;7(10):e46684. | |
Saito Y, Liang G, Egger G, et al. Specific activation of microRNA-127 with downregulation of the proto-oncogene BCL6 by chromatin-modifying drugs in human cancer cells. Cancer Cell. 2006;9(6):435–443. | |
Shi R, Chiang VL. Facile means for quantifying microRNA expression by real-time PCR. Biotechniques. 2005;39(4):519–525. | |
Herman JG, Graff JR, Myöhänen S, Nelkin BD, Baylin SB. Methylation-specific PCR: a novel PCR assay for methylation status of CpG islands. Proc Natl Acad Sci U S A. 1996;93(18):9821–9826. | |
UCSC Genome Bioinformatics [homepage on the Internet]. Santa Cruz, CA: University of California, Santa Cruz. Available from: http://genome.ucsc.edu/. Accessed May 23, 2014. | |
Zhou X, Ruan J, Wang G, Zhang W. Characterization and identification of microRNA core promoters in four model species. PLoS Comput Biol. 2007;3(3):e37. | |
Livak KJ, Schmittgen TD. Analysis of relative gene expression data using real-time quantitative PCR and the 2(-Delta Delta C(T)) Method. Methods. 2001;25(4):402–408. | |
Wilting SM, van Boerdonk RA, Henken FE, et al. Methylation-mediated silencing and tumour suppressive function of hsa-miR-124 in cervical cancer. Mol Cancer. 2010;9:167. | |
Shin VY, Jin H, Ng EK, et al. NF-ºB targets miR-16 and miR-21 in gastric cancer: involvement of prostaglandin E receptors. Carcinogenesis. 2011;32(2):240–245. | |
Guil S, Cáceres JF. The multifunctional RNA-binding protein hnRNP A1 is required for processing of miR-18a. Nat Struct Mol Biol. 2007;14(7):591–596. | |
Aqeilan RI, Calin GA, Croce CM. miR-15a and miR-16-1 in cancer: discovery, function and future perspectives. Cell Death Differ. 2010;17(2):215–220. | |
Jazdzewski K, Murray EL, Franssila K, Jarzab B, Schoenberg DR, de la Chapelle A. Common SNP in pre-miR-146a decreases mature miR expression and predisposes to papillary thyroid carcinoma. Proc Natl Acad Sci U S A. 2008;105(20):7269–7274. | |
Martello G, Rosato A, Ferrari F, et al. A MicroRNA targeting dicer for metastasis control. Cell. 2010;141(7):1195–1207. | |
Egger G, Liang G, Aparicio A, Jones PA. Epigenetics in human disease and prospects for epigenetic therapy. Nature. 2004;429(6990):457–463. | |
Bird A. DNA methylation patterns and epigenetic memory. Genes Dev. 2002;16(1):6–21. | |
Stein RA. DNA methylation profiling: a promising tool and a long road ahead for clinical applications. Int J Clin Pract. 2011;65(12):1212–1213. | |
Akhavan-Niaki H, Samadani AA. DNA methylation and cancer development: molecular mechanism. Cell Biochem Biophys. 2013;67(2):501–513. | |
Wang P, Chen L, Zhang J, et al. Methylation-mediated silencing of the miR-124 genes facilitates pancreatic cancer progression and metastasis by targeting Rac1. Oncogene. 2014;33(4):514–524. | |
Pavicic W, Perkiö E, Kaur S, Peltomäki P. Altered methylation at microRNA-associated CpG islands in hereditary and sporadic carcinomas: a methylation-specific multiplex ligation-dependent probe amplification (MS-MLPA)-based approach. Mol Med. 2011;17(7–8):726–735. | |
Guo SL, Peng Z, Yang X, et al. miR-148a promoted cell proliferation by targeting p27 in gastric cancer cells. Int J Biol Sci. 2011;7(5):567–574. | |
Duursma AM, Kedde M, Schrier M, le Sage C, Agami R. miR-148 targets human DNMT3b protein coding region. RNA. 2008;14(5):872–877. | |
Pan W, Zhu S, Yuan M, et al. MicroRNA-21 and microRNA-148a contribute to DNA hypomethylation in lupus CD4+ T cells by directly and indirectly targeting DNA methyltransferase 1. J Immunol. 2010;184(12):6773–6781. | |
Yan J, Guo X, Xia J, et al. MiR-148a regulates MEG3 in gastric cancer by targeting DNA methyltransferase 1. Med Oncol. 2014;31(3):879. | |
Xu Q, Jiang Y, Yin Y, et al. A regulatory circuit of miR-148a/152 and DNMT1 in modulating cell transformation and tumor angiogenesis through IGF-IR and IRS1. J Mol Cell Biol. 2013;5(1):3–13. |
 © 2014 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.
© 2014 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.