Back to Journals » OncoTargets and Therapy » Volume 13
Circular RNA circEXOC6B Inhibits the Progression of Ovarian Cancer by Sponging miR-421 and Regulating RUS1 Expression
Authors Wang Z, Zhang W, Fang J, Xie P, Miao M, Yang H
Received 19 December 2019
Accepted for publication 9 April 2020
Published 19 August 2020 Volume 2020:13 Pages 8233—8243
DOI https://doi.org/10.2147/OTT.S243040
Checked for plagiarism Yes
Review by Single anonymous peer review
Peer reviewer comments 2
Editor who approved publication: Dr William C. Cho
Zhonghai Wang,1 Wenmin Zhang,2 Jinchuan Fang,2 Ping Xie,2 Miao Miao,2 Hongwen Yang2
1Department of Gynecology, Huazhong University of Science and Technology, Shenzhen, Guangdong 518052, People’s Republic of China; 2Women & Children Health Institute Futian Shenzhen, Shenzhen, Guangdong 518017, People’s Republic of China
Correspondence: Hongwen Yang
Department of Gynecology, Women & Children Health Institute Futian Shenzhen, No. 2002 Jintian Road, Futian District, Shenzhen, Guangdong 518017, People’s Republic of China
Email [email protected]
Background: Evidence has been shown that circular RNAs (circRNAs) play a vital role during the development of ovarian cancer. However, the mechanism by which circEXOC6B regulates tumorigenesis of ovarian cancer remains unknown. Thus, this study aimed to investigate the role of circEXOC6B during the progression of ovarian cancer.
Materials and Methods: The dual-luciferase reporter system assay was used to determine the interaction between circEXOC6B, miR-421 and RUS1 in ovarian cancer, respectively. CCK8 and colony formatting were used to evaluate cell proliferation. Meanwhile, the expressions of RSU1, PINCH1 and ILK in SKOV3 cells were detected with Western blot.
Results: Downregulation of circEXOC6B markedly promoted the proliferation and invasion in A2780 cells. In contrast, upregulation of circEXOC6B significantly inhibited the proliferation and invasion in SKOV3 cells. Moreover, overexpression of circEXOC6B obviously induced the apoptosis of SKOV3 cells. Furthermore, luciferase reporter assay identified that miR-421 was the potential miRNA binding of circEXOC6B, and RUS1 was the potential binding target of miR-421. Mechanism analysis indicated that upregulation of circEXOC6B increased the level of RUS1 by acting as a competitive “sponge” of miR‐421.
Conclusion: In this study, we found that circEXOC6B suppressed the growth of ovarian cancer cells through upregulating RSU1 partially via sponging miR-421. Therefore, circEXOC6B might be a potential target for the treatment of ovarian cancer.
Keywords: ovarian cancer, circular RNAs, circEXOC6B, miR-421, TUS1
Introduction
Ovarian cancer is the seventh-eighth most common cancer among women.1,2 Epithelial ovarian cancer (EOC) is the most common type of ovarian cancer, accounting for 85% to 90% of ovarian malignancies.3 Ovarian cancer is generally classified into five histological subtypes according to clinical features and treatments, including endometrioid, clear cell, mucinous, high grade serous, and low grade serous.4 Surgery, radiotherapy, and chemotherapy are the current primary clinical therapies for the treatment of ovarian cancer.5,6 However, the prognosis of ovarian cancer remains poor with the 5-year survival rate of approximately 40%.7 Meanwhile, the recurrence rate is high in patients with ovarian cancer.8 Therefore, identify of novel clinical biomarkers may help to understand the diagnosis, prognosis and treatment of ovarian cancer.
Circular RNAs (circRNAs) are a kind of non-coding RNA (ncRNA) molecules without a 5′ cap and 3′ poly A tail.9 CircRNAs are more stable than linear RNAs because of the resistance to exonuclease.10 Most circRNAs derive from exons of protein-coding genes, while few circRNAs originate from introns or exon–introns.11 Previous study reported that circRNAs play important roles during the progression and development of cancers including gastric cancer, prostate cancer and colorectal cancer.12–14 Furthermore, circRNAs play important roles in ovarian tumor growth via functioning as oncogenes or tumor suppressor genes.9,15 In addition, some circRNAs have been found to regulate gene expression through functioning as competing endogenous RNAs (ceRNAs) at the transcriptional or post-transcriptional level.16 Ning et al indicated that the level of circEXOC6B was markedly downregulated in EOC specimens, compared with normal ovarian tissues.17 Meanwhile, circEXOC6B may function as a prognostic indicator in patients with EOC.17 However, the mechanisms by which circEXOC6B regulates the proliferation, apoptosis and invasion in ovarian cancer remain unclear.
In this study, we identified circEXOC6B harbored conserved miR-421 cognate sites, and then we made an assumption that circEXOC6B might function as a ceRNA for miR-421. MiR-421 has shown to play a vital role in cancer progression, such as gastric cancer, colorectal cancer and cervical cancer.18–20 Furthermore, TargetScan predicted that Ras Suppressor-1 (RUS1) was a potential target of miR-421. RSU1 functions as a tumor suppressor, which could inhibit cell migration.21 Collectively, we identified circEXOC6B might be a tumor suppressor involved in the progression of ovarian cancer via functioning as a ceRNA for miR-421, and activating RUS1 signaling.
Materials and Methods
Cell Culture
Normal ovarian epithelial IOSE80 cells were obtained from Lonza (Basel, Switzerland). Human ovarian cancer cell lines A2870, SKOV3 and OVCAR3 were provided by American Type Culture Collection (ATCC, Rockville, MD, USA). Cells were cultured in Dulbecco’s modified Eagle’s medium (DMEM, Thermo Fisher Scientific, Waltham, MA, USA) supplemented with 10% FBS and 100 U/mL of penicillin/streptomycin, and maintained at 37°C in the presence of 5% CO2.
Quantitative Real‑time Polymerase Chain Reaction (qRT‑PCR)
Trizol reagent (Thermo Fisher Scientific) was used to extract total RNA from ovarian cancer cells. High Capacity cDNA Reverse Transcription Kit (Thermo Fisher Scientific) was used to synthesize cDNA. To detect miR-421 expression, PrimeScript™ RT Master Mix Kit (Takara Bio Inc. Shiga, Japan) was used to synthesize cDNA. qPCR was performed using SYBR Premix Ex Taq™ kit (Takara). The primer sequences used in this study were as follows: circEXOC6B, forward: 5ʹ-CCTACCTGCCTCAAGTAAGCC-3ʹ; reverse: 5ʹ-CGTCACTTGATTTTGCTTCATG-3ʹ. Actin, forward: 5ʹ-GTCCACCGCAAATGCTTCTA-3ʹ; reverse: 5ʹ-TGCTGTCACCTTCACCGTTC-3ʹ. MiR-421, forward: 5ʹ-TCAACAGACATTAATTGGGCG-3ʹ; reverse: 5ʹ-CTCAACTGGTGTCGTGGAGTC-3ʹ. U6, forward: 5ʹ-CTCGCTTCGGCAGCACAT-3ʹ; reverse: 5ʹ-AACGCTTCACGAATTTGCGT-3ʹ. RNA levels were normalized to actin and U6 for circEXOC6B and miR‐421, respectively, by using 2−ΔΔCt method.
Lentivirus Production and Cell Transfection
The circEXOC6B-shRNA1 and circEXOC6B-shRNA2 plasmids were synthesized by GenePharma (Shanghai, China). For overexpression of circEXOC6B, the circEXOC6B sequence was synthesized by GenePharma, and then sub-cloned into the lentiviral expression vector. CircEXOC6B-shRNA1, circEXOC6B-shRNA2 and lenti-circEXOC6B (circEXOC6B-OE) plasmids were infected into 293T cells for 72 h. After that, the supernatants containing the retroviral particles were collected.
A2780 cells were infected with circEXOC6B-shRNA1 or circEXOC6B-shRNA2, respectively. Twenty-four hours later, cells were incubated with puromycin (2.5 μg/mL, Thermo Fisher Scientific) to select stable circEXOC6B overexpression cells for another 48 h. SKOV3 cells were infected with circEXOC6B-OE supernatants for 24 h. Later on, cells were incubated with puromycin (2.5 μg/mL, Thermo Fisher Scientific) to select stable circEXOC6B knockdown cells for another 48 h.
MiR-421 agonist and miR-421 antagonist were obtained from GenePharma. SKOV3 cells were transfected with miR-421 agonist or miR-421 antagonist for 72 h using Lipofectamine 2000 (Thermo Fisher Scientific) according to the manufacturer’s instructions.
Cell Proliferation Assay
Cell viability was determined using Cell Counting Kit 8 (Beyotime, China). A2780 cells were infected with circEXOC6B-shRNA1 and circEXOC6B-shRNA2 for 24, 48 and 72 h, respectively. In addition, SKOV3 cells were infected with circEXOC6B-OE for 24, 48 and 72 h, respectively. After that, the culture medium was replaced with fresh DMEM containing 10% CCK8 reagent and incubated for another 2 h. The OD value was detected at 450 nm by using a microplate reader (Bio-Rad Laboratories, Hercules, CA, USA).
For colony formation assay, A2780 or SKOV3 cells (5 × 103 cell/well) were plated into 6-well plates and incubated at 37°C overnight. After that, A2780 cells were infected with circEXOC6B-shRNA2 for another 3 days at 37°C. In addition, ASKOV3 cells were infected with circEXOC6B-OE for another 3 days at 37°C. Later on, A2780 or SKOV3 cells were stained with methylene blue solution for 1 h at room temperature. Finally, cells were photographed using a fluorescence microscope (Olympus, Tokyo, Japan).
Luciferase Reporter Assay
The cDNA fragments of circEXOC6B or the RSU1 3ʹ-UTR that had a miR-421 binding site were cloned and inserted into psiCHECK™-2 Vector (Promega, Madison, WI, USA) to construct luciferase vector. After that, miR-421 agonist and above luciferase vector were co-transfected into SKOV3 cells using Lipo 2000, respectively. After 24 h of incubation, luciferase activity was detected using the Dual-Luciferase Reporter Assay System (Promega, Madison, USA) with renilla luciferase activity as the endogenous control.
Fluorescence in situ Hybridization (FISH) Analysis
The oligonucleotide probe for circEXOC6B was synthesized and obtained from Biosense (Guangzhou, China). SKOV3 cells were incubated with 10 μL of probe mixture (Biosense) in the dark overnight at 37°C. Subsequently, cell nucleus was stained with DAPI for 20 min and then observed under a fluorescence microscope (Olympus) as previously described.22
Transwell Assay
Transwell invasion assay was performed on a 24‐well transwell plate (Corning New York, NY, USA) coated with Matrigel in each chamber. After that, 100 μL serum‐free medium was added into the upper chamber. Later on, the lower compartment contained 600 μL of DMEM with 10% FBS act as a chemoattractant. After 24 h of incubation, the invaded cells at the bottom were fixed with 4% paraformaldehyde and then stained with 0.2% crystal violet. Finally, cells were photographed using a microscope (Olympus).
Flow Cytometry
SKOV3 cells were collected and washed twice with pre-cold PBS. After that, Annexin V-FITC Apoptosis Detection Kit was performed to measure cell apoptosis according to the manufacturer’s instructions. The cell apoptosis rate was determined on a flow cytometer (FACScan™, BD Biosciences, Franklin Lakes, NJ, USA).
Western Blot Assay
Total protein was extracted with ice-cold RIPA buffer (Thermo Fisher Scientific) and then quantified using BCA method (Beyotime). Proteins (30 μg per lane) were loaded onto 10% SDS-PAGE. Then, the proteins were electro-transferred onto polyvinylidene difluoride (PVDF) membrane (Thermo Fisher Scientific) and blocked in 5% skimmed milk for 1 h. Later on, the membrane was incubated in primary antibodies at 4°C overnight. Rabbit against Bcl-2 (1:1000, Abcam), cleaved caspase 3 (1:1000, Abcam), RSU1 (1:1000, Abcam), PINCH1 (1:1000, Abcam), ILK (1:1000, Abcam) and β-actin (1:1000, Abcam). After that, anti-rabbit IgG HRP-conjugate secondary antibodies (Abcam) were reacted with the membrane at room temperature for 1 h. Subsequently, the membrane was visualized using an enhanced chemiluminescence (Thermo Fisher Scientific). β-actin was acted as the internal control.
Statistical Analysis
All data were repeated in triplicate. Data are presented as the mean ± standard deviation (S.D.). All statistical analyses were performed using GraphPad Prism software (version 7.0, La Jolla, CA, USA). One-way analysis of variance (ANOVA) and Tukey’s tests were carried out for multiple group comparisons. The results were considered significant at *P < 0.05.
Results
Downregulation of circEXOC6B Promoted the Proliferation of Ovarian Cancer Cells
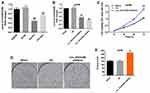
To investigate the expression of circEXOC6B in ovarian cancer cells and epithelial cells, qRT-PCR was used. As shown in Figure 1A, the level of circEXOC6B was significantly decreased in SKOV3 and OVCAR3 cells, compared with that in IOSE80 cells. Meanwhile, the level of circEXOC6B in A2780 cells was not different compared with that in IOSE80 cells (Figure 1A). In addition, the level of circEXOC6B was markedly downregulated in A2780 cells after infection with circEXOC6B-shRNA1 or circEXOC6B-shRNA2 (Figure 1B). The results indicated that circEXOC6B-shRNA2 downregulated the level of circEXOC6B more significantly than circEXOC6B-shRNA1 in A2780 cells; thus, circEXOC6B-shRNA2 was utilized in the following experiments (Figure 1B). Next, CCK-8 and colony formation assays indicated that downregulation of circEXOC6B notably promoted the proliferation of A2780 cells (Figure 1C–E). These data indicated that downregulation of circEXOC6B could promote the proliferation of ovarian cancer cells.
Overexpression of circEXOC6B Inhibited the Proliferation of Ovarian Cancer Cells
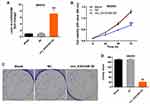
To further explore the effect of circEXOC6B on the proliferation of ovarian cancer cells, SKOV3 cells were infected with lenti-circEXOC6B. As indicated in Figure 2A, overexpression of circEXOC6B significantly upregulated the level of circEXOC6B in SKOV3 cells (Figure 2A). In addition, CCK-8 and colony formation assays indicated that the proliferation of SKOV3 cells was obviously suppressed following infection with lenti-circEXOC6B (Figure 2B–D). These data indicated that upregulation of circEXOC6B could inhibit the proliferation of ovarian cancer cells.
CircEXOC6B Functioned as a ceRNA of miR-421 in Ovarian Cancer
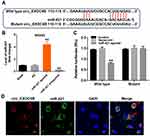
Evidence has been shown that some circRNAs can regulate gene expression through functioning as miRNA sponges.16 To investigate the miRNAs interacted with circEXOC6B, circular RNA interactome (https://circinteractome.nia.nih.gov) was performed. The data indicated that miR-421 might be a binding target of circEXOC6B (Figure 3A). In addition, the level of miR-421 was significantly increased in SKOV3 cells following transfection with miR-421 agonist, and the level of miR-421 was markedly decreased in cells following transfection with miR-421 antagonist (Figure 3B). Moreover, dual-luciferase reporter assay indicated that miR-421 agonist notably suppressed luciferase activity in SKOV3 cells at the reporter plasmid with a circEXOC6B-WT 3ʹ-UTR, while no significant inhibition was observed at the reporter plasmid with a circEXOC6B-MT 3ʹ-UTR (Figure 3C). Meanwhile, FISH assay indicated that circEXOC6B and miR-421 were partially co-localized in the cytoplasm, suggesting the direct interaction of circEXOC6B with miR-421 (Figure 3D). All these results suggested that circEXOC6B could function as a ceRNA of miR-421 in ovarian cancer.
Overexpression of circEXOC6B Inhibited the Invasion Ability of Ovarian Cancer Cells
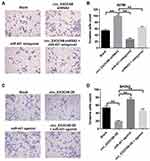
To further investigate the effect of circEXOC6B on the invasion of ovarian cancer cells, transwell invasion assay was performed. As shown in Figure 4A and B, downregulation of circEXOC6B significantly increased the invasion ability of A2870 cells, which was markedly reversed following transfection with miR-421 antagonist. In addition, overexpression of circEXOC6B notably inhibited the invasion ability of SKOV3 cells, while miR-421 agonist significantly promoted the cell invasion ability (Figure 4C and D). As expected, inhibitory effect of circEXOC6B overexpression on cell invasion was reversed by the treatment of miR-421 agonist (Figure 4C and D). These data suggested that overexpression of circEXOC6B could suppress the invasion ability of ovarian cancer cells, whereas knockdown of circEXOC6B could promote cell invasion.
Overexpression of circEXOC6B Induced the Apoptosis of Ovarian Cancer Cells
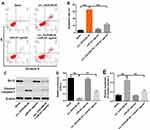
Next, to explore the effect of circEXOC6B on apoptosis of ovarian cancer cells, flow cytometry assay was applied. As shown in Figure 5A and B, overexpression of circEXOC6B significantly induced the apoptosis of SKOV3 cells, which was markedly reversed following transfection with miR-421 agonist. In addition, the level of Bcl-2 was downregulated, while the level of cleaved caspase 3 was upregulated in SKOV3 cells following infection with lenti-circEXOC6B, compared with the blank group (Figure 5C–E). Consistently, the effect of lenti-circEXOC6B on protein expressions of Bcl-2 and cleave caspase 3 in cells was notably reversed by miR-421 agonist (Figure 5C–E). These results illustrated that overexpression of circEXOC6B could induce the apoptosis of ovarian cancer cells.
CircEXOC6B Exhibited Anti-Tumor Effect Partially by Sponging miR-421 via Upregulation of RUS1
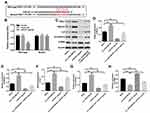
We next used TargetScan dataset (http://www.targetscan.org/vert_71/to predict the potential target of miR-421. The data indicated that RUS1 might be a potential target of miR-421 (Figure 6A). Additionally, dual-luciferase reporter assay validated that miR-421 agonist obviously inhibited the luciferase activity of psiCHECK-2-RUS1-WT, indicating that RUS1 was a binding target of miR-421 (Figure 6B). Moreover, miR-421 agonist significantly decreased the expressions of RUS1, PINCH1, ILK and E-cadherin in SKOV3 cells, but increased the expression of α-SMA (Figure 6C–H). In contrast, the expressions of RUS1, PINCH1, ILK and E-cadherin were markedly upregulated, while the expression of α-SMA was obviously downregulated in SKOV3 cells following infection with lenti-circEXOC6B. As expected, the regulated effects of lenti-circEXOC6B on these proteins were obviously abolished following transfection with miR-421 agonist in cells (Figure 6C–H). All the data suggested that CircEXOC6B exhibited anti-tumor effect were partially by sponging miR-421 via upregulation of RUS1.
Discussion
CircRNAs are widely expressed in a variety of organisms, including human cells.23 Evidence has been shown that circRNAs play an important role in the progression of human cancers.24 Meanwhile, circRNAs have been reported to exhibit some functions, such as miRNA sponges and protein sponges.25 Guan et al found that circPUM1 could promote the proliferation and invasion of ovarian cancer cells via sponging miR-615-5p and miR-6753-5p.26 Hu et al indicated that circ-ITCH could inhibit the progression of ovarian cancer via sponging miR-145.27 In this study, we found that circEXOC6B was differentially expressed between the different ovarian cell lines A2870, SKOV3 and OVCAR3, indicating that circEXOC6B expression might be related to the types of ovarian cancer. However, the potential role of circRNAs in ovarian cancer remains largely unclear.
Ning et al found that the expression of circEXOC6B was significantly decreased in epithelial ovarian cancer tissues, and low circEXOC6B expression was associated with poor prognosis in patients with ovarian cancer.17 However, the mechanism by which circEXOC6B regulates the proliferation, apoptosis and invasion in ovarian cancer cells remains unclear. In this study, we found that upregulation of circEXOC6B markedly inhibited the proliferation and invasion of SKOV3 cells. These data indicated that circEXOC6B may function as a tumor suppressor in ovarian cancer.
The circRNA-miRNA-mRNA network has been reported to involve in the mediation of tumorigenesis.28 Some circRNAs can regulate gene expression via functioning as miRNA sponges.23 For example, circGRAMD1B sponges miR-130a-3p to inhibit gastric cancer tumorigenesis.29 In the present study, luciferase reporter assay indicated that miR-421 was a potential binding target of circEXOC6B. A research showed that downregulation of miR-421 inhibited the proliferation and migration of lung cancer cells.30 Our data indicated that overexpression of miR-421 significantly increased the invasion of SKOV3 cells, suggesting that miR-421 functions as an oncogene in ovarian cancer. Meanwhile, FISH assay suggested the direct interaction of circEXOC6B with miR-421. We identified circEXOC6B as the sponge for miR-421. In addition, overexpression of circEXOC6B induced the apoptosis of SKOV3 cells, which was reversed in the presence of miR-421 agonist. These results illustrated that the circEXOC6B-miR-421 axis could play an important role in the progression of ovarian cancer.
Afterwards, TargetScan dataset was used to predict the potential target of miR-421. The data indicated that RSU1 was a direct target of miR-421. Meanwhile, luciferase reporter assays validated that miR-421 targeted RSU1 mRNA at its 3ʹ-UTR. RSU1, a tumor suppressor, which could inhibit Ras-dependent oncogenic transformation.31 RSU1 protein was identified that localized to cell-ECM adhesions.32 In addition, particularly interesting new cysteine-histidine-rich protein-1 (PINCH-1) is a cell-ECM adhesion adaptor protein.32 Evidence has been shown that RSU1 exhibited the anti-tumor effect in cancer cells via its interaction with PINCH-1.32 Moreover, the complex composed of PINCH, integrin-linked kinase (ILK) and alpha-parvin could form a stable ternary complex at cell-ECM adhesion.33 The ternary complex participated in various cellular processes, including cell spreading, migration and invasion.31 Dougherty et al indicated that RSU1 may inhibit cell migration by stabilizing the IPP adhesion complex.21 In the present study, overexpression of circEXOC6B significantly increased the expression of RSU1, PINCH-1 and ILK in SKOV3 cells, indicating that overexpression of circEXOC6B could inhibit the invasion of SKOV3 cells via upregulation of RSU1. In addition, overexpression of circEXOC6B significantly upregulated the level of epithelial marker E-cadherin, but downregulated the expression of a-SMA in SKOV3 cells indicating that lenti-circEXOC6B could inhibit the invasion ability of SKOV3 cells. Moreover, our results indicated that miR-421 exerted its oncogenic role on ovarian cancer via inhibiting RSU1. Interestingly, when treated SKOV3 cells with circEXOC6B plus miR-421 agonist, the favorable role of circEXOC6B on protein expression of RSU1 was inhibited by miR-421 overexpression. These results indicated that overexpression of circEXOC6B could inhibit the proliferation and invasion of ovarian cancer cells via regulation of miR-421/RSU1 signaling pathway.
Conclusion
In summary, we demonstrated that circEXOC6B was a tumor suppressor that inhibited the progression of ovarian cancer by sponging miR-421 and then promoting the function of RSU1. Therefore, circEXOC6B/miR-421/RSU1 axis might be a therapeutic target for the treatment of ovarian cancer.
Disclosure
The authors declare no conflicts of interest in this work.
References
1. Reid BM, Permuth JB, Sellers TA. Epidemiology of ovarian cancer: a review. (2095-3941 (Print)). Cancer Biol Med. 2017;14(1):9.
2. Bray F, Ferlay J, Soerjomataram I, et al. Global cancer statistics 2018: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J Clin. 2018;68:394–424. doi:10.3322/caac.21492.
3. Ng JS, Low JJ, Ilancheran A. Epithelial ovarian cancer. Best Pract Res Clin Obstet Gynaecol. 2012;26(3):337–345. doi:10.1016/j.bpobgyn.2011.12.005
4. Hollis RL, Gourley C, L Hollis R, Gourley C. Genetic and molecular changes in ovarian cancer. Cancer Biol Med. 2016;13(2):236–247. doi:10.20892/j.issn.2095-3941.2016.0024
5. Shi C, Yang Y, Zhang L, et al. MiR-200a-3p promoted the malignant behaviors of ovarian cancer cells through regulating PCDH9. Onco Targets Ther. 2019;12:8329–8338. doi:10.2147/OTT.S220339
6. Bristow RE, Santillan A, Salani R, et al. Intraperitoneal cisplatin and paclitaxel versus intravenous carboplatin and paclitaxel chemotherapy for Stage III ovarian cancer: a cost-effectiveness analysis. Gynecol Oncol. 2007;106(3):476–481. doi:10.1016/j.ygyno.2007.05.043
7. Zhou Y, Jin Z, Wang C. Glycogen phosphorylase B promotes ovarian cancer progression via Wnt/beta-catenin signaling and is regulated by miR-133a-3p. Biomed Pharmacother. 2019;120:109449. doi:10.1016/j.biopha.2019.109449
8. Siegel R, Ward E, Brawley O, Jemal A. Cancer statistics, 2011: the impact of eliminating socioeconomic and racial disparities on premature cancer deaths. CA Cancer J Clin. 2011;61(4):212–236. doi:10.3322/caac.20121
9. Zhang L, Zhou Q, Qiu Q, et al. CircPLEKHM3 acts as a tumor suppressor through regulation of the miR-9/BRCA1/DNAJB6/KLF4/AKT1 axis in ovarian cancer. Mol Cancer. 2019;18(1):144. doi:10.1186/s12943-019-1080-5
10. Zhang Y, Zhang XO, Chen T, et al. Circular intronic long noncoding RNAs. Mol Cell. 2013;51(6):792–806. doi:10.1016/j.molcel.2013.08.017
11. Ashwal-Fluss R, Meyer M, Pamudurti NR, et al. circRNA biogenesis competes with pre-mRNA splicing. Mol Cell. 2014;56(1):55–66. doi:10.1016/j.molcel.2014.08.019
12. Wei J, Wang J, Gao X, Qi F. Identification of differentially expressed circRNAs and a novel hsa_circ_0000144 that promote tumor growth in gastric cancer. Cancer Cell Int. 2019;19:268. doi:10.1186/s12935-019-0975-y
13. Feng Y, Yang Y, Zhao X, et al. Circular RNA circ0005276 promotes the proliferation and migration of prostate cancer cells by interacting with FUS to transcriptionally activate XIAP. Cell Death Dis. 2019;10(11):792. doi:10.1038/s41419-019-2028-9
14. Li C, He X, Zhang L, Li L, Zhao W. A pair-wise meta-analysis highlights circular RNAs as potential biomarkers for colorectal cancer. BMC Cancer. 2019;19(1):957. doi:10.1186/s12885-019-6136-9
15. Zong ZH, Du YP, Guan X, Chen S, Zhao Y. CircWHSC1 promotes ovarian cancer progression by regulating MUC1 and hTERT through sponging miR-145 and miR-1182. J Exp Clin Cancer Res. 2019;38(1):437. doi:10.1186/s13046-019-1437-z
16. Hsiao KY, Lin YC, Gupta SK, et al. Noncoding effects of circular RNA CCDC66 promote colon cancer growth and metastasis. Cancer Res. 2017;77(9):2339–2350. doi:10.1158/0008-5472.CAN-16-1883
17. Ning L, Long B, Zhang W, et al. Circular RNA profiling reveals circEXOC6B and circN4BP2L2 as novel prognostic biomarkers in epithelial ovarian cancer. Int J Oncol. 2018;53(6):2637–2646. doi:10.3892/ijo.2018.4566
18. Chen J, Wu L, Sun Y, et al. Mir-421 in plasma as a potential diagnostic biomarker for precancerous gastric lesions and early gastric cancer. Peer J. 2019;7:e7002. doi:10.7717/peerj.7002.
19. Xue L, Yang D. MiR-421 inhibited proliferation and metastasis of colorectal cancer by targeting MTA1. (1107-0625 (Print)). J BUON. 2018;23:1633–1639.
20. Zhu J, Han S. Lidocaine inhibits cervical cancer cell proliferation and induces cell apoptosis by modulating the lncRNA-MEG3/miR-421/BTG1 pathway. (1943-8141 (Print)). Am J Transl Res. 2019;11(9):5404.
21. Dougherty GW, Jose C, Gimona M, Cutler ML. The Rsu-1-PINCH1-ILK complex is regulated by Ras activation in tumor cells. Eur J Cell Biol. 2008;87(8–9):721–734. doi:10.1016/j.ejcb.2008.02.011
22. Liang M, Liu Z, Lin H, et al. High-throughput sequencing reveals circular RNA hsa_circ_0000592 as a novel player in the carcinogenesis of gastric carcinoma. Biosci Rep. 2019;39(6). doi:10.1042/BSR20181900.
23. Memczak S, Jens M, Elefsinioti A, et al. Circular RNAs are a large class of animal RNAs with regulatory potency. Nature. 2013;495(7441):333–338. doi:10.1038/nature11928
24. Yang R, Xing L, Zheng X, et al. The circRNA circAGFG1 acts as a sponge of miR-195-5p to promote triple-negative breast cancer progression through regulating CCNE1 expression. Mol Cancer. 2019;18(1):4. doi:10.1186/s12943-018-0933-7
25. Liu J, Zhao K, Huang N, Zhang N. Circular RNAs and human glioma. Cancer Biol Med. 2019;16(1):11–23. doi:10.20892/j.issn.2095-3941.2018.0425
26. Guan X, Zong ZH, Liu Y, et al. circPUM1 promotes tumorigenesis and progression of ovarian cancer by sponging miR-615-5p and miR-6753-5p. Mol Ther Nucleic Acids. 2019;18:882–892. doi:10.1016/j.omtn.2019.09.032
27. Hu J, Wang L, Chen J, et al. The circular RNA circ-ITCH suppresses ovarian carcinoma progression through targeting miR-145/RASA1 signaling. Biochem Biophys Res Commun. 2018;505(1):222–228.
28. Pan G, Hu T, Chen X, Zhang C. Upregulation of circMMP9 promotes osteosarcoma progression via targeting miR-1265/CHI3L1 axis. Cancer Manag Res. 2019;11:9225–9231. doi:10.2147/CMAR.S226264
29. Dai X, Guo X, Liu J, et al. Circular RNA circGRAMD1B inhibits gastric cancer progression by sponging miR-130a-3p and regulating PTEN and p21 expression. Aging (Albany NY). 2019;11(21):9689–9708. doi:10.18632/aging.102414
30. Li Y, Cui X, Li Y, Zhang T, Li S. Upregulated expression of miR-421 is associated with poor prognosis in non-small-cell lung cancer. Cancer Manag Res. 2018;10:2627–2633. doi:10.2147/CMAR.S167432
31. Zacharia LC, Stylianopoulos T, Gkretsi V. Ras suppressor-1 (RSU-1) in cancer cell metastasis: friend or foe? Crit Rev Oncog. 2017;22(3–4):249–253. doi:10.1615/CritRevOncog.2018024231
32. Phuah NH, Azmi MN, Awang K, Nagoor NH. Suppression of microRNA-629 enhances sensitivity of cervical cancer cells to 1’S-1ʹ-acetoxychavicol acetate via regulating RSU1. Onco Targets Ther. 2017;10:1695–1705. doi:10.2147/OTT.S117492
33. Legate KR, Montanez E, Kudlacek O, Fassler R. ILK, PINCH and parvin: the tIPP of integrin signalling. Nat Rev Mol Cell Biol. 2006;7(1):20–31. doi:10.1038/nrm1789
 © 2020 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.
© 2020 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.