Back to Journals » OncoTargets and Therapy » Volume 13
CircDDX42 Accelerates the Development of Pancreatic Cancer via miR-613/ID4/PI3K/AKT Axis
Received 30 September 2019
Accepted for publication 15 January 2020
Published 28 October 2020 Volume 2020:13 Pages 10945—10957
DOI https://doi.org/10.2147/OTT.S233000
Checked for plagiarism Yes
Review by Single anonymous peer review
Peer reviewer comments 2
Editor who approved publication: Dr Federico Perche
Zhen Yan,1,2,* Heliang Yin,1,2,* Guoying Lin1,2
1Department of General Surgery, The First Hospital of Qiqihar, Qiqihar 161005, People’s Republic of China; 2Department of General Surgery, Qiqihar Hospital Affiliated to Southern Medical University, Qiqihar 161005, People’s Republic of China
*These authors contributed equally to this work
Correspondence: Heliang Yin; Guoying Lin
Department of General Surgery, The First Hospital of Qiqihar, Qiqihar 161005, People’s Republic of China
Tel +86 452-2459787
Email [email protected]; [email protected]
Background: Pancreatic cancer (PC) is one of the fatal cancers globally. CircDEAD-box helicase 42 (circDDX42) has been reported to play an oncogenic role in many cancers. The purpose of our study was to explore the relationship between circDDX42 and PC development and the potential mechanism by which circDDX42 modulating the progression of PC.
Methods: The enrichment of circDDX42, miR-613 and inhibitor of DNA binding 4 (ID4) was determined by quantitative real-time polymerase chain reaction (qRT-PCR) in PC tissues and cells. The proliferation, apoptosis and metastasis of PC cells were examined by 3-(4,5-Dimethylthiazol-2-yl)-2,5-diphenyltetrazolium bromide (MTT), Western blot, flow cytometry and transwell migration and invasion assays, respectively. The binding sites between miR-613 and circDDX42 or ID4 were predicted by Starbase bioinformatic software, and dual-luciferase reporter assay was conducted to verify the combination between miR-613 and circDDX42 or ID4. Western blot was carried out to detect the abundance of ID4, p-phosphatidylinositol 3-kinase (p-PI3K), PI3K, p-AKT serine/threonine kinase (p-AKT) and AKT in PC cells. The in vivo role of circDDX42 was verified through using murine xenograft model.
Results: The level of circDDX42 was enhanced in PC tissues and cells compared with that in matching normal tissues and HPDE cells. CircDDX42 promoted the proliferation and metastasis and suppressed the apoptosis of PC cells. CircDDX42 could sponge miR-613, and miR-613 was negatively regulated by circDDX42 in PC cells. MiR-613 suppressed the progression of PC. ID4 was a direct target of miR-613. ID4 was inversely modulated by miR-613 and positively regulated by circDDX42 in PC cells. ID4 played an oncogenic role in the tumorigenesis of PC. CircDDX42/miR-613/ID4 axis regulated the activation of PI3K/AKT pathway in PC cells. ID4 facilitated the progression of PC via activating PI3K/AKT signal pathway. CircDDX42 promoted the tumor growth of PC in vivo.
Conclusion: CircDDX42 accelerated the proliferation and metastasis while impeded the apoptosis of PC cells via circDDX42/miR-613/ID4/PI3K/AKT axis. This axis might be a promising target for PC therapy.
Keywords: pancreatic cancer, circDDX42, miR-613, ID4, proliferation, apoptosis, metastasis, PI3K-AKT signal pathway
Introduction
There were 458,918 new cases and 432,242 deaths of pancreatic cancer (PC) globally in 2018.1 Although the therapeutic methods have been innovated, the 5-year survival rate of patients with advanced-stage PC remains dismal due to tumor recurrence and metastasis.2 Therefore, it is pivotal for us to better understand the molecular mechanism behind PC progression.
Circular RNAs (circRNAs) are a class of non-coding RNAs (ncRNAs) and are marked by the covalently closed loop structure.3 CircRNAs are involved in the growth, metastasis of multiple cancers.4–9 Currently, circRNAs have been reported to serve as RNA sponges for microRNAs (miRNAs), and the circRNAs/miRNAs axis could modulate the initiation and progression of many cancers.10 Hence, circRNAs might function as diagnostic biomarkers of a variety of cancers.11,12 CircDEAD-box helicase 42 (circDDX42; CircBase ID: hsa_circ_0007534) has been reported to play an oncogenic role in multiple cancers, including breast cancer (BC), glioma, osteosarcoma and pancreatic ductal adenocarcinoma (PDAC).13–16 Nevertheless, the role of circDDX42 in the development of PC remains poorly understood.
MiRNAs are another class of ncRNAs, and miRNAs could restrain the translation of messenger RNAs (mRNAs) by binding to their 3ʹ untranslated region (UTR).17–19 Accumulating articles claimed that the low expression of miRNAs and the up-regulation of target proteins were related to the occurrence and progression of many cancers.20–24 MiR-613 played a suppressive role in many cancers, including gastric cancer (GC), liver cancer and PC.25–28 However, the precise mechanism by which miR-613 inhibiting the progression of PC is not fully addressed.
Inhibitor of DNA binding 4 (ID4) is a member of ID family. ID4 was involved in individual development and tumorigenesis.29 Cheng et al found that SNHG7 accelerated the growth of PC cells via miR-342-3p/ID4 axis.30 Zhang et al demonstrated that ID4 facilitated the proliferation of hepatocellular carcinoma (HCC) cells. However, the function of ID4 in PC remains to be determined.
In this study, we first determined the expression of circDDX42 in PC tissues and cells. Knockdown experiments were performed to evaluate the function of circDDX42 on the proliferation, apoptosis and metastasis of PC cells. In addition, we explored the downstream genes and signal pathway of circDDX42 to further illustrate the mechanism by which circDDX42 promoting the progression of PC.
Materials and Methods
Tissue Specimens
We collected forty PC tissues and paired non-tumor tissues from PC patients who had received resection in The First Hospital of Qiqihar. Written informed consents were acquired from patients who participated in the present study. All protocols in this study had gotten the permission from the Ethics Committee of The First Hospital of Qiqihar.
Cell Culture
PC cell lines SW1990 and CFPAC-1 and normal pancreatic duct epithelial cell line HPDE were acquired from BeNa Culture Collection (Beijing, China). All cells were cultured in Roswell Park Memorial Institute-1640 (RPMI-1640) medium (Gibco, Carlsbad, CA, USA) added with 10% fetal bovine serum (FBS; Gibco), penicillin (100 units/mL) and streptomycin (100 μg/mL) at 37°C atmosphere with 5% CO2.
Quantitative Real-Time Polymerase Chain Reaction (qRT-PCR)
RNA sample was isolated with TRIzol (Invitrogen, Carlsbad, CA, USA) followed by removal of DNA with DNase. Complementary DNA (cDNA) was synthesized using M-MLV reverse transcriptase kit (Invitrogen). Glyceraldehyde-3-phosphate dehydrogenase (GAPDH) or U6 was considered as the internal control. The 2−ΔΔCt method was used for quantification of the expression of circDDX42, miR-613 and ID4.31 The primers used in this study are listed in Table 1.
 |
Table 1 Primers Used in qRT-PCR |
Cell Transfection
PC cells were seeded into 6-well plates, and transfection was performed through using Lipofectamine 2000 (Invitrogen) when the cells were grown to 70% confluence. Small interference RNA negative control (si-NC), small interference RNA against circDDX42 (si-circDDX42), small hairpin RNA negative control (sh-NC), shRNA targeting circDDX42 (sh-circDDX42), small interference RNA against ID4 (si-ID4), pcDNA negative control (pcDNA-NC) and ID4 overexpression plasmid (pcDNA-ID4) were obtained from Genepharma (Shanghai, China). MiRNA negative control (miR-NC), miR-613, anti-miR-NC and anti-miR-613 were purchased from Ribobio (Guangzhou, China).
3-(4,5-Dimethylthiazol-2-yl)-2,5-Diphenyltetrazolium Bromide (MTT) Assay
After transfection for 0 hrs, 24 hrs, 48 hrs and 72 hrs, 20 µL MTT (Invitrogen) was added to the culture medium of the wells of 96-well plates, and cells were incubated for 4 h. Dimethyl sulfoxide (DMSO; Sigma, St. Louis, MO, USA) was added to the wells to dissolve the formazan after discarding the supernatant. The optical density (OD) value at 490 nm was measured by a microplate reader.
Western Blot Assay
Proteins were extracted using cell lysis solution. The concentration of proteins was measured by BCA method. Equal amounts of proteins were separated by sodium dodecyl sulfate polyacrylamide gel electrophoresis (SDS-PAGE) and transferred on polyvinylidene fluoride (PVDF) membranes (Millipore, Billerica, MA, USA). After blocking with 5% non-fat skim milk for 1 h, the membranes were incubated with specific primary antibody targeting Ki-67 (ab16667, Abcam, Cambridge, MA, USA), PCNA (ab92552, Abcam), ID4 (ab49261, Abcam), AKT serine/threonine kinase (AKT) (ab8805, Abcam), p-AKT (ab38449, Abcam), phosphatidylinositol 3-kinase (PI3K; ab180967, Abcam), p-PI3K (ab182651, Abcam) or GAPDH (ab181602, Abcam) at 4°C for 12 h followed by the incubation with the secondary antibody (ab205718, Abcam) for 1 h at room temperature. The immunoreactive protein signal was visualized using the enhanced chemiluminescent (ECL) system (Beyotime).
Cell Apoptosis Analysis
Cell apoptosis was measured by using Annexin V-fluorescein isothiocyanate (FITC) Apoptosis Detection Kit (Invitrogen). PC cells were harvested after transfection for 72 h through centrifuging at 800 ×g, and the above cells were mixed with Annexin V-FITC and propidine iodide (PI; Solarbio, Beijing, China) simultaneously at room temperature in the dark for 10 min. The apoptotic PC cells were distinguished from necrotic or normal PC cells through the analysis of flow cytometer (BD Biosciences, San Jose, CA, USA).
Transwell Migration and Invasion Assays
Transwell chambers were used in transwell migration assay. PC cells suspended in medium without serum were added to the upper chambers. 10% FBS served as the chemotactic factor, and culture medium supplemented with 10% FBS was added to the lower chambers. Cotton swab was used to remove the cells remained in the upper surface of polycarbonate membrane, and the migrated PC cells were stained and counted using a microscope.
For detection of the invasion capacity of PC cells, matrigel (BD Biosciences) was added to the upper chambers to pre-coat the polycarbonate membrane before seeding PC cells. The other protocol of transwell invasion assay was the same as above.
Dual-Luciferase Reporter Assay
The cDNA sequences of circDDX42 (wild-type or mutant type) were embedded into the pmirGLO luciferase reporter vector (Promega, Madison, WI, USA). The luciferase reporter vectors were co-transfected with miR-613 or miR-NC into SW1990 and CFPAC-1 cells. Luciferase activity was detected through dual-luciferase reporter assay system (Promega). The validation of the binding between miR-613 and ID4 was conducted following the same protocol.
In vivo Xenograft Assay
This study was approved by the Animal Ethics Committee of the The First Hospital of Qiqihar. Nude mice from the Chinese Academy of Medical Sciences (Beijing, China) were subcutaneously injected SW1990 cells stably expressing sh-circDDX42. A vernier caliper was used to measure the volume of tumors from sh-NC group and sh-circDDX42 group every week. The weight of tumors was detected by using an analytical balance. QRT-PCR and Western blot were performed to detect relevant expression of RNAs and proteins.
Statistical Analysis
All data were analyzed using GraphPad Prism7 software and were presented as mean ± standard deviation (SD). P value was calculated using Student’s t-test and one-way analysis of variance (ANOVA) followed by Turkey’s test. Spearman correlation coefficient was used to evaluate the relationship between the expression of miR-613 and ID4 or circDDX42 in PC tissues. P<0.05 indicated as statistically significant.
Results
CircDDX42 Promotes the Proliferation and Metastasis While Suppresses the Apoptosis of PC Cells
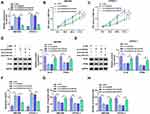
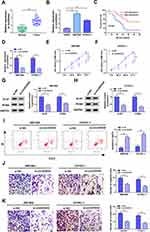
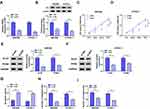
QRT-PCR was performed to explore the expression pattern of circDDX42 in PC tissues and corresponding normal tissues. As indicated in Figure 1A, circDDX42 was abnormally up-regulated in PC tissues compared with that in adjacent normal tissues. Meanwhile, we found that the level of circDDX42 was also elevated in PC cells compared with that in normal pancreatic duct epithelial cells HPDE (Figure 1B). PC patients with high expression of circDDX42 possessed a low survival rate (Figure 1C).
To investigate the biological role of circDDX42 in PC cells, MTT, Western blot, flow cytometry and transwell migration and invasion assays were conducted to measure the proliferation, apoptosis and metastasis of PC cells transfected with si-NC or si-circDDX42. The level of circDDX42 was reduced in PC cells transfected with si-circDDX42 (Figure 1D). As showed in Figure 1E and F, the knockdown of circDDX42 prominently restrained the proliferation of PC cells. Besides, the abundance of proliferation-related proteins (Ki-67 and PCNA) was down-regulated by the transfection of si-circDDX42, indicating that circDDX42 promoted the proliferation of PC cells (Figure 1G and H). As mentioned in Figure 1I, the interference of circDDX42 accelerated the apoptosis of SW1990 and CFPAC-1 cells. Additionally, the depletion of circDDX42 inhibited the migration and invasion of PC cells (Figure 1J and K). Taken together, circDDX42 played an oncogenic role in PC cells.
MiR-613 Is a Direct Target of circDDX42
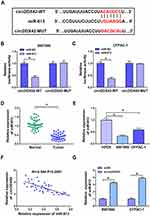
MiR-613 was predicted as a target of circDDX42 by Starbase online software, and the binding sites between miR-613 and circDDX42 are displayed in Figure 2A. The luciferase reporter vector containing the wild-type or mutant type binding sites of circDDX42, namelycircDDX42-WT or circDDX42-MUT, was co-transfected with miR-NC or miR-613 into SW1990 and CFPAC-1 cells. As shown in Figure 2B and C, luciferase activity was significantly declined with the accumulation of miR-613 in the circDDX42-WT group compared with that in the circDDX42-MUT group, indicating that miR-613 was a direct target of circDDX42 in PC cells.
We measured the enrichment of miR-613 in PC tissues and cells. As indicated in Figure 2D and E, the level of miR-613 was declined in PC tissues and cells compared with that in matching normal tissues and HPDE cells. Subsequently, we found an inverse correlation between the expression of miR-613 and the abundance of circDDX42 in PC tissues (Figure 2F). Meanwhile, the abundance of miR-613 was negatively regulated by circDDX42 in PC cells (Figure 2G). Collectively, miR-613 was a direct target of circDDX42 and was inversely modulated by circDDX42 in PC cells.
The Depletion of miR-613 Alleviates the Suppressive Effects of circDDX42 Intervention on the Proliferation and Metastasis and the Promoting Impact on the Apoptosis of PC Cells
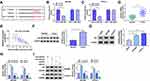
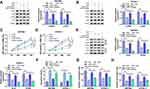
We wondered whether miR-613 was involved in circDDX42-mediated proliferation, apoptosis and metastasis of PC cells. SW1990 and CFPAC-1 cells were transfected with si-NC, si-circDDX42, si-circDDX42 + anti-miR-NC orsi-circDDX42 + anti-miR-613, respectively. As indicated in Figure 3A, the abundance of miR-613 was increased with the knockdown of circDDX42, and the inhibition of miR-613 reversed the effect of circDDX42 interference on the abundance of miR-613 in SW1990 and CFPAC-1 cells. MTT assay showed that the proliferation of PC cells was inhibited by the transfection of si-circDDX42, and it was recovered by the co-transfection of si-circDDX42 and anti-miR-613 (Figure 3B and C). The depletion of circDDX42 reduced the abundance of Ki-67 and PCNA in PC cells, and the addition of anti-miR-613 recovered the abundance of the two proteins, indicating that the inhibition of miR-613 attenuated the inhibitory effect of circDDX42 interference on the growth of PC cells (Figure 3D and E).
Besides, the depletion of miR-613 reversed the promoting effect of circDDX42 interference on the apoptosis ofSW1990 and CFPAC-1 cells (Figure 3F). Transwell migration and invasion assays indicated that the transfection of anti-miR-613 abolished the suppressive impact of circDDX42 inhibition on the metastasis of PC cells (Figure 3G and H). Collectively, circDDX42 accelerated the proliferation and metastasis while inhibited the apoptosis of PC cells through sponging miR-613.
ID4 Is a Functional Target of miR-613
To illustrate the mechanism by which miR-613 inhibiting the progression of PC, we aimed to find the downstream gene of miR-613. The binding sites between miR-613 and ID4 were predicted by Starbase software (Figure 4A). The wild-type or mutant type binding sites of ID4 3ʹ UTR were inserted into luciferase reporter vector, generating ID4 3ʹ UTR-WT or ID4 3ʹ UTR-MUT plasmid, respectively. SW1990 and CFPAC-1 cells were co-transfected with ID4 3ʹ UTR-WT or ID4 3ʹ UTR-MUT and miR-NC or miR-613. The luciferase activity was significantly decreased in PC cells co-transfected with miR-613 and ID4 3ʹ UTR-WT, whereas it remained unchanged in the two PC cells co-transfected with miR-613 and ID4 3ʹ UTR-MUT, demonstrating that miR-613 directly bound to ID4 in SW1990 and CFPAC-1 cells (Figure 4B and C).
QRT-PCR was carried out to measure the abundance of ID4 mRNA in PC tissues and adjacent normal tissues. As showed in Figure 4D, the enrichment of ID4mRNA was higher in PC tissues than that in corresponding normal tissues. The expression of miR-613 was negatively correlated with the abundance of ID4 in PC tissues (Figure 4E). The abundance of ID4 protein was elevated in PC tissues and cells compared with that in paired normal tissues and HPDE cells (Figure 4F and G).
To clarify the modulatory relationship between ID4 and miR-613 in PC cells, the abundance of ID4 mRNA and protein was measured in SW1990 and CFPAC-1 cells transfected with anti-miR-NC, anti-miR-613, miR-NC or miR-613. As indicated in Figure 4H and I, the enrichment of ID4 was increased by the inhibition of miR-613, and the overexpression of miR-613 reduced the mRNA and protein expression of ID4. These findings suggested that ID4 was a direct target of miR-613 in PC cells, and there was a negative correlation between the expression of miR-613 and the abundance of ID4 in PC tissues.
The Abundance of ID4 Is Regulated by circDDX42/miR-613 Axis in PC Cells
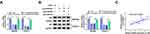
As mentioned above, there was an inverse correlation between the enrichment of miR-613 and the expression of circDDX42 or ID4. To further confirm the regulatory relationship among circDDX42, miR-613 and ID4, we conducted the following experiments. SW1990 and CFPAC-1 cells were transfected with si-NC, si-circDDX42, si-circDDX42 + anti-miR-NC or si-circDDX42 + anti-miR-613, respectively. As showed in Figure 5A and B, the abundance of ID4 mRNA and protein was down-regulated with the interference of circDDX42, and it was enhanced with the addition of anti-miR-613 in PC cells. As expected, the abundance of circDDX42 was positively correlated with the level of ID4 in PC tissues (Figure 5C).
ID4 Accelerates the Proliferation and Metastasis While Impedes the Apoptosis of PC Cells
To investigate the function of ID4 on the proliferation, apoptosis and metastasis of PC cells, SW1990 and CFPAC-1 cells transfected with si-NC or si-ID4 were used for the following experiments. We first assessed the knockdown efficiency of si-ID4 in the two PC cells, and the abundance of ID4 mRNA and protein was notably decreased by the transfection of si-ID4 in PC cells (Figure 6A and B). As displayed in Figure 6C and D, the depletion of ID4 impeded the proliferation of SW1990 and CFPAC-1 cells. Western blot assay revealed that the levels of proliferation-related proteins (Ki-67 and PCNA) were declined in si-ID4 group, suggesting that ID4 promoted the proliferation of PC cells (Figure 6E and F). Cell apoptosis was dramatically increased in SW1990 and CFPAC-1 cells transfected with si-ID4 (Figure 6G). As indicated in Figure 6H and I, the silencing of ID4 restrained the metastasis of PC cells. Taken together, ID4 accelerated the proliferation and metastasis while suppressed the apoptosis of PC cells.
CircDDX42/miR-613/ID4 Axis Regulates the Activation of PI3K/AKT Pathway
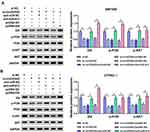
To explore the downstream pathway by which ID4 promoting the proliferation and metastasis and impeding the apoptosis of PC cells, SW1990 and CFPAC-1 cells were transfected with si-NC, si-circDDX42, si-circDDX42 + anti-miR-NC, si-circDDX42 + anti-miR-613, si-circDDX42 + pcDNA-NC or si-circDDX42 + pcDNA-ID4. The phosphorylation of PI3K and AKT was restrained with the interference of circDDX42, and it was recovered with the co-transfection of si-circDDX42 and anti-miR-613 or pcDNA-ID4 in PC cells (Figure 7A and B). These findings indicated that PI3K/AKT pathway could be modulated by circDDX42/miR-613/ID4 axis in PC cells.
ID4 Promotes the Activation of PI3K/AKT Signaling
To clarify the regulatory relationship between ID4 and PI3K/AKT pathway in PC cells, the inhibiting agent (LY294002) and activating agent (740Y-P) of PI3K/AKT pathway were used in the following experiments. As mentioned in Figure 8A and B, ID4 intervention reduced the phosphorylation of PI3K and AKT, while the addition of 740Y-P recovered the phosphorylation of PI3K and AKT. Besides, the addition of LY294002 down-regulated the levels of p-PI3K and p-AKT in PC cells compared with that in si-ID4 and 740Y-P group. Functional experiments showed that the stimulation of 740Y-P reversed the inhibitory effect of ID4 interference on the proliferation of PC cells (Figure 8C–E), while the stimulation of LY294002 showed opposite results to 740Y-P. The apoptosis was triggered in si-ID4 group, and the addition of 740Y-P declined the apoptotic rate of PC cells (Figure 8F). The introduction of LY294002 showed reverse results on the apoptosis of PC cells relative to 740Y-P + si-ID4 group. As exhibited in Figure 8G and H, the suppressive influence of ID4 intervention on the migration and invasion of PC cells was overturned by the addition of 740Y-P, and the metastasis was restrained in si-ID4 + LY294002 group than that in si-ID4 + 740Y-P group. These results suggested that ID4 promoted the proliferation and metastasis while blocked the apoptosis of PC cells via PI3K/AKT pathway.
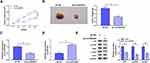
CircDDX42 Silencing Inhibits the PC Tumor Growth via miR-613/ID4/PI3K/AKT Axis in vivo
To explore whether circDDX42 exerted the same role in vivo, we used a murine xenograft model. Tumors were lesser in sh-circDDX42 group than that in the sh-NC group (Figure 9A and B). The levels of circDDX42, ID4, p-PI3K and p-AKT were decreased in tumors from sh-circDDX42 group in comparison with that in sh-NC group (Figure 9C and E). As shown in Figure 9D, a significant up-regulation of miR-613 was observed in sh-circDDX42 group. Taken together, circDDX42 accelerated the PC tumor growth via miR-613/ID4/PI3K/AKT signaling.
Discussion
CircRNAs could serve as miRNA sponges to exert their diverse functions.6 CircRNA/miRNA axis was involved in the occurrence and development of multiple cancers.7 For instance, Wang et al proved that the expression of circRNA-000911 was notably decreased in BC, and circRNA-000911 suppressed the progression of BC via sponging miR-449a.32 In this study, we found that the abundance of circDDX42 was elevated in PC tissues and cells compared with that in adjacent normal tissues and normal pancreatic duct epithelial cells HPDE. Besides, the expression of circDDX42 was negatively correlated with the prognosis of PC patients. Loss-of-function experiments indicated that circDDX42 promoted the proliferation and metastasis while inhibited the apoptosis of PC cells.
MiR-613 was predicted as a target of circDDX42 by Starbase online software, and the combination between miR-613 and circDDX42 was validated by dual-luciferase reporter assay. To clarify the role of miR-613 in PC, we first measured the expression of miR-613 in PC tissues and cells. The level of miR-613 was lower in PC tissues and cells than that in paired normal tissues and HPDE cells. The statistical analysis revealed that there was a negative correlation between the expression of miR-613 and circDDX42 in PC tissues. MiR-613 was identified as a tumor suppressor gene in many cancers. Xue et al found that the overexpression of miR-613 enhanced the sensitivity of GC cells to cisplatin via SOX9.25 Cai et al claimed that lncRNA HOTAIR promoted the progression of PC through up-regulating the expression of Notch3 by sponging miR-613.28 Consistent with the above findings, we found that the depletion of miR-613 alleviated the suppressive effects of circDDX42 interference on the proliferation and metastasis and the promoting impact on the apoptosis of PC cells, indicating that miR-613 functioned as the downstream gene of circDDX42 to suppress the growth and motility and facilitate the apoptosis of PC cells.
To illustrate the underlying molecular mechanism by which circDDX42/miR-613 regulating the progression of PC, we aimed to find the target of miR-613. ID4 was predicted to be a target of miR-613, and dual-luciferase reporter assay confirmed the combination between miR-613 and ID4. ID4 has been reported to be an oncogene in PC and HCC.30,33 Consistent with the above results, the abundance of ID4 was elevated in PC tissues and cells, and it promoted the proliferation and metastasis while impeded the apoptosis of PC cells.
PI3K-AKT pathway is involved in cell survival, growth and motility. This signal pathway was frequently activated in PC.34 The inhibition of miR-613 or the overexpression of ID4 could attenuate the inhibitory impact of circDDX42 silencing on the activation of PI3K-AKT pathway. We further found that ID4 accelerated the progression of PC via activating PI3K/AKT pathway. The in vivo assay also confirmed the oncogenic role of circDDX42. Nevertheless, the mechanism by which ID4 regulating the activation of PI3K/AKT pathway remains elusive. More efforts are needed to investigate the regulatory relationship between ID4 and PI3K/AKT signaling.
In summary, this study elucidated the biological role of circDDX42 in PC cells. CircDDX42 promoted the proliferation and metastasis while restrained the apoptosis of PC cells via miR-613/ID4/PI3K/AKT axis. The circDDX42/miR-613/ID4/PI3K/AKT axis might provide new insight in understanding the pathogenesis of PC.
Disclosure
The authors declare that they have no financial conflicts of interest in this work.
References
1. Bray F, Ferlay J, Soerjomataram I, et al. Global cancer statistics 2018: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J Clin. 2018;68(6):394–424. doi:10.3322/caac.21492
2. Coveler AL, Herman JM, Simeone DM, et al. Localized pancreatic cancer: multidisciplinary management. Am Soc Clin Oncol Educ Book. 2016;35:e217–e226. doi:10.14694/EDBK_160827
3. Ebbesen KK, Kjems J, Hansen TB. Circular RNAs: identification, biogenesis and function. Biochim Biophys Acta. 2016;1859(1):163–168. doi:10.1016/j.bbagrm.2015.07.007
4. Qian L, Yu S, Chen Z, et al. The emerging role of circRNAs and their clinical significance in human cancers. Biochim Biophys Acta Rev Cancer. 2018;1870(2):247–260. doi:10.1016/j.bbcan.2018.06.002
5. Chen LL, Yang L. Regulation of circRNA biogenesis. RNA Biol. 2015;12(4):381–388. doi:10.1080/15476286.2015.1020271
6. Fu L, Jiang Z, Li T, et al. Circular RNAs in hepatocellular carcinoma: functions and implications. Cancer Med. 2018;7:3101–3109. doi:10.1002/cam4.2018.7.issue-7
7. Meng S, Zhou H, Feng Z, et al. CircRNA: functions and properties of a novel potential biomarker for cancer. Mol Cancer. 2017;16(1):94. doi:10.1186/s12943-017-0663-2
8. Zhang HD, Jiang LH, Sun DW, Hou JC, Ji ZL. CircRNA: a novel type of biomarker for cancer. Breast Cancer. 2018;25(1):1–7. doi:10.1007/s12282-017-0793-9
9. Kristensen LS, Hansen TB, Veno MT, Kjems J. Circular RNAs in cancer: opportunities and challenges in the field. Oncogene. 2018;37(5):555–565. doi:10.1038/onc.2017.361
10. Chan JJ, Tay Y. Noncoding RNA:RNA regulatory networks in cancer. Int J Mol Sci. 2018;19(5):1310. doi:10.3390/ijms19051310
11. Li P, Chen S, Chen H, et al. Using circular RNA as a novel type of biomarker in the screening of gastric cancer. Clin Chim Acta. 2015;444:132–136. doi:10.1016/j.cca.2015.02.018
12. Gao JL, Chen G, He HQ, et al. CircRNA as a new field in human disease research. Zhongguo Zhong Yao Za Zhi. 2018;43(3):457–462. doi:10.19540/j.cnki.cjcmm.20171106.012
13. Song L, Xiao Y. Downregulation of hsa_circ_0007534 suppresses breast cancer cell proliferation and invasion by targeting miR-593/MUC19 signal pathway. Biochem Biophys Res Commun. 2018;503(4):2603–2610. doi:10.1016/j.bbrc.2018.08.007
14. Li GF, Li L, Yao ZQ, et al. Hsa_circ_0007534/miR-761/ZIC5 regulatory loop modulates the proliferation and migration of glioma cells. Biochem Biophys Res Commun. 2018;499(4):765–771. doi:10.1016/j.bbrc.2018.03.219
15. Li B, Li X. Overexpression of hsa_circ_0007534 predicts unfavorable prognosis for osteosarcoma and regulates cell growth and apoptosis by affecting AKT/GSK-3beta signaling pathway. Biomed Pharmacother. 2018;107:860–866. doi:10.1016/j.biopha.2018.08.086
16. Hao L, Rong W, Bai L, et al. Upregulated circular RNA circ_0007534 indicates an unfavorable prognosis in pancreatic ductal adenocarcinoma and regulates cell proliferation, apoptosis, and invasion by sponging miR-625 and miR-892b. J Cell Biochem. 2019;120(3):3780–3789. doi:10.1002/jcb.v120.3
17. Li Z, Lei H, Luo M, et al. DNA methylation downregulated mir-10b acts as a tumor suppressor in gastric cancer. Gastric Cancer. 2015;18(1):43–54. doi:10.1007/s10120-014-0340-8
18. Xiong X, Ren HZ, Li MH, et al. Down-regulated miRNA-214 induces a cell cycle G1 arrest in gastric cancer cells by up-regulating the PTEN protein. Pathol Oncol Res. 2011;17(4):931–937. doi:10.1007/s12253-011-9406-7
19. He XP, Shao Y, Li XL, et al. Downregulation of miR-101 in gastric cancer correlates with cyclooxygenase-2 overexpression and tumor growth. FEBS J. 2012;279(22):4201–4212. doi:10.1111/febs.12013
20. Zhang BG, Li JF, Yu BQ, et al. microRNA-21 promotes tumor proliferation and invasion in gastric cancer by targeting PTEN. Oncol Rep. 2012;27(4):1019–1026. doi:10.3892/or.2012.1645
21. Tsai MM, Huang HW, Wang CS, et al. MicroRNA-26b inhibits tumor metastasis by targeting the KPNA2/c-jun pathway in human gastric cancer. Oncotarget. 2016;7(26):39511–39526. doi:10.18632/oncotarget.8629
22. Fan L, Wu Q, Xing X, et al. MicroRNA-145 targets vascular endothelial growth factor and inhibits invasion and metastasis of osteosarcoma cells. Acta Biochim Biophys Sin (Shanghai). 2012;44(5):407–414. doi:10.1093/abbs/gms019
23. Mao JH, Zhou RP, Peng AF, et al. microRNA-195 suppresses osteosarcoma cell invasion and migration in vitro by targeting FASN. Oncol Lett. 2012;4(5):1125–1129. doi:10.3892/ol.2012.863
24. Ge H, Li B, Hu WX, et al. MicroRNA-148b is down-regulated in non-small cell lung cancer and associated with poor survival. Int J Clin Exp Pathol. 2015;8(1):800–805.
25. Xue M, Li G, Sun P, et al. MicroRNA-613 induces the sensitivity of gastric cancer cells to cisplatin through targeting SOX9 expression. Am J Transl Res. 2019;11(2):885–894.
26. Li B, Liu D, Yang P, et al. miR-613 inhibits liver cancer stem cell expansion by regulating SOX9 pathway. Gene. 2019;707:78–85. doi:10.1016/j.gene.2019.05.015
27. Liu H, Chen K, Wang L, et al. miR-613 inhibits Warburg effect in gastric cancer by targeting PFKFB2. Biochem Biophys Res Commun. 2019;515(1):37–43. doi:10.1016/j.bbrc.2019.05.001
28. Cai H, Yao J, An Y, et al. LncRNA HOTAIR acts a competing endogenous RNA to control the expression of notch3 via sponging miR-613 in pancreatic cancer. Oncotarget. 2017;8(20):32905–32917. doi:10.18632/oncotarget.16462
29. Benezra R, Davis RL, Lockshon D, et al. The protein Id: a negative regulator of helix-loop-helix DNA binding proteins. Cell. 1990;61(1):49–59. doi:10.1016/0092-8674(90)90214-Y
30. Cheng D, Fan J, Ma Y, et al. LncRNA SNHG7 promotes pancreatic cancer proliferation through ID4 by sponging miR-342-3p. Cell Biosci. 2019;9:28. doi:10.1186/s13578-019-0290-2
31. Livak KJ, Schmittgen TD. Analysis of relative gene expression data using real-time quantitative PCR and the 2(-Delta Delta C(T)) Method. Methods. 2001;25(4):402–408. doi:10.1006/meth.2001.1262
32. Wang H, Xiao Y, Wu L, et al. Comprehensive circular RNA profiling reveals the regulatory role of the circRNA-000911/miR-449a pathway in breast carcinogenesis. Int J Oncol. 2018;52(3):743–754. doi:10.3892/ijo.2018.4265
33. Zhang Y, Zhang LX, Liu XQ, et al. Id4 promotes cell proliferation in hepatocellular carcinoma. Chin J Cancer. 2017;36(1):19. doi:10.1186/s40880-017-0186-7
34. Ebrahimi S, Hosseini M, Shahidsales S, et al. Targeting the Akt/PI3K signaling pathway as a potential therapeutic strategy for the treatment of pancreatic cancer. Curr Med Chem. 2017;24(13):1321–1331. doi:10.2174/0929867324666170206142658
 © 2020 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.
© 2020 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.