Back to Journals » OncoTargets and Therapy » Volume 13
Circ_0006528 Contributes to Paclitaxel Resistance of Breast Cancer Cells by Regulating miR-1299/CDK8 Axis
Authors Liu G, Zhang Z, Song Q, Guo Y, Bao P, Shui H
Received 7 March 2020
Accepted for publication 7 August 2020
Published 24 September 2020 Volume 2020:13 Pages 9497—9511
DOI https://doi.org/10.2147/OTT.S252886
Checked for plagiarism Yes
Review by Single anonymous peer review
Peer reviewer comments 2
Editor who approved publication: Dr Yao Dai
Guoqi Liu,1 Zhenxing Zhang,1 Qing Song,2 Yanling Guo,1 Puqiang Bao,1 Huifeng Shui1
1Department of Integrated Traditional Chinese and Western Medicine in Oncology, Luoyang Central Hospital Affiliated to Zhengzhou University, Luoyang, People’s Republic of China; 2Department of Oncology and Hematology, No. 989 Hospital of Joint Logistics Support Force of Chinese People’s Liberation Army, Pingdingshan, People’s Republic of China
Correspondence: Huifeng Shui
Department of Integrated Traditional Chinese and Western Medicine in Oncology, Luoyang Central Hospital Affiliated to Zhengzhou University, No. 288 Zhongzhou Middle Road, Luoyang 471009, Henan, People’s Republic of China
Tel +86-379-63892209
Email [email protected]
Background: Circular RNAs (circRNAs) have been reported to be involved in regulating the development of breast cancer. Paclitaxel (PTX) can be used for the chemotherapy of breast cancer. The study aimed to explore the role and mechanism of circ_0006528 in PTX-resistant breast cancer progression.
Methods: The levels of circ_0006528, microRNA-1299 (miR-1299) and cyclin-dependent kinase 8 (CDK8) were measured by quantitative real-time polymerase chain reaction (qRT-PCR). RNase R treatment was used to confirm that the circ_0006528 was a circular RNA. PTX resistance and cell proliferation were determined by Cell counting kit-8 (CCK-8) assay. Cell apoptosis, migration and invasion were analyzed by flow cytometry and Transwell assays, respectively. The levels of all proteins were examined by Western blot. The interaction between circ_0006528 and miR-1299 or CDK8 was predicted by online database confirmed by dual-luciferase reporter and RNA immunoprecipitation (RIP) assays. Xenograft mice model was constructed to reveal the role of circ_0006528 on tumor growth in vivo.
Results: Circ_0006528 was significantly up-regulated and miR-1299 was down-regulated in PTX-resistant breast cancer tissues and cells compared with control groups. CDK8 protein expression was dramatically upregulated in PTX-resistant breast cancer tissues and cells as compared to control groups. Loss-of-function experiments revealed that circ_0006528 knockdown decreased IC50 value of PTX and restrained proliferation, migration, invasion and autophagy, whereas induced apoptosis of PTX-resistant breast cancer cells in vitro. The inhibitory effects of sh-circ_0006528 on the progression of PTX-resistant breast cancer cells were reversed by decreasing miR-1299 or increasing CDK8 expression. Furthermore, circ_0006528 could modulate CDK8 expression by sponging miR-1299. Circ_0006528 silencing impeded the growth of PTX-resistant tumors by regulating miR-1299/CDK8 axis in vivo.
Conclusion: Circ_0006528 partially contributed to PTX resistance of breast cancer cells through up-regulating CDK8 expression by sponging miR-1299.
Keywords: breast cancer, paclitaxel resistance, circ_0006528, miR-1299, CDK8
Introduction
As the most common tumor in women’s gynecological diseases, breast cancer is still a thorny problem.1 Although techniques containing surgery, chemotherapy and radiotherapy have been improved, they still remain unsatisfactory, especially in chemotherapy. Chemoresistance is a major obstacle to clinical treatment.2 For example, paclitaxel (PTX) resistance to cancers could affect the treatment of cancers, thereby resulting in poor survival rate.3 Several studies have shown that breast cancer cells PTX were resistant to the chemotherapeutic drug PTX.4 The underlying mechanism of PTX resistance to cancers is complex, therefore, elucidating the mechanism of chemical resistance is necessary for effective treatment of breast cancer.
Growing evidences indicated that noncoding circular RNAs (circRNAs) can act as competing endogenous RNAs (ceRNAs) of microRNAs (miRNAs) to regulate the development of tumors by participating in drug resistance.5 For example, circRNA PVT1 could promote PTX resistance in gastric cancer cells by modulating ZEB1 expression through sponging miR-124-3p.6 It has also been found that circRNA CELSR1 impaired proliferation and apoptosis of ovarian cancer cells and accelerated PTX resistance via modulating miR-1252/FOXR2 pathway.7 Another example was that circPVT1 silencing could attenuate doxorubicin and cisplatin resistance in osteosarcoma cells by decreasing ABCB1 expression.8 In a previous study, it was validated that circ_0006528 could facilitate that progression of breast cancer via targeting miR-7-5p through activating MAPK/ERK pathway.9 However, whether circ_0006528 can mediate PTX resistance in breast cancer has not been studied.
MiR-1299 has been widely reported as a tumor inhibitor. For colon cancer, miR-1299 hindered the cells growth through down-regulating the expression of STAT3.10 For prostate cancer, miR-1299 was found to restrain the proliferation and metastasis of cells.11 Also, Meng et al supported that miR-1299 accelerated starvation and Rapamycin-induced autophagy of esophageal squamous cell carcinoma cells.12 Wang et al reported that miR-107 participated in regulating the sensitivity of breast cancer cells to PTX,13 which aroused our curiosity to explore the role of miR-1299 in PTX-resistant breast cancer.
Cyclin-dependent kinase8 (CDK8) belongs to the CDK family, and the members of this protein family influence a variety of biological behaviors, such as angiogenesis, stem cell self-renewal and sperm formation.14 Emerging studies indicated that CDK8 was overexpressed and accelerated the progression of tumor cells in multiple cancers, such as colon cancer,15 pancreatic cancer16 and glioma.17 Importantly, Li et al found that CDK8 was associated with sensitivity of PTX in NCI60 cells.18 But the interaction among circ_0006528, miR-1299 and CDK8 in breast cancer remains unknown. This study aimed to explore the function of circ_0006528 in PTX resistance of breast cancer, and clarify the potential mechanism of circ_0006528 in PTX-resistant breast cancer.
Materials and Methods
Tissue Samples
Tumor tissues and corresponding normal tissues were collected from 48 patients with breast cancer (33 PTX-chemosensitive patients and 15 PTX-chemoresistance patients) at Luoyang Central Hospital Affiliated to Zhengzhou University. Any treatment has not been conducted on these patients before the collection of tissues, and the patients signed the written informed consents. All the experiments in the present study were approved by the Ethics committee of Luoyang Central Hospital Affiliated to Zhengzhou University. Animal studies were performed in compliance with the ARRIVE guidelines and the Basel Declaration. All animals received humane care according to the National Institutes of Health (USA) guidelines.
PTX-Resistant Cells Construction and Culture
MCF10A, BT-549 and ZR-75-30 cell lines were purchased from American Tissue Culture Collection (ATCC, Manassas, VA, USA). These cells were cultured in Roswell Park Memorial Institute 1640 (RPMI 1640, Hyclone, South Logan, UT, USA) medium which included 10% fetal bovine serum (FBS, Gibco, Carlsbad, CA, USA) at 37°C. Besides, the PTX-resistant cell lines (BT-549/PTX and ZR-75-30/PTX) were established by successive adding paclitaxel. The PTX resistance was maintained by co-culture 1 μmol/L paclitaxel in RPMI 1640 (Hyclone) medium.
Quantitative Real-Time Polymerase Chain Reaction (qRT-PCR) and RNase R Treatment
Total RNA from milled tissues and harvested cells was extracted by Trizol (Invitrogen, Carlsbad, CA, USA) with RNeasy Mini Kit (QIAGEN, Shanghai, China). Complementary DNA (cDNA) was reversely transcribed from RNA by Prime Script RT Master Mix (Thermo Fisher Scientific, Waltham, MA, USA) and qRT-PCR was performed on 7500 Real-Time PCR System (Applied Biosystems, Foster City, CA, USA) with SYBR Select Master Mix (Applied Biosystems). The relative levels of circ_0006528 and CDK8 were normalized to glyceraldehyde-3-phosphate dehydrogenase (GAPDH) expression using 2−∆∆Ct method, and miR-1299 expression was normalized to U6. Primer sequences were synthesized by Sangon Biotech (Sangon, Shanghai, China) as following: circ_0006528, forward (F), 5ʹ-TACGGAGTCACTGCCTTACG-3ʹ; reverse (R), 5ʹ-TGCTATCCTAGAGAAGCTGCC-3ʹ. GAPDH, F, 5ʹ-GGAAAGCTGTGGCGTGAT-3ʹ; R, 5ʹ-AAGGTGGAAGAATGGGAGTT-3ʹ. MiR-1299, F, 5ʹ-ACACTCCAGCTGGGTTCTGGAAUUCTC-3ʹ; R, 5ʹ-CAGTGCGTGTCGTGGAGT-3ʹ. U6, F, 5ʹ-CTCGCTTCGGCAGCACA-3ʹ; R, 5ʹ- AACGCTTCACGAATTTGCGT-3ʹ. CDK8, F, 5ʹ-GCTGATAGGAAGGTGTGGCT-3ʹ; R, 5ʹ-GCAAAGCCCATGTCAGCAAT-3ʹ.
The circular structure of circ_0006528 was confirmed by RNase R (Geneseed, Guangzhou, China) treatment. The principle was that RNase R could digest linear RNA molecules, but not circular RNA molecules.19 In brief, the total RNA was incubated with RNase R in a 10 μL volume. After inactivation of RNase R, the treated RNAs were detected by qRT-PCR.
Transfection
Lentivirus-sh-circ_0006528, scrambled its matched negative control (sh-control), lentivirus-sh-CDK8 and the negative control (sh-control) were constructed by Thermo Fisher Scientific. MiR-1299-mimics, scrambled its control NC-mimics, miR-1299 inhibitor, scrambled its control NC inhibitor, overexpression plasmid of CDK8 (pcDNA3.1/CDK8) and its control pcDNA3.1 were bought from GenePharma (Shanghai, China). Transfection experiment in BT-549/PTX and ZR-75-30/PTX cells was performed by Lipofectamine 2000 (Thermo Fisher Scientific).
PTX Sensitivity and Cell Counting Kit-8 (CCK-8) Assays
The cell 50% inhibitory concentration (IC50) value of PTX and proliferation were examined by CCK-8 assay. For detecting IC50 value, the transfected BT-549/PTX and ZR-75-30/PTX cells (5×103 per well) were seeded into the 96-well plates for 24-h incubation. After treated with different concentrations of PTX, cells were continued to culture for 48 h. Subsequently, 10 μL CCK-8 solution (Dojindo Molecular Technologies, Shanghai, China) was added to the cells for another 4 h. The optical density (OD) at 450 nm was detected by a microplate reader (Promega Corporation, Fitchburg, WI, USA). The IC50 value of PTX was analyzed by the relative survival curve. For the cell proliferation assay, the transfected cells (5×103 per well) were seeded into 96-well plates. Ten microliters of CCK-8 were added at different time points (0 h, 24 h, 4872 h) after transfection, the detection was performed also as described above.
Flow Cytometry Assay
Cell apoptosis in the present study was assessed using an Annexin V fluorescein isothiocyanate (FITC)/propidium iodide (PI) apoptosis detection kit (Solarbio, Beijing, China). Briefly, the inoculated plates treated with BT-549/PTX and ZR-75-30/PTX cells (3×105 per well) were placed in an incubator for 48 h at 37°C. The cells were then harvested and stained with Annexin V-FITC and PI for 20 min without light and analyzed using a FACS Caliber flow cytometer (FlowCam, Shanghai, China).
Transwell Assay
Transwell assay was applied to examine the abilities of cells migration and invasion, the only difference was that detection of cell invasion required embedding Matrigel (BD, Franklin Lakes, NJ, USA). The two transfected cells were seeded into the upper chamber at a density of 1×105 per well with serum-free medium, while the bottom chamber was added with 600 μL of 10% FBS medium. After incubation for 24 h, cells migrated or invaded to the bottom chamber were stained with 0.1% crystal violet (Solarbio). Then, the cell number was examined by the Countess automatic cell counter (Invitrogen) with five random fields.
Western Blot
Proteins were obtained from tissues and cells by RIPA Buffer (Beyotime, Shanghai, China). Sodium dodecyl sulfate-polyacrylamide gel electrophoresis (SDS-PAGE, Beyotime) was applied to separate protein. Subsequently, proteins were transferred to 0.22 μm polyvinylidene difluoride (PVDF, Beyotime) membranes. After blocked in 5% non-fat milk for 2 h, the membranes were incubated overnight at 4°C with the primary antibodies against multi-drug resistance gene1 (MDR1, 1:200, Thermo Fisher Scientific), Light chain3-I (LC3-I, 1:2000, Abcam, Cambridge, MA, USA), LC3-II (1:1000, Abcam), P62 (2 μg/mL, Abcam), CDK8 (1:2000, Abcam) or GAPDH (1:3000, Abcam). The membranes were probed with horseradish peroxidase-conjugated goat anti-rabbit second antibody (1:4000, Abcam) for 1 h. The bands were observed by ECL reagent (R&S, Shanghai, China) and analyzed using ImageJ analysis software.
Dual-Luciferase Reporter Assay
The binding sites between circ_0006528 and miR-1299 were predicted by Circinteractome online database and the target gene of miR-1299 was predicted by TargetScanHuman 7.2 online database. The wild-type (WT) sequence of circ_0006528 containing miR-1229 binding sites and its mutant (MUT) sequence were cloned to the pmirGLO vector (Promega Corporation), named as circ_0006528-WT and circ_0006528-MUT. The 3ʹ untranslated region (UTR) of CDK8 containing miR-1299 binding sites and its mutant sequence were also implanted into the pmirGLO vector, named as CDK8-WT and CDK8-MUT. These plasmids were co-transfected with miR-1299-mimics or NC-mimics into BT-549/PTX and ZR-75-30/PTX cells, and the cells were further cultured for 48 h, then the relative luciferase activity was detected by a dual-luciferase reporter kit (Promega Corporation).
RNA Immunoprecipitation (RIP) Assay
RIP assay was performed using a RIP kit (Millipore, Billerica, MA, USA). BT-549/PTX and ZR-75-30/PTX cells were lysed by lysis buffer and magnetic beads were conjugated to anti-Ago2 antibody or IgG antibody. After digestion with protease K, the RNA was extracted by TRIzol (Invitrogen). Then, qRT-PCR assay was used to measure the expression of circ_0006528, miR-1299 or CDK8.
In vivo Experiment
ZR-75-30/PTX cells (2×106) transfected with lentivirus-mediated sh-circ_0006528 or sh-control were injected subcutaneously into the flank of male BALB/c nude mice (4 weeks old, n=8). The tumor size was measured once a week for a total of 4 times. The tumor volume was calculated according to the following formula: volume = length × width2/2. The tumor weight was weighed after the mice were euthanized, and the resected tumor tissues were frozen for qRT-PCR or Western blot analysis. The animal experiments were permitted by the Animal Care and Use Committee of Luoyang Central Hospital Affiliated to Zhengzhou University.
Statistical Analysis
All data were generated from at least three separate experiments and appeared as the mean ± standard deviation (SD). Student’s t-test was performed with the SPSS 18.0 software for statistical analysis. P value less than 0.05 was considered to be significant.
Results
Circ_0006528 Was Up-Regulated in PTX-Resistant Breast Cancer Tissues and Cells
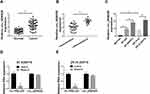
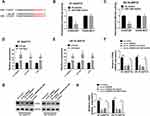
To study whether circ_0006528 was associated with breast cancer, we first examined the expression of circ_0006528 in breast cancer tissues by qRT-PCR. As shown in Figure 1A, circ_0006528 was up-regulated in breast cancer tissues compared with normal tissues (n=48). QRT-PCR was then performed to detect the expression of circ_0006528 in PTX-chemosensitive (n=33) and PTX-chemoresistance (n=15) breast cancer tissues. The results showed that circ_0006528 expression in PTX-chemoresistance group was higher than that in PTX-chemosensitive group (Figure 1B). Meanwhile, we found that circ_0006528 expression was up-regulated in PTX-resistant breast cancer cell lines (BT-549/PTX and ZR-75-30/PTX) in contrast to their parental cell lines (MCF10A, BT-549 and ZR-75-30) (Figure 1C). To validate that circ_0006528 was a circular RNA, total RNA from BT-549/PTX and ZR-75-30/PTX cells were treated with RNase R, which degraded linear RNA without affecting circular RNA. As expected, lnc PRELID2 expression in BT-549/PTX and ZR-75-30/PTX cells was significantly down-regulated after treatment with RNase R, while no significant change was observed in circ_0006528 expression (Figure 1D and E). These results demonstrated that dysregulation of circ_0006528 might be related to PTX resistance in breast cancer.
Silencing Circ_0006528 Repressed Proliferation, Migration, Invasion and Autophagy, Whereas Induced Apoptosis in PTX-Resistant Breast Cancer Cells in vitro
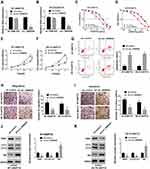
Given that circ_0006528 was up-regulated in PTX-resistant breast cancer, in subsequent experiments, the transfection efficiency of sh-circ_0006528 was firstly determined. QRT-PCR results showed that sh-circ_0006528 down-regulated the expression of circ_0006528 in BT-549/PTX and ZR-75-30/PTX cells, whereas the expression of lnc PRELID2 was no obvious change (Figure 2A and B). Then, the IC50 value was detected by CCK-8 assay for detecting the effect of circ_0006528 knockdown on PTX sensitive in breast cancer cells. As shown in Figure 2C and D, the IC50 value of PTX was declined in BT-549/PTX and ZR-75-30/PTX cells after circ_0006528 was knocked down. Moreover, CCK-8 assay showed that the proliferation of BT-549/PTX and ZR-75-30/PTX cells was markedly impaired by circ_0006528 knockdown (Figure 2E and F). The results of flow cytometry assay showed that circ_0006528 silencing in BT-549/PTX and ZR-75-30/PTX cells accelerated cell apoptosis (Figure 2G). Besides, Transwell assay was used to examine the migration and invasion of PTX-resistant breast cancer cells. The results indicated that circ_0006528 silencing inhibited the migration and invasion of BT-549/PTX and ZR-75-30/PTX cells (Figure 2H and I). Additionally, we examined the levels of the drug resistance-associated protein MDR1 and the autophagy-related proteins LC3-I, LC3-II and P62 by Western blot. Light chain3 (LC3) existed in the form of LC3-I under normal homeostatic conditions and bound with phosphatidylethanolamine (PE) to form the membrane-bound form LC3-II upon autophagy induction.20 As shown in Figure 2J and K, circ_0006528 knockdown in BT-549/PTX and ZR-75-30/PTX cells decreased the protein level of MDR1 and LC3-II/LC3-I ratio, and increased the protein level of P62, indicating that circ_0006528 could also induce autophagy. These results suggest that the down-regulation of circ_0006528 inhibited the progression of PTX-resistant breast cancer cells in vitro.
Circ_0006528 Directly Associated with miR-1299
We then searched the miRNAs associated with circ_0006528 by Circinteractome online database to explore the potential mechanism of circ_0006528 in PTX-resistant breast cancer progression. Results showed that miR-1299 might be a candidate (Figure 3A). The results of dual-luciferase reporter assay showed that miR-1299-mimics notably reduced the luciferase activity of BT-549/PTX and ZR-75-30/PTX cells in circ_0006528-WT group compared with circ_0006528-MUT group (Figure 3B and C). Also, RIP assay showed that the levels of circ_0006528 and miR-1299 in BT-549/PTX and ZR-75-30/PTX cells were obviously enhanced in anti-Ago2 group compared with anti-IgG group (Figure 3D and E). Subsequently, we validated that the expression pattern of miR-1299 was contrary to circ_0006528 in PTX-resistant breast cancer. And we found that miR-1299 expression was reduced in breast cancer tissues compared to normal tissues (Figure 3F) and was lower in PTX-chemoresistance group than that in PTX-chemosensitive group (Figure 3G). MiR-1299 expression was also down-regulated in PTX-resistant breast cancer cell lines (BT-549/PTX and ZR-75-30/PTX) relative to their parental cell lines (MCF10A, BT-549 and ZR-75-30) (Figure 3H). In addition, circ_0006528 knockdown significantly increased miR-1299 expression (Figure 3I) in BT-549/PTX and ZR-75-30/PTX. All these data indicated that circ_0006528 directly associated with miR-1299 and negatively regulated its expression in PTX-resistant breast cancer cells.
MiR-1299 Silencing Reversed the Inhibitory Effect of Circ_0006528 Knockdown on the Progression of PTX-Resistant Breast Cancer Cells
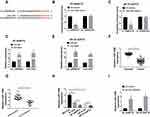
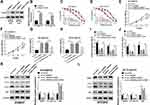
In view of the regulatory effect of circ_0006528 on miR-1299 expression, the effect of circ_0006528 on the progression of PTX-resistant breast cancer cells was correlative with miR-1299 to be further explored. As shown in Figure 4A and B, the inhibition of sh-circ_0006528 on the IC50 value of PTX was weakened by miR-1299 inhibitor in BT-549/PTX and ZR-75-30/PTX cells. CCK-8 assays showed that sh-circ_0006528 inhibited cell viability, whereas this effect was decreased by miR-1299 inhibitor in BT-549/PTX and ZR-75-30/PTX cells (Figure 4C and D). Flow cytometry analysis revealed that miR-1299 inhibitor hindered the promotion effect of circ_0006528 knockdown on cell apoptosis in BT-549/PTX and ZR-75-30/PTX cells (Figure 4E and F). Transwell assay revealed that knockdown of circ_0006528 impaired the migration and invasion of BT-549/PTX and ZR-75-30/PTX cells, which were then reversed by miR-1299 inhibitor (Figure 4G and H). Also, the effects of sh-circ_0006528 on the decrease of MDRI protein expression, the increase of P62 protein expression and the reduction of LC3-II/LC3-I ratio were overturned by interfering miR-1299 (Figure 4I and J). These results investigated that circ_0006528 promoted the progression of PTX-resistant breast cancer cells by targeting miR-1299.
Circ_0006528 Positively Regulated CDK8 Expression via Sponging miR-1299
We then sought to identify the downstream genes of circ_0006528 and found that there were putative binding sites between miR-1299 and 3ʹUTR of CDK8 by TargetScanHuman 7.2 online database (Figure 5A). Dual-luciferase reporter assay confirmed the molecular binding within miR-1299 and CDK8 (Figure 5B and C). Importantly, RIP assay results demonstrated that the levels of circ_0006528, miR-1299 and CDK8 in BT-549/PTX and ZR-75-30/PTX cells were dramatically increased in anti-Ago2 group than that in control group (Figure 5D and E). Furthermore, circ_0006528 silencing down-regulated the mRNA and protein levels of CDK8 in BT-549/PTX and ZR-75-30/PTX cells, while miR-1299 inhibitor reversed these effects (Figure 5FH). Taken together, the results indicated that circ_0006528 positively regulated CDK8 expression by sponging miR-1299, suggesting the possibility of circ_0006528/miR-1299/CDK8 pathway in PTX-resistant breast cancer cells.
CDK8 Promoted Proliferation, Migration, Invasion and Autophagy, Whereas Suppressed Apoptosis in PTX-Resistant Breast Cancer Cells
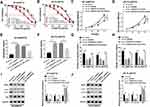
To investigate the functional effects of CDK8 on breast cancer progression, we examined its expression in breast cancer tissues, PTX-resistant tissues and cells. The results of qRT-PCR and Western blot showed that the mRNA and protein levels of CDK8 in breast cancer tissues and PTX-chemoresistance tissues were higher than that in normal tissues and PTX-chemosensitive tissues, respectively (Figure 6AC). Consistently, the protein level of CDK8 was elevated in PTX-resistant breast cancer cell lines (BT-549/PTX and ZR-75-30/PTX) compared to their parental cell lines (MCF10A, BT-549 and ZR-75-30) (Figure 6D). Considering that CDK8 was highly expressed in PTX-resistant breast cancer cell lines, the functional deletion experiments were performed. As shown in Figure 6E, CDK8 protein expression was obviously reduced in BT-549/PTX and ZR-75-30/PTX cells transfected with sh-CDK8 relative to that in cells transfected with sh-control. Then, we found that CDK8 knockdown decreased the IC50 value of PTX in BT-549/PTX and ZR-75-30/PTX cells (Figure 6F). The results of CCK-8, Flow cytometry and Transwell assays, respectively, indicated that the interference of CDK8 hindered proliferation (Figure 6G), migration (Figure 6I) and invasion (Figure 6J), whereas led to cell apoptosis (Figure 6H) in BT-549/PTX and ZR-75-30/PTX cells. Moreover, the down-regulation of CDK8 also reduced MDR1 protein level, upregulated P62 protein level and dwindled the ratio of LC3-II/LC3-I (Figure 6K and L). These data showed that CDK8 knockdown repressed breast cancer progression.
Circ_0006528 Promoted the Progression of PTX-Resistant Breast Cancer Cells by Up-Regulating the Expression of CDK8
Subsequently, the interaction between circ_0006528 and CDK8 on breast cancer progression was further studied. Western blot assay was first used to detect CDK8 expression in BT-549/PTX and ZR-75-30/PTX cells transfected with pcDNA3.1/CDK8 or pcDNA3.1. As shown in Figure 7A and B, CDK8 protein expression was enormously upregulated after pcDNA3.1/CDK8 transfection. IC50 curve test showed that the inhibitory effect of sh-circ_0006528 on the IC50 value in PTX in BT-549/PTX and ZR-75-30/PTX cells was inverted by CDK8 overexpression (Figure 7C and D). CCK-8 results showed that overexpression of CDK8 attenuated the suppressive impact of sh-circ_0006528 on proliferation of BT-549/PTX and ZR-75-30/PTX cells (Figure 7E and F). Flow cytometry revealed that circ_0006528 knockdown accelerated the apoptosis of BT-549/PTX and ZR-75-30/PTX cells, while this effect was alleviated by increasing CDK8 expression (Figure 7G and H). The results of Transwell assay showed that the inhibition of circ_0006528 silencing on migration and invasion of BT-549/PTX and ZR-75-30/PTX cells was reserved by up-regulating CDK8 expression (Figure 7I and J). Meanwhile, the up-regulation of P62 expression and the reduction of MDR1 expression and the ratio of LC3-II/LC3-I were observed after sh-circ_0006528 transfection in BT-549/PTX and ZR-75-30/PTX cells; however, CDK8 overexpression overturned the effects of sh-circ_0006528 on the levels of these proteins (Figure 7K and L). All these results revealed that circ_0006528 knockdown repressed breast cancer progression by regulating CDK8 expression.
Circ_0006528 Promoted the Tumor Growth of PTX-Resistant Breast Cancer Cells by Regulating miR-1299/CDK8 Axis in vivo
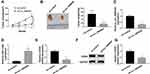
Xenograft mice assay in vivo was then performed to test the role of circ_0006528 on tumor growth using ZR-75-30/PTX cells in vivo. The results showed that circ_0006528 knockdown significantly reduced tumor volume (Figure 8A). Then, the excised tumors were measured and weighted, results revealed that the size and weight of tumors were dramatically decreased by circ_0006528 silencing (Figure 8B). QRT-PCR results showed that the level of circ_0006528 was downregulated by sh-circ_0006528 in removed tumor tissues from nude mice (Figure 8C). Subsequently, we found that miR-1299 expression was upregulated (Figure 8D) and CDK8 expression at mRNA and protein levels was down-regulated by circ_0006528 knockdown in vivo (Figure 8EG). These data indicated that circ_0006528 silencing suppressed the tumor growth of PTX-resistant breast cancer cells by regulating miR-1299/CDK8 axis in vivo.
 |
Figure 9 The regulatory mechanism of circ_0006528 in PTX-resistant breast cancer. |
Discussion
The mortality rate in women with breast cancer remains high worldwide.21 Latterly, drug chemotherapy was frequently used for the treatment of breast cancer patients.22 However, the chemoresistance makes most patients fail treatment. And the survival rates are extremely low.23 Circ_0006528 overexpression in PTX-resistant breast cancer cells attracted our attention.
Accumulating studies showed that circRNAs played a pivotal regulatory role in the chemoresistance of various tumors. For instance, circRNA_101505 expression was restrained in cisplatin-resistant hepatocellular carcinoma (HCC) tissues and cells, and up-regulation of circRNA_101505 impaired the growth of HCC cells in vivo and in vitro, which augmented cisplatin toxicity.24 Besides, Xu et al detected thousands of circRNAs changed significantly in A549/Taxol cells through high-throughput circRNA microarrays.25 In our present study, circ_0006528 was overexpressed in PTX-resistant breast cancer.
Paclitaxel has been shown to be useful in the chemotherapy of multiple cancers, including breast cancer,26 it retards cell cycle progression and induces apoptosis to inhibit tumor cell growth.27 In order to reveal the function of circ_0006528 in PTX-resistant breast cancer cells, loss-of-function assays were performed, and we found that its knockdown decreased the IC50 value of PTX as well as hindered cell proliferation, migration, invasion and autophagy, and induced cell apoptosis in breast cancer in vitro. Therefore, we hypothesized that circ_0006528 might exist in breast cancer as an oncogene, and its biological function might be related to PTX resistance.
Subsequently, we studied the potential mechanism of circ_0006528 in PTX-resistant breast cancer cells. The most classic study on the mechanism of circRNA was that circRNA acted as the ceRNA of miRNA to regulate the protein expression of target mRNA.28 Recently, Sang et al indicated that circRNA_0025202 increased FOXO3a expression via sponging miR-182-5p in breast cancer.29 We found that there were binding sites between circ_0006528 and miR-1299 through bioinformatics prediction tools, and confirmed circ_0006528 directly associated with miR-1299. As expected, miR-1299 was down-regulated in PTX-resistant breast cancer, and its silencing overturned the suppressive effect of sh-circ_0006528 on the breast cancer progression. Our finding was similar to previous studies, circRNA ciRS-7 could act as the ceRNA of miR-1299 to promote migration and invasion of triple-negative breast cancer cells.30 Taken together, circ_0006528 might regulate the resistance of breast cancer cells to PTX by binding to miR-1299.
Previously, a study showed that CDK8 was associated with the sensitivity of PTX.18 In addition, Tang et al reported that CDK8 was associated with autophagy.31 It is interesting that miR-1299 was able to target the 3ʹ-UTR of CDK8 in our results, and we found that circ_0006528 knockdown down-regulated CDK8 expression by sponging miR-1299. Additionally, down-regulation of CDK8 also retarded cell progression, which was in agreement with previous reports. Song et al demonstrated that CDK8 was increased in the trastuzumab-resistant breast cancer cells, and when CDK8 was silenced, the cell migration and invasion were restrained.32 Simultaneously, overexpression of CDK8 reversed the inhibition effect of sh-circ_0006528 on cell progression. Finally, in vivo experiments showed that circ_0006528 knockdown could repress the growth of PTX-resistant tumors through sponging miR-1299 and increasing the expression of CDK8. These results signified that circ_0006528 promoted cell proliferation, migration, invasion and autophagy, whereas inhibited cell apoptosis by sponging miR-1299 to upregulate CDK8 expression, which promoted PTX resistance in breast cancer (Figure 9).
Conclusion
Our results clarified the function of circ_0006528 in the PTX-resistant breast cancer. Circ_0006528 facilitated the PTX resistance in breast cancer tumorigenesis through modulating miR-1299/CDK8 axis, suggesting circ_0006528 might be a novel target in the treatment of PTX-resistant breast cancer.
Highlights
- Circ_0006528 was first verified to affect the progression of PTX-resistant breast cancer cells by targeting miR-1299.
- Circ_0006528 knockdown reduced autophagy of PTX-resistant breast cancer cells.
- The regulatory mechanism of circ_0006528 sponging miR-1229 to enhance CDK8 expression was first validated.
Abbreviations
circRNAs, circular RNAs; CDK8, cyclin-dependent kinase8; IC50, the cell 50% inhibitory concentration; RIP, RNA immunoprecipitation.
Data Sharing Statement
The analyzed data sets generated during the present study are available from the corresponding author on reasonable request.
Ethics Approval
The present study was approved by the ethical review committee of Luoyang Central Hospital Affiliated to Zhengzhou University.
Author Contributions
All authors contributed to data analysis, drafting or revising the article, have agreed on the journal to which the article will be submitted, gave final approval of the version to be published, and agree to be accountable for all aspects of the work.
Funding
No funding was received.
Disclosure
The authors declare that they have no competing interests for this work.
References
1. Ganz PA, Goodwin PJ. Breast cancer survivorship: where are we today? Adv Exp Med Biol. 2015;862:1–8. doi:10.1007/978-3-319-16366-6_1
2. Sutter SA, Slinker A, Balumuka DD, et al. Surgical management of breast cancer in africa: a continent-wide review of intervention practices, barriers to care, and adjuvant therapy. J Glob Oncol. 2017;3(2):162–168. doi:10.1200/JGO.2016.003095
3. Jiang N, Dong XP, Zhang SL, et al. Triptolide reverses the Taxol resistance of lung adenocarcinoma by inhibiting the NF-kappaB signaling pathway and the expression of NF-kappaB-regulated drug-resistant genes. Mol Med Rep. 2016;13(1):153–159. doi:10.3892/mmr.2015.4493
4. Sun Y, Yan L, Guo J, et al. Downregulation of SRSF3 by antisense oligonucleotides sensitizes oral squamous cell carcinoma and breast cancer cells to paclitaxel treatment. Cancer Chemother Pharmacol. 2019;84(5):1133–1143. doi:10.1007/s00280-019-03945-9
5. Huang X, Li Z, Zhang Q, et al. Circular RNA AKT3 upregulates PIK3R1 to enhance cisplatin resistance in gastric cancer via miR-198 suppression. Mol Cancer. 2019;18(1):71. doi:10.1186/s12943-019-0969-3
6. Yan-Yan L, Li-Ying Z, Wen-Zhen D. Circular RNA circ-PVT1 contributes to paclitaxel resistance of gastric cancer cells through the regulation of ZEB1 expression by sponging miR-124-3p. Biosci Rep. 2019;39:12. doi:10.1042/BSR20193045
7. Shu Z, Jie C, Chen-Lian Q, et al. CircCELSR1 (hsa_circ_0063809) contributes to paclitaxel resistance of ovarian cancer cells by regulating FOXR2 expression via miR-1252. Mol Ther Nucleic Acids. 2019;19:718–730. doi:10.1016/j.omtn.2019.12.005
8. Kun-Peng Z, Xiao-Long M, Chun-Lin Z. Overexpressed circPVT1, a potential new circular RNA biomarker, contributes to doxorubicin and cisplatin resistance of osteosarcoma cells by regulating ABCB1. Int J Biol Sci. 2018;14(3):321–330. doi:10.7150/ijbs.24360
9. Gao D, Qi X, Zhang X, et al. hsa_circRNA_0006528 as a competing endogenous RNA promotes human breast cancer progression by sponging miR-7-5p and activating the MAPK/ERK signaling pathway. Mol Carcinog. 2019;58(4):554–564. doi:10.1002/mc.22950
10. Wang Y, Lu Z, Wang N, et al. MicroRNA-1299 is a negative regulator of STAT3 in colon cancer. Oncol Rep. 2017;37(6):3227–3234. doi:10.3892/or.2017.5605
11. Zhang FB, Du Y, Tian Y, et al. MiR-1299 functions as a tumor suppressor to inhibit the proliferation and metastasis of prostate cancer by targeting NEK2. Eur Rev Med Pharmacol Sci. 2019;23(2):530–538. doi:10.26355/eurrev_201901_16865
12. Meng L, Liu S, Ding P, et al. Circular RNA ciRS-7 inhibits autophagy of ESCC cells by functioning as miR-1299 sponge to target EGFR signaling. J Cell Biochem. 2020;121(2):1039–1049. doi:10.1002/jcb.29339
13. Wang G, Ma C, Shi X, et al. miR-107 enhances the sensitivity of breast cancer cells to paclitaxel. Open Med. 2019;14:456–466. doi:10.1515/med-2019-0049
14. Lim S, Kaldis P. Cdks, cyclins and CKIs: roles beyond cell cycle regulation. Development. 2013;140(15):3079–3093. doi:10.1242/dev.091744
15. Liang J, Chen M, Hughes D, et al. CDK8 selectively promotes the growth of colon cancer metastases in the liver by regulating gene expression of TIMP3 and matrix metalloproteinases. Cancer Res. 2018;78(23):6594–6606. doi:10.1158/0008-5472.CAN-18-1583
16. Wei R, Kong L, Xiao Y, et al. CDK8 regulates the angiogenesis of pancreatic cancer cells in part via the CDK8-beta-catenin-KLF2 signal axis. Exp Cell Res. 2018;369(2):304–315. doi:10.1016/j.yexcr.2018.05.033
17. Zhang J-F, Zhang J-S, Zhao Z-H, et al. MicroRNA-770 affects proliferation and cell cycle transition by directly targeting CDK8 in glioma. Cancer Cell Int. 2018;18(1):195. doi:10.1186/s12935-018-0694-9
18. Li Z, Zhang Y, Zhang Z, et al. A four-gene signature predicts the efficacy of paclitaxel-based neoadjuvant therapy in human epidermal growth factor receptor 2-negative breast cancer. J Cell Biochem. 2019;120(4):6046–6056. doi:10.1002/jcb.27891
19. Xu L, Feng X, Hao X, et al. CircSETD3 (Hsa_circ_0000567) acts as a sponge for microRNA-421 inhibiting hepatocellular carcinoma growth. J Exp Clin Cancer Res. 2019;38(1):98. doi:10.1186/s13046-019-1041-2
20. Lai YC, Chuang YC, Chang CP, et al. Macrophage migration inhibitory factor has a permissive role in concanavalin A-induced cell death of human hepatoma cells through autophagy. Cell Death Dis. 2015;6:e2008. doi:10.1038/cddis.2015.349
21. Desantis CE, Ma J. Breast cancer statistics. CA Cancer J Clin. 2019;69(6):438–451. doi:10.3322/caac.21583
22. Chen G, Guo Z, Liu M, et al. Clinical value of capecitabine-based combination adjuvant chemotherapy in early breast cancer: a meta-analysis of randomized controlled trials. Oncol Res. 2017;25(9):1567–1578. doi:10.3727/096504017X14897173032733
23. Chen G, Sun L, Han J, et al. RILPL2 regulates breast cancer proliferation, metastasis, and chemoresistance via the TUBB3/PTEN pathway. Am J Cancer Res. 2019;9(8):1583–1606.
24. Luo Y, Fu Y, Huang R, et al. CircRNA_101505 sensitizes hepatocellular carcinoma cells to cisplatin by sponging miR-103 and promotes oxidored-nitro domain-containing protein 1 expression. Cell Death Discov. 2019;5:121. doi:10.1038/s41420-019-0202-6
25. Xu N, Chen S, Liu Y, et al. Profiles and bioinformatics analysis of differentially expressed circrnas in taxol-resistant non-small cell lung cancer cells. Cell Physiol Biochem. 2018;48(5):2046–2060. doi:10.1159/000492543
26. Greenlee H, Dupont-Reyes MJ, Balneaves LG, et al. Clinical practice guidelines on the evidence-based use of integrative therapies during and after breast cancer treatment. CA Cancer J Clin. 2017;67(3):194–232. doi:10.3322/caac.21397
27. Cretella D, Fumarola C, Bonelli M, et al. Pre-treatment with the CDK4/6 inhibitor palbociclib improves the efficacy of paclitaxel in TNBC cells. Sci Rep. 2019;9(1):13014. doi:10.1038/s41598-019-49484-4
28. Li X, Ding J, Wang X, et al. NUDT21 regulates circRNA cyclization and ceRNA crosstalk in hepatocellular carcinoma. Oncogene. 2020;39(4):891–904. doi:10.1038/s41388-019-1030-0
29. Sang Y, Chen B, Song X, et al. CircRNA_0025202 regulates tamoxifen sensitivity and tumor progression via regulating the miR-182-5p/FOXO3a axis in breast cancer. Mol Ther. 2019;27(9):1638–1652. doi:10.1016/j.ymthe.2019.05.011
30. Sang M, Meng L, Liu S, et al. Circular RNA ciRS-7 maintains metastatic phenotypes as a ceRNA of miR-1299 to target MMPs. Mol Cancer Res. 2018;16(11):1665–1675. doi:10.1158/1541-7786.MCR-18-0284
31. Tang HW, Hu Y, Chen CL, et al. The TORC1-regulated CPA complex rewires an RNA processing network to drive autophagy and metabolic reprogramming. Cell Metab 2018;27(5):1040–1054.e1048. doi:10.1016/j.cmet.2018.02.023
32. Song W, Wu S, Wu Q, et al. The microRNA-141-3p/CDK8 pathway regulates the chemosensitivity of breast cancer cells to trastuzumab. J Cell Biochem. 2019;120(8):14095–14106. doi:10.1002/jcb.28685
 © 2020 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.
© 2020 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.