Back to Journals » Research and Reports in Neonatology » Volume 4
Biomarkers for neonatal sepsis: recent developments
Authors Mally P, Xu J, Hendricks-Muñoz K
Received 2 May 2014
Accepted for publication 2 July 2014
Published 17 September 2014 Volume 2014:4 Pages 157—168
DOI https://doi.org/10.2147/RRN.S48316
Checked for plagiarism Yes
Review by Single anonymous peer review
Peer reviewer comments 3
Pradeep Mally,1 Jie Xu,2 Karen D Hendricks-Muñoz2
1Department of Pediatrics, Division of Neonatology, New York University School of Medicine, New York, NY, USA; 2Department of Pediatrics, Division of Neonatal Medicine, Virginia Commonwealth University School of Medicine, Richmond, VA, USA
Abstract: As a leading cause of neonatal morbidity and mortality, neonatal sepsis remains a significant global health challenge. Despite recent advances in the management of neonatal sepsis, including use of more potent antibiotics, timely identification continues to be a frequent and challenging problem in the management of the newborn or high-risk neonate in the neonatal intensive care unit. Lack of specific early objective diagnostic evaluations or specific signs and symptoms, especially in the preterm infant, impedes early identification. However, emerging technologies linked with enhanced understanding of the immature and developing neonatal immune system responses to early infection provide an opportunity to develop critically needed biomarkers to improve early identification in this high-risk population. This review will focus on the field of neonatal sepsis biomarker development, identifying current promising biomarkers that have been investigated and widely utilized, as well as provide insight into recent advances and the rapidly evolving technologies that are being exploited in biomarker development to improve diagnosis, treatment, and prognosis in neonatal sepsis.
Keywords: biomarker, cytokines, neonatal sepsis, recent developments, morbidity, mortality, neonates
Introduction
Neonatal sepsis can be devastating, leading to high morbidity and mortality in newborns, and is recognized as a global health challenge.1,2 The definition of early-onset sepsis is variable from ≤3 days (American Academy of Pediatrics definition) to ≤7 days (Centers for Disease Control definition based on epidemiology studies).3,4
The overall incidence of early-onset neonatal sepsis, occurring at ≤3 days of life, in North America is 0.76–0.77 cases per 1,000 live births, with a mortality rate of 24.4%.5 Globally more than a million newborns die in Third World countries each year from infections, with a risk of neurodevelopmental impairment seen in survivors.6–9 Those infants requiring intensive care in neonatal intensive care units (NICUs) have increased susceptibility to late-onset sepsis, occurring at >3 days of life, especially premature very low birth weight infants, in whom rates approach 36%.10 Despite recent advances in the management of neonatal sepsis, including use of more potent antibiotics and an array of sophisticated biomarkers to diagnose sepsis, timely identification continues to be a frequent and challenging problem in the management of the newborn or high-risk neonate in the NICU.11–13 The complex interaction between the functionally immature immune system of the newborn and the developing premature neonate linked with a wide spectrum of potential infecting organisms, ranging from hospital-acquired infections to those acquired from the mother transplacentally or during the birth process, contributes significantly to the limited diagnostic capabilities and adverse outcomes in this cohort population.14 Without early objective and specific diagnostic predictors that remain abnormal for a significant time to allow detection of neonatal sepsis, delay in therapy increases mortality and morbidity risk.15
Factors that delay diagnosis and initiation of therapy include lack of specific clinical features, as the infant often presents with subtle and nonspecific clinical signs and symptoms.16 Despite the low incidence of culture-proven neonatal sepsis, at approximately two per 1,000 live births,17 because of diagnostic testing limitations a significant number of neonates (up to 7%–13%) are routinely evaluated and treated for possible neonatal sepsis. Typical diagnostic parameters depend on conventional laboratory tests that are routinely serum based, such as white blood count (WBC), absolute neutrophil count (ANC), immature/total neutrophil (I/T) ratio, and variably obtained C-reactive protein (CRP)18,19 However, these conventional sepsis evaluation parameters have low sensitivity and are nonspecific, often demonstrating increased level response to various other neonatal conditions such as meconium aspiration, prolonged rupture of membranes, asphyxia, and the birth process.6,8,17,20–22 Indeed, recent information has suggested that the diagnostic accuracy of WBC, ANC, and I/T parameters may better predict sepsis using age-specific ratio normograms rather than fixed normal ranges.23 The definitive conventional diagnosis of sepsis rests upon isolation of pathogenic bacteria in blood cultures, which has low sensitivity and lacks timeliness to influence initiation of antibiotic therapy.24 Further diagnostic limitations of the blood culture method include a higher incidence of false negative results, due to low blood volume drawn for culture and antenatal antibiotic use that may influence subsequent bacterial growth.17 As a result, early antibiotic therapy is frequently initiated for presumed infection or delayed due to uncertainty increasing disease risk.
Thus, early, accurate, and rapid diagnosis of neonatal sepsis remains a major diagnostic challenge in neonatology, revealing the need for reliable and timely diagnostic biomarkers to enable clinicians to efficiently diagnose sepsis risk during the early phases of sepsis, provide effective antibiotic management tailored to causative organisms, and provide a useful guide for therapy during recovery. The objective of this review is to provide a review of biomarker developments for diagnosis and treatment of neonatal sepsis.
Methods
The studies used to evaluate biomarkers of neonatal sepsis were determined through a computer-based literature search in PubMed, the Cochrane Library, and Google. Initial search terms used in the retrieval of studies included “sepsis,” “neonatal,” “biomarkers,” “infant,” “omic,” and combinations of these terms. A specific further detailed literature search was used to identify and retrieve individual biomarker references as well as identify evolving novel techniques.
Biomarkers of sepsis
The properties of an ideal diagnostic biomarker include excellence in sensitivity and negative predicative value as well as excellent specificity and positive predictive value. Biomarker levels should change early in the disease course and remain altered for a period of time, to give an opportunity for clinicians to measure these biomarkers to optimize clinical management, monitor disease progress, and guide antimicrobial treatment.20 Discrimination between etiologies of sepsis such as viral, bacterial, or fungal etiology would be a valuable characteristic to assist clinicians in timely directed antibiotic therapy or antibiotic stewardship to avoid excess antibiotic use. Biomarkers should also assist in prediction of disease severity at the onset of infection and predict later prognosis with therapy. Ideally, measured biomarkers should be stable compounds that can be quantified with an easy method of measurement and have a quick turnaround time with low cost, so as to be used effectively as a routine measurement. Characteristics of an ideal biomarker are summarized in Table 1.
  | Table 1 Characteristics of an ideal biomarker for neonatal sepsis |
The long-established, widely used diagnostic practice for neonatal sepsis evaluation is the white cell analysis in the hematological counts, including the total WBC, ANC, and I/T ratio. Neutropenia is more predictive of sepsis than neutrophilia, which may be reflective of neonatal stress during birth, neonatal asphyxia, and also maternal hypertension.25 The use of the immature neutrophil count and I/T ratio continue to be useful in discriminating between host sepsis probability and are valuable screens, especially in the preterm infant.26–30 Furthermore, the presence of thrombocytopenia can provide an alert for illness severity.26 The addition of other biomarkers, such as CRP, may further enhance predictability of sepsis and assist in therapeutic management for the neonate.26 Though valuable, these hematologic tests do not provide absolute specific or sensitive diagnostic accuracy to assist in the decision to initiate treatment, particularly in the early phase of early-onset sepsis. As a result, promising biomarkers that target components of the complex early inflammatory response cascade provide the prospect to target early immune host response in the developmental stages of sepsis.
In the neonate, immune function is suboptimal and gestational age dependent, which contributes to a newborn’s enhanced susceptibility to infection and increased risk of morbidity and mortality from sepsis.31–34 Although the distinct neonatal innate immune system remains incompletely understood, dramatic progress in molecular characterization of this complex, intricate immune system has been delineated over the past decade, paving the way for the development, identification, and use of a variety of biomarkers with the potential to assist clinicians in the diagnosis of sepsis.35–40 Recent progressive advancement in biochemical and genetic research has led to the development of more sophisticated classes of biomarkers such as cytokines,41 chemokines,42,43 cell surface markers,44,45 and acute phase reactants. Additional sepsis biomarker candidates using genomics, proteomics, nucleic acid-based molecular techniques, and metabolomics are currently being evaluated, with the potential to revolutionize the diagnostic approach to neonatal sepsis and its management.
Common biomarkers used in neonatal sepsis
C-reactive protein
Among the acute phase reactants, CRP, produced in the liver, is a frequently used laboratory test for the diagnosis of neonatal sepsis.41,45–51 This biomarker has a half-life of 24–48 hours. Importantly, CRP takes 10–12 hours to respond after an infection, making it an unreliable marker of the initial stages of an acute infection.52 Given its response rate, serial CRP levels in combination with the absolute and complete WBC and I/T ratio has been widely used as a negative predictor of sepsis 24–48 hours after the onset of symptoms.53 Utilized in this manner, CRP can be a reliable late marker for sepsis with changing patterns or continuous decreased levels useful to monitor progress or guide clinicians in decisions related to duration of antibiotic treatment.54–56 In neonates, CRP has a higher sensitivity and specificity as a diagnostic marker of late onset sepsis than the neutrophil count and I/T ratio. A variety of noninfectious neonatal conditions such as meconium aspiration syndrome, delayed transition after birth, hemolysis, tissue injury, or surgery can also increase CRP reference values, including premature infant exposure to glucocorticoids, making it a nonspecific biomarker and presently an unreliable test to diagnose infectious risk during the early phase of a presumed neonatal infection.41,47,49
Procalcitonin
Procalcitonin (PCT) is the peptide prohormone of calcitonin and an acute phase reactant, independent of calcitonin levels, that is associated with the immunomodulation and vascular response associated with systemic inflammatory response syndrome (SIRS). Produced by monocytes and hepatocytes, concentrations of PCT increase early within 2–4 hours, after an exposure to a bacterial pathogen during the acute stage of sepsis. Levels peak at 6–8 hours and remain elevated for the next 24 hours, with a half-life of 24–30 hours.57 Uniquely, PCT responses and its concentrations do not appear to be affected by the gestational age at birth. However, there is a continuous change in the reference interval during the first 48 hours after birth. Additionally, identifying antibiotics identify total PCT, which includes PCT-1 and the major circulating form PCT-3.17 The discriminating responses of these PCT forms during development and with gestational age maturity may improve the specificity in early sepsis diagnosis.17 The early acute phase response kinetics of PCT makes it an appealing predictive biomarker for early diagnosis of neonatal sepsis as compared to the usefulness of CRP.57,58 Indeed, the diagnostic utility of PCT in early-onset infection has a sensitivity of 92%, specificity of 97%, positive predictive value of 94%, and negative predictive value of 96%.59 The response of PCT to early infection is more rapid than that of serum CRP levels, making it more useful for the clinicians in the diagnostic workup of a neonate. Furthermore, serum concentrations of PCT remain high when compared to other biomarkers such as tumor necrosis factor alpha (TNFα) and interleukin (IL)-6, making it more useful in predicting the severity of infection and outcomes response to treatment.59,60 However, PCT reliability as a single biomarker of neonatal infection is limited by its nonspecific elevations in healthy neonates over the first 48 hours of life. Furthermore, it can be falsely elevated in other noninfective conditions such as intracranial hemorrhage, birth asphyxia, and conditions associated with neonatal hypoxemia.61–64 Used in conjunction with CRP, a positive PCT level (≥2.5 ng/mL) increased the diagnosis of bacterial infection from 39% to 92%, while a negative CRP level (<40 mg/L) added diagnostic value through its negative predictive value, providing an avenue to guide antibiotic therapy or predict clinical disease course.65,66 Additional studies aimed at understanding species variability are needed to enhance the usefulness of PCT in diagnosis and prognosis of neonatal sepsis.
Developing biomarkers
Acute phase proteins and other proteins
Serum amyloid A
Serum amyloid A (SAA) is an apolipoprotein produced in the liver and an early acute phase reactant that has been studied, though not extensively, in neonates. SAA is derived from a variety of other tissues such as endothelial cells, monocytes, and smooth muscle cells and regulated by cytokines IL1 and IL6 as well as TNFα.67
Serum levels of SAA have a wide range, which increase with age from birth to adolescence; SAA is released as a response to infection and injury.68,69 SAA levels may be affected by hepatic function and host nutritional status, which may limit usefulness in late neonatal sepsis where hepatic dysfunction and nutritional status may be decreased.70 The des-arginine variant of SAA holds promise as a marker of acute and chronic inflammation.71 SAA is a reliable screening marker throughout the first 24 hours after the onset of infection, when compared to other biomarkers such as CRP and IL-6 with sensitivities that are suboptimal at different stages of an infection. In a study of 104 full-term infants <72 hours of age, levels of SAA at 0, 24, and 48 hours rose earlier than CRP levels with better diagnostic accuracy for predicting early-onset sepsis, including a sensitivity of 96% versus 30%, similar specificity, and greater positive predictive value (85% versus 78%), making SAA a superior marker of infection.72 Furthermore, serum levels of SAA show greater accuracy in the early suspicion of sepsis and are inversely associated with neonatal mortality and infection, enhancing their reliability and potential to prognosticate mortality in the early phase of an acute infection.73 Finally, the ability for this biomarker to be readily incorporated in clinical diagnostic evaluations increases its value as a potential biomarker.70
Lipopolysaccharide-binding protein
Lipopolysaccharide-binding protein (LPB), primarily produced by hepatocytes but also by epithelial and muscle cells, is a soluble pattern-recognition molecule important for interaction with endotoxin of gram-negative bacterial infections. LPB recognizes microbial-associated molecular patterns of bacteria to transport endotoxin to CD14 immune effector cells in response to infections.74,75 Binding to the lipopolysaccharide component of the bacteria, LPB forms a complex linked to the host macrophage to initiate a response to an acute infection.61 Levels of LPB peak early, within 6–8 hours, after an acute infection. As a result, LPB has a higher sensitivity and negative predictive value as a diagnostic test in early-onset sepsis when compared to other reactants such as CRP and PCT. Additionally, serum LBP levels are available using standardized analytical methods of clinical practice (Immulite, Siemens Healthcare) and have less physiological fluctuations during the first 48 hours of a newborn’s life. The test is less influenced by other obstetrical events when compared to other acute phase reactants, thus enhancing its applicability as a superior diagnostic marker of sepsis.14,76 Although LPB has promising potential, further research is required for neonatal sepsis evaluation before LBP can be used widely by the clinicians.
Apart from the acute-phase reactants that have been previously discussed, several other reactants show potential and have been studied as biomarkers of sepsis but are not routinely used by clinicians primarily because of low yield in diagnosing acute neonatal infection and low clinical availability compared to other sophisticated markers that possess better individual sensitivity and specificity in diagnosing infection. Some of these biomarkers that may prove to be valuable in combination with other sepsis markers are α-1 antitrypsin, fibronectin, haptoglobulin, lactoferrin, and neopterin.46,77–79
Cytokines and chemokines
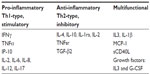
Chemokines are cytokines that have an ability to direct white cell migration. All cytokines are cellular signaling proteins that play a crucial role in modulating the host immunological response to infection.13,80,81 The development of a host immune response to a pathogen is dependent on host pathogen identification or host tissue damage identification. This identification allows the host to generate “danger” signals that are captured by pattern recognition receptors such as toll-like receptors (TLR), nucleotide-binding oligomerization domain-like receptors, and retinoic-acid-inducible protein 1-like receptors, present on the cell surface, to convey specificity in the innate immune response.82,83 As part of an intricate system of innate host immunity, activation of TLR’s by invading pathogens results in production of cytokines, chemokines, and coagulation proteins as well as complement; necessary to combat infections. Leukocyte immune regulation and trafficking into areas of tissue infection and injury is primarily controlled by chemokines and cytokines, which are broadly divided into the subsets of pro-inflammatory (Th1-type, stimulatory), anti-inflammatory (Th2-type, inhibitory) or multifunction depending on the final balance of their effects on the immune system (see Table 2 for cytokine classification). As functionally classified, proinflammatory cytokines include cytokines such as interferon-gamma, TNFα, inducible protein -10 (IP-10), and IL-2, IL-6, IL-8, IL-12, and IL-17. Multiple function inflammatory cytokines include cytokines such as IL-1β, monocyte chemoattractant protein (MCP-1), and soluble CD40 ligand (sCD40L) and growth factors, IL-3, granulocyte-colony stimulating factors and their secondary mediators, nitric oxide, thromboxanes, leukotrienes, platelet-activating factor, prostaglandins, and complements. These multiple function secondary mediators cause activation of the coagulation cascade, the complement cascade, as well as participate in the production of prostaglandins, leukotrienes, proteases, and oxidants. As a result of these highly regulated factors, most adverse effects of sepsis such as SIRS, disseminated intravascular coagulation (DIC), septic shock, multiple organ dysfunction syndrome, and complement activated response syndrome are associated with an imbalance in the production of pro-inflammatory mediators84–87 as well as the counterbalance synthesis of anti-inflammatory cytokines. Anti-inflammatory cytokines include IL-4, IL-10, tumor necrosis factor soluble receptor, IL-1 receptor alpha, and transforming growth factor beta 2 (TGF- β2). Thus, the evolution of disease and clinical symptoms during neonatal sepsis is dependent upon a complex and delicate balance between the pro-inflammatory, anti-inflammatory, and multiple function cytokines based on their final effect on the immune system.
IL-6
Among the group of pro-inflammatory cytokines, IL-6 has been widely investigated for its potential use as a biomarker of early neonatal sepsis.63,88–92 During the acute phase of an infection, B and T lymphocytes are stimulated to produce IL-6 cytokine, which in turn induces hepatocyte production of acute phase reactants such as CRP.93,94 As an early phase biomarker, IL-6 has superior sensitivity (90%) compared to CRP, with a negative predictive value of 91%.14,15,44 The limitations of using IL-6 solely as an early biomarker of neonatal sepsis is its very short half-life, with circulating concentrations that decrease precipitously following initiation of antimicrobial treatment to undetectable levels after 24 hours of life. This narrow window of opportunity has led to the use of IL-6 in conjunction with other biomarkers to improve its diagnostic usefulness in neonatal sepsis.95,96 The combined use of IL-6 with TNFα appears to convey greater sensitivity in diagnosing early infection than using either of these biomarkers alone. Indeed, combinations of IL-6, TNFα, and CRP led to sensitivities and negative predictive values that increased close to 90% in diagnosing early-onset neonatal infection.92
IL-8
IL-8, the only interleukin that belongs to the chemokine family, is a frequently studied pro-inflammatory cytokine for use as a marker of neonatal sepsis, with a sensitivity of 90% and specificity between 75%–100%. IL-8 regulates leukocyte migration and activation and has been extensively investigated in neonatal infection.97–100 Serum concentrations of IL-8 rise within 2–4 hours of an infection, and rapidly decline by 4 hours, making it useful as an early marker of infection, with greater sensitivity than CRP.90 Ng demonstrated that the use of IL-8 in conjunction with CRP as a biomarker of sepsis enhanced its diagnostic utility and reduced the use of antibiotics for presumed infection in neonates.11 Thus combining early and later cytokines may be a fruitful area of exploration in the early diagnosis of neonatal sepsis.
Anti-inflammatory cytokines
The inflammatory process is highly regulated by anti-inflammatory mediators such as IL-10 and TGF-β. Immunologically, these cytokines prevent an exaggerated pro-inflammatory response in reaction to pathogen invasion.101 In the premature infant, the ability to mount an aggressive anti-inflammatory response is limited, leading to increased susceptibility to target organ injury with excessive SIRS. Thus, in this population, serum values of anti-inflammatory cytokines have been recently studied to explore their ability to prognosticate improvement and survival.102,103 In the infant, an elevated anti-inflammatory (IL-10) to pro-inflammatory (TNFα) ratio is associated with severe late onset sepsis, similar to adult responses where elevated ratios were associated with greater morbidity.45,102 Implementation of anti- and pro-inflammatory ratio analysis emphasizes the importance of host immune balance in disease development and progression, providing an avenue to guide therapy in the prediction of later morbidity and mortality.
Other chemokines
Given their role in leukocyte migration, other chemokines investigated as markers of early neonatal infection are IP-10, monokine induced by interferon-gamma, regulated on activation, normal T cell expressed and secreted (RANTES), and MCP-1.43 Their use in combination with other inflammatory markers demonstrate better diagnostic utility in determining infection risk. Although further studies are needed to validate use in routine clinical practice, the combined measurements of IL-6, IL-10, and RANTES at the onset of infection potentially predicts greater morbidity in sepsis associated with DIC, suggesting that this methodology of targeting early inflammatory processes that include regulation of leukocyte activation holds promise as a diagnostic aid for clinicians in the future.42
Cell-surface antigens
Circulating inflammatory cells such as neutrophils, lymphocytes, monocytes, and natural killer cells express cell surface antigens, after activation by microbial products, that can be detected by flow cytometric technology.21 Several cell surface antigens such as CD11b, CD 14, CD64, CD32, CD16, CD69, and sCD163 have been identified to be promising in the detection of congenital sepsis, as well as early and late onset neonatal sepsis.45,93,104 Of this group, the most promising is neutrophil CD64.45,105 CD64 is a high affinity Fc receptor for immunoglobulin G that increases expression in response to infection.20,106 CD64 is a sensitive biomarker for diagnosing early-onset sepsis and intra-abdominal infections and is activated even before a rise in CRP levels is detected. CD64 has been shown to have a sensitivity of 80% and a negative predictive value of 89% in diagnosing early-onset sepsis and 24 hours after the infection, when both values rose to the high 90s.45,104 CD64 in combination with IL-6 or CRP had demonstrated a sensitivity and a negative predictive value close to 100% in diagnosing early-onset sepsis.47 Similar to CD64, CD11b up regulation represents an early-onset response to neonatal infection. CD11b differs from CD64 in that expression is variable, fluctuating frequently, and its expression is influenced by noninfectious conditions such as respiratory distress syndrome, making it a less reliable biomarker for early infection.80,81 CD14, a surface antigen present on inflammatory response cell surface membranes, appears to be involved in bacterial endotoxin signaling.107 The soluble fraction soluble CD14 subtype, sCD14-ST, or presepsin, is specifically developed in sepsis. Comparative studies with CRP, PCT, and IL6 using enzyme-linked immunoassay demonstrated that sCD14-ST had greater sepsis diagnostic sensitivity. Thus, this new biomarker may be a useful addition in early sepsis evaluation and awaits further testing.107 Soluble CD163 is a glycoprotein receptor that functions in the clearance of circulating free hemoglobin important in decreasing the effects of hemolysis-mediated oxidative damage.108 In its immune function, sCD 163 is a macrophage cell surface glycoprotein receptor with the potential to bind gram-negative and gram-positive bacteria and promote proinflammatory cytokines TNFα, IL-1β, IL-6, and IL-10.109 Recent studies have identified a valuable role for sCD163 in early-onset bacterial sepsis diagnosis where use of sCD163 improved differentiation between the noninfected and infected neonates before antibiotic initiation.110
There is tremendous potential in identification and use of surface antigen markers (Table 3). This technology, though not widely available for the routine diagnostic workup for neonatal sepsis, may be valuable in the future given the advantages of rapid presence during host response to sepsis, small blood sampling volumes, prompt turnover time, and wide time frame for blood sampling.
  | Table 3 Characteristics of common biomarkers |
Other biomarkers of interest
Other recent biomarkers of interest that are being explored include pentraxin 3 (PTX3), angiopoietins, suPAR, and soluble triggering receptor expressed on myeloid cells-1 (sTREM-1), and deserve mention here. PTX is an acute phase glycoprotein in the family of CRP and shares 98% homology with tumor necrosis factor. Synthesis occurs in endothelial and inflammatory cells in response to endotoxin and cytokines. High levels of PTX3 have been identified as a valid prognostic indicator in pediatric meningococcal shock, and usefulness is being explored in the neonatal population.111 Angiopoietins (Ang)-1 and Ang-2 are antagonistic endothelial cell growth factors widely studied in proliferative diseases such as retinopathy.112 Both angiopoietins are important in vascular permeability and function, with Ang-2 associated with greater vascular permeability such as host responses to TNFα and IL-6. Lower Ang-1 and higher Ang-2 concentrations predict poor prognostic outcome in children with sepsis, with low levels of Ang-1 predicting high mortality. Thus, Ang-1 and Ang-2 may be useful biomarkers for response to therapy in the neonate with sepsis.113 The soluble form of the urokinase-type plasminogen activator receptor, suPAR, is expressed on immune and endothelial cells. Existing in three forms with little concentration variability, suPAR has a variety of immune regulatory roles.114 Used in the early diagnosis of neonatal sepsis, suPAR levels correlated well with CRP levels in infected infants with greater levels in septic infants at admission. Post recovery, suPAR decreased but remained greater than in control infants, suggesting a biomarker role in infection diagnosis but less reliable for antibiotic stewardship or host response to therapy.115 Soluble triggering receptor is a newly explored biomarker of the immunoglobulin family expressed by phagocytes (sTREM-1).116 Involved in the innate inflammatory response and sepsis, sTREM-1 may be useful as a biomarker of early sepsis and sepsis severity. sTREM appears to correlate with infant white cell counts and ratios. At concentration limits of 310 pg/mL, sensitivity and specificity reached 100%, suggesting a role in early diagnosis of neonatal sepsis.117
Technical frontiers in diagnosing neonatal sepsis
Several novel approaches and techniques that have cutting-edge potential to provide rapid and specific identification of pathogens have generated a great deal of interest amongst researchers and clinicians interested in neonatal sepsis diagnosis. These include combinations of biomarkers studied by proteomics-based research and identification of sepsis based on gene expression profiling.
Molecular technology
A major disadvantage of using the conventional gold standard method of blood culture in diagnosing infection is the prolonged periods of time needed for incubation and identification of isolates, the false negative results that are obtained when pretreated with antibiotics, and the technical difficulties in obtaining adequate sample volumes. Thus, the application of molecular techniques and biomarker panels may have the opportunity to avoid these problems to rapidly predict sepsis. Emerging molecular techniques using multiplex platforms that can measure multiple markers such as protein, deoxyribonucleic acid (DNA), and ribonucleic acid (RNA) using technologies such as fluorescence in situ hybridization (FISH), quantitative polymerase chain reaction (qPCR), 16 S rRNA, and miRNA detection can revolutionize the diagnostic approach to neonatal sepsis.84 qPCR is a rapid test that can be used to detect bacterial DNA in body fluids of the host suspected of having an infection. FISH can significantly reduce the time required to identify organisms isolated in culture. The use of the probe-based specific qPCR can reduce diagnostic time, help in identifying bacterial species, and improve specificity and positive predictive value.84 Molecular technology also provides opportunities to further understanding of underlying pathological events that can be useful in development of new diagnostic approaches, algorithms, or score systems. Limitations of this technology for rapid bacterial detection are the lack of availability of all existing genetic sequences for microbiota, many of which are undiscovered. Additionally, resistant cell wall structures of certain bacteria limit the digestion, DNA extraction, and analysis, requiring further technological refinement before this application is widely used in the neonatal population.84
Use of other body fluids in diagnosis of neonatal sepsis
Given their noninvasive collection and ease of accessibility, development of biomarkers for neonatal sepsis has recently focused on the use of saliva and urine as surrogates for early infection detection or infection risk. Saliva is an abundant body fluid that is readily accessible for use in the analysis of systemic illness in the critically ill infant. Recent technological advances in the “omics” described below provide the opportunity to evaluate salivary potential as a tool in the analysis of neonatal sepsis.118,119 Current applications have demonstrated that saliva can be used for assessment of the microbiome response to environmental exposures, and also as a tool to study cytokine changes in disease risk states, providing an avenue open to exploration in early diagnosis of neonatal sepsis risk in the future.120,121 Urine is an additional body fluid that is readily accessible and noninvasively collected to be used as a marker for infection analysis. Development and use of biomarker panels in biofluid analysis hold the potential to provide early detection of infection risk that may be valuable in clinical decision-making. Recently, urinary cytokines IL8, IP10, and MCP-1 have been shown to be elevated during early presumed infection in a cohort of healthy and at-risk term infants, identifying a role for urine as a potential biomarker in infant early infection.122 These and other findings suggest that other body fluids may hold promise as clinical adjuncts to understand the infant’s physiological responses to neonatal sepsis and assist in the development of potential biomarkers of neonatal sepsis early diagnosis or effective treatment management.122
“Omics” technology: genomics, proteomics, metabolomics
Recent advances in “omics” research present new opportunities to complement current traditional approaches with rapidly available information related to expression of a large number of proteins or functioning metabolites that are characteristic for neonatal sepsis. This frontier is an exciting focus for researchers, who study neonatal sepsis and provide avenues to integrate systems biology or “omics” technology into clinical care. From a practical standpoint for neonatal sepsis, these available technologies provide the capability to unify multiple levels of information (genomic, proteomic, and metabolomics) to form a more complete understanding of the sepsis condition.123 Genomics technology has the potential to identify genes that demonstrate altered regulation during infection. As the circulating concentrations of inflammatory markers such as the cytokines, chemokines, and acute phase reactants may not fully represent the complete host response to an infection, these technologies push researchers to focus on chemokine mRNA expression and its relation to infection and other stressors such as birth asphyxia.11,85 Recent reports demonstrate an association of elevated IL-8 mRNA expression in neonates exposed to perinatal infection compared to mRNA expression of IL-8 and MCP-1 expression changes in neonates with perinatal asphyxia.85 Proteomic and metabolomics profiling technologies provide insight into the functional expression of proteins or metabolites present in a biological sample. The aim of proteomics is to obtain insight into the functional expression of proteins during neonatal sepsis, to identify different pattern of protein expression in different individuals with and without diseases, and to discover host response biomarkers that will be useful in the diagnostic process of an acute infection. Proteomic processing high throughput technology involves separation of proteins based on intrinsic properties such as molecular weight, isoelectric point, or affinity to metals or antibodies. This information offers a new approach to identify protein signatures and functionally expressed metabolites occurring during specific disease states. Metabolomics technology, using spectrometric techniques such as nuclear magnetic resonance and mass spectrometry, provides a window to gene and environmental (eg, neonatal sepsis) interactions using a variety of samples (blood, urine, or tissue).124,125 Metabolites by nature are dynamic and functional as they have specific connections and patterns for the host that are identified during clinical sepsis and can be expressed rapidly, varying within seconds. Thus metabolites provide valuable information related to host response during the course of sepsis. “Omics” technology offer expanded understanding of host physiology. Current limitations of these technologies are adequate “disease exposed” samples and appropriate controls and skilled analysis of the complex generated datasets. For some “omics,” such as metabolomics, the analysis is time consuming in their current form and not useful for early decisions to initiate sepsis therapy. Nevertheless, “omics” technologies is an investigational area that holds unique opportunities to simultaneously identify and translate pathologic infection-host influences into future diagnostic biomarker developments to target early identification of neonatal sepsis in the future.126
Conclusion
The conventional hematological and microbiological techniques that are routinely used to diagnose neonatal sepsis remain unreliable in the face of an associated high mortality and serious morbidity. The search for an ideal biomarker or biomarkers that provide early, specific, and reliable identification of the neonate at risk for infection has been enhanced by the potential use of unique technological developments and evolving understanding of current biomarker strengths and limitations. Advanced understanding of the neonatal immune system capabilities in response to infection risk has further led to identification of several promising potential biomarkers that may translate to improved diagnosis, treatment, and prognosis of neonatal sepsis in the future. Despite many promising candidate biomarkers, to date no single biomarker, combination of biomarkers, or score system can exclusively be considered in the accurate diagnosis of early neonatal sepsis. Only CRP and PCT have undergone sufficient studies and continue to be studied with serial measurements. Combined with other biomarkers or scoring systems, these biomarkers may offer improved diagnostic and treatment sensitivity. For instance, persistently normal CRP levels have a strong negative predictive value, which can add value when linked with other markers. Future studies are needed to outline PCT responses and PCT species changes during infant development as well as create gestational age reference intervals to assist clinicians in understanding the usefulness of PCT for early neonatal sepsis diagnosis. Several biomarkers are nonspecific, impacted by host environmental exposures, host changing developmental profiles, or by clinical therapies or conditions. Our knowledge of the pathophysiology of neonatal sepsis is expanding, and exploratory research using available molecular microbial techniques as well as the use of “omics” technological capabilities to identify disease profiles has great potential to advance diagnostic capabilities in the future. The clinician remains challenged in the need for optimal biomarker evidence from randomized control trials, which are extremely limited in this critical population. The unique advantages of metabolomics technology are that this technique offers a dynamic view of host functional responses during health and disease to offer early and rapid identification of sepsis.124,125 These advances should provide the diagnostic accuracy needed for early identification and assist clinicians in understanding neonatal host immune responses to appropriately guide antibiotic and other therapies in the care of the infant who presents with neonatal sepsis.
Disclosure
The authors report no conflicts of interest in this work.
References
Stoll BJ, Hansen N, Fanaroff AA, et al. Late-onset sepsis in very low birth weight neonates: the experience of the NICHD Neonatal Research Network. Pediatrics. 2002;110(2 Pt 1):285–291. | |
Tsai MH, Chu SM, Lee CW, et al. Recurrent late-onset sepsis in the neonatal intensive care unit: incidence, clinical characteristics and risk factors. Clin Microbiol Infect. 2014. | |
Polin RA. Committee on Fetus and Newborn. Management of neonates with suspected or proven early-onset bacterial sepsis. Pediatrics. 2012;129(5):1006–1015. | |
Centers for Disease Control and Prevention (CDC). Group B Strep Infection in Newborns [webpage on the Internet]. Available at: http://www.cdc.gov/groupbstrep/about/newborns-pregnant.html. Last updated November 18, 2010. | |
Ohlsson A, Shah VS. Intrapartum antibiotics for known maternal Group B streptococcal colonization. Cochrane Database Syst Rev. 2014;6:CD007467. | |
Garcia-Prats JA, Cooper TR, Schneider VF, Stager CE, Hansen TN. Rapid detection of microorganisms in blood cultures of newborn infants utilizing an automated blood culture system. Pediatrics. 2000;105(3 Pt 1):523–527. | |
Stoll BJ, Hansen NI, Adams-Chapman I, et al. National Institute of Child Health and Human Development Neonatal Research Network. Neurodevelopmental and growth impairment among extremely low-birth-weight infants with neonatal infection. JAMA. 2004;292(19):2357–2365. | |
Ganatra HA, Stoll BJ, Zaidi AK. International perspective on early-onset neonatal sepsis. Clin Perinatol. 2010;37(2):501–523. | |
Pappas A, Kendrick DE, Shankaran S, et al. Eunice Kennedy Shriver National Institute of Child Health and Human Development Neonatal Research Network. Chorioamnionitis and early childhood outcomes among extremely low-gestational-age neonates. JAMA Pediatr. 2014;168(2):137–147. | |
Stoll BJ, Hansen NI, Bell EF, et al. Neonatal outcomes of extremely preterm infants from the NICHD Neonatal Research Network. Pediatrics. 2010;126(3):443–456. | |
Ng PC. Diagnostic markers of infection in neonates. Arch Dis Child Fetal Neonatal Ed. 2004;89(3):F229–F235. | |
Mehr S, Doyle LW. Cytokines as markers of bacterial sepsis in newborn infants: a review. Pediatr Infect Dis J. 2000;19(9):879–887. | |
Commins SP, Borish L, Steinke JW. Immunologic messenger molecules: cytokines, interferons, and chemokines. J Allergy Clin Immunol. 2010; 125(2 Suppl 2):S53–S72. | |
D’Alquen D, Kramer BW, Seidenspinner S, et al. Activation of umbilical cord endothelial cells and fetal inflammatory response in preterm infants with chorioamnionitis and funisitis. Pediatr Res. 2005;57(2):263–269. | |
Ng PC, Lam HS. Diagnostic markers for neonatal sepsis. Curr Opin Pediatr. 2006;18(2):125–131. | |
Lever A, Mackenzie I. Sepsis: definition, epidemiology, and diagnosis. BMJ. 2007;335(7625):879–883. | |
Buttery JP. Blood cultures in newborns and children: optimising an everyday test. Arch Dis Child Fetal Neonatal Ed. 2002;87(1):F25–F28. | |
Christensen RD, Baer VL, Gordon PV, et al. Reference ranges for lymphocyte counts of neonates: associations between abnormal counts and outcomes. Pediatrics. 2012;129(5):e1165–e1172. | |
Chiesa C, Natale F, Pascone R, et al. C reactive protein and procalcitonin: reference intervals for preterm and term newborns during the early neonatal period. Clin Chim Acta. 2011;412(11–12):1053–1059. | |
Ng PC, Lam HS. Biomarkers for late-onset neonatal sepsis: cytokines and beyond. Clin Perinatol. 2010;37(3):599–610. | |
Lam HS, Ng PC. Biochemical markers of neonatal sepsis. Pathology. 2008;40(2):141–148. | |
van Dissel JT, van Langevelde P, Westendorp RG, Kwappenberg K, Frölich M. Anti-inflammatory cytokine profile and mortality in febrile patients. Lancet. 1998;351(9107):950–953. | |
Newman TB, Puopolo KM, Wi S, Draper D, Escobar GJ. Interpreting complete blood counts soon after birth in newborns at risk for sepsis. Pediatrics. 2010;126(5):903–909. | |
Griffin MP, Lake DE, Bissonette EA, Harrell FE, O’Shea TM, Moorman JR. Heart rate characteristics: novel physiomarkers to predict neonatal infection and death. Pediatrics. 2005;116(5):1070–1074. | |
Philip AG, Hewitt JR. Early diagnosis of neonatal sepsis. Pediatrics. 1980;65(5):1036–1041. | |
Makkar M, Gupta C, Pathak R, Garg S, Mahajan NC. Performance evaluation of hematologic scoring system in early diagnosis of neonatal sepsis. J Clin Neonatol. 2013;2(1):25–29. | |
Rodwell RL, Leslie AL, Tudehope DI. Early diagnosis of neonatal sepsis using a hematologic scoring system. J Pediatr. 1988;112(5):761–767. | |
Polin RA, Randis TM. Biomarkers for late-onset neonatal sepsis. Genome Med. 2010;2(9):58. | |
Khair KB, Rahman MA, Sultana T, Roy CK, Rahman MQ, Ahmed AN. Early diagnosis of neonatal septicemia by hematologic scoring system, C-reactive protein and serum haptoglobin. Mymensingh Med J. 2012;21(1):85–92. | |
Khair KB, Rahman MA, Sultana T, Roy CK, Rahman MQ, Ahmed AN. Early diagnosis of neonatal septicemia by hematologic scoring system, C-reactive protein and serum haptoglobin. Mymensingh Med J. 2012;21(1):85–92. | |
Wynn JL, Neu J, Moldawer LL, Levy O. Potential of immunomodulatory agents for prevention and treatment of neonatal sepsis. J Perinatol. 2009;29(2):79–88. | |
Levy O. Innate immunity of the newborn: basic mechanisms and clinical correlates. Nat Rev Immunol. 2007;7(5):379–390. | |
Adkins B, Leclerc C, Marshall-Clarke S. Neonatal adaptive immunity comes of age. Nat Rev Immunol. 2004;4(7):553–564. | |
Kollmann TR, Crabtree J, Rein-Weston A, et al. Neonatal innate TLR-mediated responses are distinct from those of adults. J Immunol. 2009;183(11):7150–7160. | |
Rittirsch D, Flierl MA, Ward PA. Harmful molecular mechanisms in sepsis. Nat Rev Immunol. 2008;8(10):776–787. | |
Hotchkiss RS, Karl IE. The pathophysiology and treatment of sepsis. N Engl J Med. 2003;348(2):138–150. | |
Abraham E, Singer M. Mechanisms of sepsis-induced organ dysfunction. Crit Care Med. 2007;35(10):2408–2416. | |
Cinel I, Dellinger RP. Advances in pathogenesis and management of sepsis. Curr Opin Infect Dis. 2007;20(4):345–352. | |
Jean-Baptiste E. Cellular mechanisms in sepsis. J Intensive Care Med. 2007;22(2):63–72. | |
Sriskandan S, Altmann DM. The immunology of sepsis. J Pathol. 2008;214(2):211–223. | |
Ng PC, Cheng SH, Chui KM, et al. Diagnosis of late onset neonatal sepsis with cytokines, adhesion molecule, and C-reactive protein in preterm very low birthweight infants. Arch Dis Child Fetal Neonatal Ed. 1997;77(3):F221–F227. | |
Ng PC, Li K, Leung TF, et al. Early prediction of sepsis-induced disseminated intravascular coagulation with interleukin-10, interleukin-6, and RANTES in preterm infants. Clin Chem. 2006;52(6):1181–1189. | |
Ng PC, Li K, Chui KM, et al. IP-10 is an early diagnostic marker for identification of late-onset bacterial infection in preterm infants. Pediatr Res. 2007;61(1):93–98. | |
Turunen R, Andersson S, Nupponen I, Kautiainen H, Siitonen S, Repo H. Increased CD11b-density on circulating phagocytes as an early sign of late-onset sepsis in extremely low-birth-weight infants. Pediatr Res. 2005;57(2):270–275. | |
Ng PC, Li K, Wong RP, Chui KM, Wong E, Fok TF. Neutrophil CD64 expression: a sensitive diagnostic marker for late-onset nosocomial infection in very low birthweight infants. Pediatr Res. 2002;51(3):296–303. | |
Speer C, Bruns A, Gahr M. Sequential determination of CRP, alpha 1- antitrypsin and haptoglobin in neonatal septicaemia. Acta Paediatr Scand. 1983;72(5):679–683. | |
Seibert K, Yu VY, Doery JC, Embury D. The value of C-reactive protein measurement in the diagnosis of neonatal infection. J Paediatr Child Health. 1990;26(5):267–270. | |
Berger C, Uehlinger J, Ghelfi D, Blau N, Fanconi S. Comparison of C-reactive protein and white blood cell count with differential in neonates at risk for septicaemia. Eur J Pediatr. 1995;154(2):138–144. | |
Da Silva O, Ohlsson A, Kenyon C. Accuracy of leukocyte indices and C-reactive protein for diagnosis of neonatal sepsis: a critical review. Pediatr Infect Dis J. 1995;14(5):362–366. | |
Bilavsky E, Yarden-Bilavsky H, Ashkenazi S, Amir J. C-reactive protein as a marker of serious bacterial infections in hospitalized febrile infants. Acta Paediatr. 2009;98(11):1776–1780. | |
Cetinkaya M, Ozkan H, Köksal N, Celebi S, Hacimustafaoglu M. Comparison of serum amyloid A concentrations with those of C-reactive protein and procalcitonin in diagnosis and follow-up of neonatal sepsis in premature infants. J Perinatol. 2009;29(3):225–231. | |
Hofer N, Zacharias E, Müller W, Resch B. An update on the use of C-reactive protein in early-onset neonatal sepsis: current insights and new tasks. Neonatology. 2012;102(1):25–36. | |
Clyne B, Olshaker JS. The C-reactive protein. J Emerg Med. 1999;17(6):1019–1025. | |
Pourcyrous M, Bada HS, Korones SB, Baselski V, Wong SP. Significance of serial C-reactive protein responses in neonatal infection and other disorders. Pediatrics. 1993;92(3):431–435. | |
Kawamura M, Nishida H. The usefulness of serial C-reactive protein measurement in managing neonatal infection. Acta Paediatr. 1995;84(1):10–13. | |
Franz AR, Steinbach G, Kron M, Pohlandt F. Reduction of unnecessary antibiotic therapy in newborn infants using interleukin-8 and C-reactive protein as markers of bacterial infections. Pediatrics. 1999;104(3 Pt 1):447–453. | |
Tang BM, Eslick GD, Craig JC, McLean AS. Accuracy of procalcitonin for sepsis diagnosis in critically ill patients: systematic review and meta-analysis. Lancet Infect Dis. 2007;7(3):210–217. | |
Luzzani A, Polati E, Dorizzi R, Rungatscher A, Pavan R, Merlini A. Comparison of procalcitonin and C-reactive protein as markers of sepsis. Critical Care Medicine. 2003;31(6):1737–1741. | |
Chiesa C, Panero A, Rossi N, et al. Reliability of procalcitonin concentrations for the diagnosis of sepsis in critically ill neonates. Clin Infect Dis. 1998;26(3):664–672. | |
Whicher J, Bienvenu J, Monneret G. Procalcitonin as an acute phase marker. Ann Clin Biochem. 2001;38(Pt 5):483–493. | |
Pavcnik-Arnol M, Hojker S, Derganc M. Lipopolysaccharide-binding protein in critically ill neonates and children with suspected infection: comparison with procalcitonin, interleukin-6, and C-reactive protein. Intensive Care Med. 2004;30(7):1454–1460. | |
van Rossum AMC, Wulkan RW, Oudesluys-Murphy AM. Procalcitonin as an early marker of infection in neonates and children. Lancet Infect Dis. 2004;4(10):620–630. | |
Resch B, Gusenleitner W, Müller W. Procalcitonin, interleukin-6, C-reactive protein and leukocyte counts in infants with bronchiolitis. Pediatr Infect Dis J. 2003;22(5):475–476. | |
Enguix A, Rey C, Concha A, Medina A, Coto D, Diéguez MA. Comparison of procalcitonin with C-reactive protein and serum amyloid for the early diagnosis of bacterial sepsis in critically ill neonates and children. Intensive Care Med. 2001;27(1):211–215. | |
Simon L, Saint-Louis P, Amre DK, Lacroix J, Gauvin F. Procalcitonin and C-reactive protein as markers of bacterial infection in critically ill children at onset of systemic inflammatory response syndrome. Pediatr Crit Care Med. 2008;9(4):407–413. | |
Kordek A, Hałasa M, Podraza W. Early detection of an early onset infection in the neonate based on measurements of procalcitonin and C-reactive protein concentrations in cord blood. Clin Chem Lab Med. 2008;46(8):1143–1148. | |
Yuan H, Huang J, Lv B, et al. Diagnosis value of the serum amyloid A test in neonatal sepsis: a meta-analysis. Biomed Res Int. 2013;2013:520294. | |
He R, Sang H, Ye RD. Serum amyloid A induces IL-8 secretion through a G protein-coupled receptor, FPRL1/LXA4R. Blood. 2003;101(4):1572–1581. | |
Lannergård A, Friman G, Ewald U, Lind L, Larsson A. Serum amyloid A (SAA) protein and high-sensitivity C-reactive protein (hsCRP) in healthy newborn infants and healthy young through elderly adults. Acta Paediatr. 2005;94(9):1198–1202. | |
Yamada T, Miyake N, Itoh K, Igari J. Further characterization of serum amyloid A4 as a minor acute phase reactant and a possible nutritional marker. Clin Chem Lab Med. 2001;39(1):7–10. | |
Ng PC, Ang IL, Chiu RW, et al. Host-response biomarkers for diagnosis of late-onset septicemia and necrotizing enterocolitis in preterm infants. J Clin Invest. 2010;120(8):2989–3000. | |
Arnon S, Litmanovitz I, Regev RH, Bauer S, Shainkin-Kestenbaum R, Dolfin T. Serum amyloid A: an early and accurate marker of neonatal early-onset sepsis. J Perinatol. 2007;27(5):297–302. | |
Arnon S, Litmanovitz I, Regev R, Lis M, Shainkin-Kestenbaum R, Dolfin T. The prognostic virtue of inflammatory markers during late-onset sepsis in preterm infants. J Perinat Med. 2004;32(2):176–180. | |
Delsesto D, Opal SM. Future perspectives on regulating pro-and anti-inflammatory responses in sepsis. Contrib Microbiol. 2011;17:137–156. | |
Behrendt D, Dembinski J, Heep A, Bartmann P. Lipopolysaccharide binding protein in preterm infants. Arch Dis Child Fetal Neonatal Ed. 2004;89(6):F551–F554. | |
Turner D, Hammerman C, Rudensky B, Schlesinger Y, Goia C, Schimmel MS. Procalcitonin in preterm infants during the first few days of life: introducing an age related nomogram. Arch Dis Child Fetal Neonatal Ed. 2006;91(4):F283–F286. | |
Gerdes JS, Polin RA. Sepsis screen in neonates with evaluation of plasma fibronectin. Pediatr Infect Dis J. 1987;6(5):443–446. | |
Jurges ES, Henderson DC. Inflammatory and immunological markers in preterm infants: correlation with disease. Clin Exp Immunol. 1996;105(3):551–555. | |
Scott PH. Plasma lactoferrin levels in newborn preterm infants: effect of infection. Ann Clin Biochem. 1989;26(Pt 5):412–415. | |
Nupponen I, Andersson S, Järvenpää AL, Kautiainen H, Repo H. Neutrophil CD11b expression and circulating interleukin-8 as diagnostic markers for early-onset neonatal sepsis. Pediatrics. 2001;108(1):E12. | |
Weirich E, Rabin RL, Maldonado Y, et al. Neutrophil CD11b expression as a diagnostic marker for early-onset neonatal infection. J Pediatr. 1998;132(3 Pt 1):445–451. | |
Brown GD. Dectin-1: a signalling non-TLR pattern-recognition receptor. Nat Rev Immunol. 2006;6(1):33–43. | |
Kawai T, Akira S. The roles of TLRs, RLRs and NLRs in pathogen recognition. Int Immunol. 2009;21(4):317–337. | |
Wu YD, Chen LH, Wu XJ, et al. Gram stain-specific-probe-based real-time PCR for diagnosis and discrimination of bacterial neonatal sepsis. J Clin Microbiol. 2008;46(8):2613–2619. | |
Petrakou E, Mouchtouri A, Levi E, Lipsou N, Xanthou M, Fotopoulos S. Interleukin-8 and monocyte chemotactic protein-1 mRNA expression in perinatally infected and asphyxiated preterm neonates. Neonatology. 2007;91(2):107–113. | |
Lawn JE, Cousens S, Zupan J; Lancet Neonatal Survival Steering Team. 4 million neonatal deaths: when? Where? Why? Lancet. 2005;365(9462):891–900. | |
Martinot A, Leclerc F, Cremer R, Leteurtre S, Fourier C, Hue V. Sepsis in neonates and children: definitions, epidemiology, and outcome. Pediatr Emerg Care. 1997;13(4):277–281. | |
Lehrnbecher T, Schrod L, Kraus D, Roos T, Martius J, von Stockhausen HB. Interleukin-6 and soluble interleukin-6 receptor in cord blood in the diagnosis of early onset sepsis in neonates. Acta Paediatr. 1995;84(7):806–808. | |
Procianoy RS, Silveira RC. The role of sample collection timing on interleukin-6 levels in early-onset neonatal sepsis. J Pediatr (Rio J). 2004;80(5):407–410. | |
Resch B, Gusenleitner W, Müller WD. Procalcitonin and interleukin-6 in the diagnosis of early-onset sepsis of the neonate. Acta Paediatr. 2003;92(2):243–245. | |
Chiesa C, Pellegrini G, Panero A, et al. C-reactive protein, interleukin-6, and procalcitonin in the immediate postnatal period: influence of illness severity, risk status, antenatal and perinatal complications, and infection. Clin Chem. 2003;49(1):60–68. | |
Silveira RC, Procianoy RS. Evaluation of interleukin-6, tumour necrosis factor-alpha and interleukin-1beta for early diagnosis of neonatal sepsis. Acta Paediatr. 1999;88(6):647–650. | |
Hodge G, Hodge S, Han P, Haslam R. Multiple leucocyte activation markers to detect neonatal infection. Clin Exp Immunol. 2004;135(1):125–129. | |
Kishimoto T. The biology of interleukin-6. Blood. 1989;74(1):1–10. | |
Küster H, Weiss M, Willeitner AE, et al. Interleukin-1 receptor antagonist and interleukin-6 for early diagnosis of neonatal sepsis 2 days before clinical manifestation. Lancet. 1998;352(9136):1271–1277. | |
Raynor LL, Saucerman JJ, Akinola MO, Lake DE, Moorman JR, Fairchild KD. Cytokine screening identifies NICU patients with Gram-negative bacteremia. Pediatr Res. 2012;71(3):261–266. | |
Franz AR, Sieber S, Pohlandt F, Kron M, Steinbach G. Whole blood interleukin 8 and plasma interleukin 8 levels in newborn infants with suspected bacterial infection. Acta Paediatr. 2004;93(5):648–653. | |
Reinsberg J, Dembinski J, Dorn C, Behrendt D, Bartmann P, van Der Ven H. Determination of total interleukin-8 in whole blood after cell lysis. Clin Chem. 2000;46(9):1387–1394. | |
Orlikowsky TW, Neunhoeffer F, Goelz R, et al. Evaluation of IL-8-concentrations in plasma and lysed EDTA-blood in healthy neonates and those with suspected early onset bacterial infection. Pediatr Res. 2004;56(5):804–809. | |
Fotopoulos S, Mouchtouri A, Xanthou G, Lipsou N, Petrakou E, Xanthou M. Inflammatory chemokine expression in the peripheral blood of neonates with perinatal asphyxia and perinatal or nosocomial infections. Acta Paediatr. 2005;94(6):800–806. | |
Schultz C, Strunk T, Temming P, Matzke N, Härtel C. Reduced IL-10 production and -receptor expression in neonatal T lymphocytes. Acta Paediatr. 2007;96(8):1122–1125. | |
Ng PC, Li K, Wong RP, et al. Proinflammatory and anti-inflammatory cytokine responses in preterm infants with systemic infections. Arch Dis Child Fetal Neonatal Ed. 2003;88(3):F209–F213. | |
Schultz C, Temming P, Bucsky P, Göpel W, Strunk T, Härtel C. Immature anti-inflammatory response in neonates. Clin Exp Immunol. 2004;135(1):130–136. | |
Ng PC, Li G, Chui KM, et al. Neutrophil CD64 is a sensitive diagnostic marker for early-onset neonatal infection. Pediatr Res. 2004;56(5):796–803. | |
Bhandari V, Wang C, Rinder C, Rinder H. Hematologic profile of sepsis in neonates: neutrophil CD64 as a diagnostic marker. Pediatrics. 2008;121(1):129–134. | |
Wang K, Bhandari V, Chepustanova S, et al. Which biomarkers reveal neonatal sepsis? PLoS One. 2013;8(12):e82700. | |
Shozushima T, Takahashi G, Matsumoto N, Kojika M, Okamura Y, Endo S. Usefulness of presepsin (sCD14-ST) measurements as a marker for the diagnosis and severity of sepsis that satisfied diagnostic criteria of systemic inflammatory response syndrome. J Infect Chemother. 2011;17(6):764–769. | |
Graversen JH, Madsen M, Moestrup SK. CD163: a signal receptor scavenging haptoglobin-hemoglobin complexes from plasma. Int J Biochem Cell Biol. 2002;34(4):309–314. | |
Fabriek BO, Dijkstra CD, van den Berg TK. The macrophage scavenger receptor CD163. Immunobiology. 2005;210(2–4):153–160. | |
Prashant A, Vishwanath P, Kulkarni P, et al. Comparative assessment of cytokines and other inflammatory markers for the early diagnosis of neonatal sepsis-a case control study. PLoS One. 2013;8(7):e68426. | |
Sprong T, Peri G, Neeleman C, et al. Pentraxin 3 and C-reactive protein in severe meningococcal disease. Shock. 2009;31(1):28–32. | |
Fiedler U, Reiss Y, Scharpfenecker M, et al. Angiopoietin-2 sensitizes endothelial cells to TNF-alpha and has a crucial role in the induction of inflammation. Nat Med. 2006;12(2):235–239. | |
Mankhambo LA, Banda DL, Jeffers G, et al; IPD Study Group. The role of angiogenic factors in predicting clinical outcome in severe bacterial infection in Malawian children. Crit Care. 2010;14(3):R91. | |
Eugen-Olsen J. suPAR – a future risk marker in bacteremia. J Intern Med. 2011;270(1):29–31. | |
Siahanidou T, Margeli A, Tsirogianni C, et al. Clinical value of plasma soluble urokinase-type plasminogen activator receptor levels in term neonates with infection or sepsis: a prospective study. Mediators Inflamm. 2014;2014:375702. | |
Wu Y, Wang F, Fan X, et al. Accuracy of plasma sTREM-1 for sepsis diagnosis in systemic inflammatory patients: a systematic review and meta-analysis. Crit Care. 2012;16(6):R229. | |
Adly AA, Ismail EA, Andrawes NG, El-Saadany MA. Circulating soluble triggering receptor expressed on myeloid cells-1 (sTREM-1) as diagnostic and prognostic marker in neonatal sepsis. Cytokine. 2014;65(2):184–191. | |
Romano-Keeler J, Wynn JL, Maron JL. Great expectorations: the potential of salivary ‘omic’ approaches in neonatal intensive care. J Perinatol. 2014;34(3):169–173. | |
Castagnola M, Inzitari R, Fanali C, et al. The surprising composition of the salivary proteome of preterm human newborn. Mol Cell Proteomics. 2011;10(1):M110.003467. | |
Hendricks-Muñoz KD, Perez-Perez G, Xu J, Kim Y, Louie M. Maternal antenatal treatments influence initial oral microbial acquisition in preterm infants. Am J Perinatol. 2013;30(1):47–52. | |
Benitz WE. Adjunct laboratory tests in the diagnosis of early-onset neonatal sepsis. Clin Perinatol. 2010;37(2):421–438. | |
Suguna Narasimhulu S, Hendricks-Muñoz KD, Borkowsky W, Mally P. Usefulness of urinary immune biomarkers in the evaluation of neonatal sepsis: a pilot project. Clin Pediatr (Phila). 2013;52(6):520–526. | |
Fanos V, Van den Anker J, Noto A, Mussap M, Atzori L. Metabolomics in neonatology: fact or fiction? Semin Fetal Neonatal Med. 2013;18(1):3–12. | |
Dessì A, Corsello G, Stronati M, et al. New diagnostic possibilities in systemic neonatal infections: metabolomics. Early Hum Dev. 2014; 90 Suppl 1:S19–S21. | |
Fanos V, Caboni P, Corsello G, et al. Urinary (1)H-NMR and GC-MS metabolomics predicts early and late onset neonatal sepsis. Early Hum Dev. 2014;90 Suppl 1:S78–S83. | |
Buhimschi IA, Buhimschi CS. The role of proteomics in the diagnosis of chorioamnionitis and early-onset neonatal sepsis. Clin Perinatol. 2010;37(2):355–374. |
 © 2014 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.
© 2014 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.