Back to Journals » OncoTargets and Therapy » Volume 13
Biological Roles and Mechanisms of Circular RNA in Human Cancers
Received 7 October 2019
Accepted for publication 20 February 2020
Published 9 March 2020 Volume 2020:13 Pages 2067—2092
DOI https://doi.org/10.2147/OTT.S233672
Checked for plagiarism Yes
Review by Single anonymous peer review
Peer reviewer comments 3
Editor who approved publication: Dr Jianmin Xu
Qing Tang, Swei Sunny Hann
Laboratory of Tumor Biology, The Second Clinical College of Guangzhou University of Chinese Medicine, Guangzhou 510120, Guangdong Province, People’s Republic of China
Correspondence: Swei Sunny Hann
Laboratory of Tumor Biology, The Second Clinical College of Guangzhou University of Chinese Medicine, No. 111, Dade Road, Guangzhou 510120, Guangdong Province, People’s Republic of China
Tel +86 20-39318472
Email [email protected]
Abstract: Circular RNA (circRNA) is an intriguing class of RNA with covalently closed-loop structure and is highly stable and conservative. As new members of the ncRNAs, the function, mechanism, potential diagnostic biomarker, and therapeutic target have raised increased attention. Most circRNAs are presented with characteristics of abundance, stability, conservatism, and often exhibiting tissue/developmental-stage-specific manner. Over 30,000 circRNAs have been identified with their unique structures to maintain stability more easily than linear RNAs. An increased numbers of circRNAs are dysregulated and involved in several biological processes of malignance, such as tumorigenesis, growth, invasion, metastasis, apoptosis, and vascularization. Emerging evidence suggests that circRNAs play important roles by acting as miRNA sponge or protein scaffolding, autophagy regulators, and interacting with RNA-binding protein (RBP), which may potentially serve as a novel promising biomarker for prevention, diagnosis and therapeutic target for treatment of human cancer with great significance either in scientific research or clinic arena. This review introduces concept, major features of circRNAs, and mainly describes the major biological functions and clinical relevance of circRNAs, as well as expressions and regulatory mechanisms in various types of human cancer, including pathogenesis, mode of action, potential target, signaling regulatory pathways, drug resistance, and therapeutic biomarkers. All of which provide evidence for the potential utilities of circRNAs in the diagnosis and treatment of cancer.
Keywords: circRNA, cancer, miRNA sponge, protein scaffolding, gene splicing and transcription, biomarker, therapeutic target
Introduction
CircRNA has a covalent closed-loop structure and is highly stable and conservative special RNA, which exists widely in various tissues and organs with variable expression levels, and broadly participates in the occurrence and development of diseases including cancer in various ways. Currently, with the development of deep RNA sequencing (RNA-seq) technologies and novel bioinformatic approaches, over 30,000 circRNAs have been identified with the unique structures and have attracted increasing attention given its high biological and functional interest. CircRNAs are resistant to exonuclease RNase R and maintain stability more easily than linear RNAs. Increasing evidences have shown that part of circRNAs containing miRNA binding sites may act as sponge to interact with miRNA and regulate its biological functions, thereby affecting the expression and function of its downstream target genes.1,2 CircRNAs with binding sites of enzymes and substrates may act as proteins scaffolding to mediate the interaction of protein-protein. For example, circFoxo3, containing the binding sites of mouse double minute 2 (MDM2) and p53, was involved in the interaction between MDM2 and p53 by functioning as a protein scaffold.3,4 Moreover, circRNA also play important roles in regulating biological functions of cancers through involvement in gene alternative splicing, transcription and translation, cell autophagy and interacting with RNA-binding proteins (RBPs).4–6 In this review, we describe the major biological functions and clinical relevance of circRNAs, as well as its expression and regulatory mechanisms in various types of human cancer including pathogenesis, mode of action, potential target, signaling regulatory pathways, therapeutic biomarkers, drug resistance and clinical application.
Major Features of circRNAs
CircRNAs, first identified in RNA viruses in 19767 and once considered “splicing noise” in organisms, have recently become interest research focus as the results of improvements in high-throughput sequencing technology and bioinformatics, circRNAs have become a research hotspot.4 As a new type of RNA molecule, circRNAs are single-stranded circularized RNA with no 5′ caps and 3′ poly(A) tails; and commonly generated from the precursor mRNA (pre-mRNA) by a process called back-splicing in which an upstream acceptor site is joined with a donor site.8 Most circRNAs are evolutionarily conserved across species.9 The vast majorities of circRNAs are often located in the cytoplasm, which are derived from known protein-coding genes containing one or several exons that are toward the 5′ of the gene and are flanked by longer introns. The long introns containing flank regions that will become circRNAs usually contain specific sequences which induce circRNA formation either by complementarity and/or by binding to circRNA-promoting factors, and are generally expressed in cell type-specific and tissue-specific manners.10,11 Based on the different structures, and cycling mechanisms, circRNA molecules are divided into four categories: exonic circRNAs (ecRNAs), intronic circRNAs (ciRNAs), exon-intron circRNAs (eIciRNAs), and intergenic circRNAs. Unlike linear RNAs, circRNAs are stable and resistant to exonucleases (including RNase R) due to the lack of a poly(A) tail and have longer median half-life than that of their linear mRNAs due to the lack of free 3′ or 5′ ends, which makes them resistant to regular mechanisms of linear RNA decay.10–12
Functions of cirRNAs
Acting as miRNA Sponges
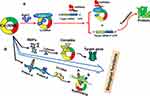
Accumulating evidence has revealed that numerous circRNAs regulate gene expression by functioning as miRNA sponge molecules. Compared with other competing endogenous RNAs (ceRNAs) (such as lncRNA or pseudogenes), circRNAs exhibit a greater preference to bind to miRNAs and are called “super sponges.”13 Many circRNAs contain miRNA response elements and binding sites, therefore, acting as “miRNA sponge” is the most important function and mechanism of regulating the growth and progression of human cancer (Figure 1). For example, circ_0004771 was identified to restrain cell proliferation and accelerate cell apoptosis in breast cancer cells. circ_0004771 acted as a miRNA sponge to decrease expression of miR-653, this in turn targeted mesenchymal marker zinc finger E-box-binding homeobox 2 (ZEB2) gene expression by binding to its 3ʹ-untranslated region (3ʹUTR) of ZEB2 mRNA.14 In addition, circAGFG1 was correlated with advanced clinical stage, poor prognosis and pathological grade of triple-negative breast cancer patients, mechanistically, circAGFG1 might directly sponge miR-195-5p, which targeted and repressed cyclin E1 expression.15
Regulating Gene Splicing, Transcription and Translation
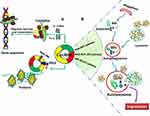
Although most circRNA is located in cytoplasm, some are still existed in nucleus. The part of circRNA retained in nucleus acted as transcriptional or splicing regulators to interfere with gene expression and involved in alternative splicing and transcription process (Figure 2). For example, circPOK, derived from the Zbtb7a gene in tumor cells, was involved in gene transcription by encoding the pokemon transcription factor, thereby regulating the pro-proliferative and pro-angiogenic factors through activating the interleukin enhancer-binding factor 2 and 3 complex (ILF2/3) complex.16 Moreover, circITGA7 was found to increase the transcription of its host gene integrin alpha 7 (ITGA7) by inhibiting a transcription factor RAS-responsive element-binding protein 1 (RREB1) through Ras pathway.17 In addition, circ-UBR5 was significantly decreased in non-small cell lung cancer (NSCLC) tissues and associated with tumor differentiation. Mechanically, circ-UBR5 might be involved in RNA splicing regulatory process through binding to splicing regulatory factor QKI, NOVA alternative splicing regulator 1 (NOVA1) and U1 small nuclear RNA (snRNA) in the NSCLC cells.18 Furthermore, circRNA containing AUG sites and open reading frame (ORF) could be driven by the internal ribosome entry site (IRES) and translated into the functional protein (Figure 2). However, the relevance of role of circRNA translational function needs to be further explored in terms of mediating occurrence and progression of human cancer.3,4 Overall, gene alternative splicing, transcription and translation are important processes to exert biological functions in cancers, while circRNA plays a crucial role in these processes (Figure 2).
Acting as Autophagy Regulator
Autophagy is a highly conserved and successive self-degradative process that plays an important role in cellular stress responses and survival, which often occurs during tumorigenesis, progression, metastasis and chemotherapy leading to drug resistance in the treatment of cancer.19,20 Emerging evidence showed that circRNA was involved in the tumor autophagy affecting occurrence and progression of human cancer (Figure 2). For example, circHIPK3 was found to act as an autophagy regulator in STK11 mutant lung cancer cells. The results showed that missing expression of circHIPK3 could induce cell autophagy through regulating the miR124-3p/signal transducer and activator of transcription 3 (STAT3)/protein kinase AMP-activated catalytic subunit alpha 2 (PRKAA)/AMP-activated protein kinase (AMPKα) signaling regulatory pathways. Moreover, the ratio of circHIPK3 to linHIPK3 (liner HIPK3) reflected the level of autophagy in cancer cells.21 Moreover, circ_104075 was correlated with apoptosis and autophagy of glioma cells. Overexpression of circ_104075 neutralized the inhibitory effects of matrine on proliferation and promoted the cell autophagy in glioma U251 cells.22 Nevertheless, the research of circRNA in the tumor autophagy process is still in the initial stage, the true role and function, potential mechanism underlying this required to be explored in the future.
Interacting with RNA-Binding Proteins (RBPs) and Acting as Protein Scaffolding
RNA-binding proteins (RBPs) are a group of proteins widely involved in gene transcription and translation. Increasing evidences suggested that circRNA could bind RBPs and affect its function23 (Figure 1). CircRNA could also sequester, store and sort RBPs and thus control the intracellular localization.24,25 Conversely, RBPs could also regulate the function and expression level of circRNA. RNA-binding protein 3 (RBM3) dynamically regulated the production of SCD-circRNA2, rooted in the 3ʹ-UTR of the stearoyl-CoA desaturase (SCD) gene, thereby ultimately regulating proliferation in hepatocellular carcinoma cells.26 Recent studies have shown that RBP quaking could also modulate the formation of circRNA through forming RNA-protein complexes (RPCs).27 Moreover, RNA-binding motif protein 20 (RBM20) was associated with the formation of subset of circRNAs and formed the class of RBM20-dependent circRNAs.28 Thus, circRNA and RPBs are closely associated with each other in the occurrence and development of cancer. Overall, circRNAs are abnormally expressed and related to the occurrence and progression of human cancer via influencing cell growth, proliferation, migration, invasion, and other pathological processes (Table 1). In addition, circRNAs were also correlated with clinical relevance, such as TNM stage, lymph node metastasis, differentiation, tumor size and overall survival (Table 2). All of these provided evidences for the potential biomarker and therapeutic target in the diagnosis and treatment of human cancers. In addition to interacting with RBPs, another function of circRNA is its interaction with protein. It can function as protein sponges by adsorbing one or more proteins via the binding sites, thereby directly mediating the interaction between proteins by acting as protein scaffolding, thus regulating gene expression (Figure 1). For example, cyclin-dependent kinase 2 (CDK2) and p21 proteins were associated with cell cycle regulation in tumor. CircFOXO3 could mediate the formation of circFOXO3-p21-CDK2 ternary complex by serving as scaffolding, which blocked the function of CDK2, thereby affecting the cell cycle progression of cancer.29
 |  |  |
Table 1 Expression and Functional Characterization of circRNAs in Cancers |
 |
Table 2 The Roles in Clinical Relevance and Prognosis of circRNAs in Human Cancers |
circRNAs in Human Cancers
Lung Cancer
Lung cancer is the leading cause of cancer death and accounts for approximately 13% of all cancer cases and 23% of all cancer-related deaths worldwide.30 Increasing evidence suggested that circRNA might participate in the cell proliferation, migration and invasion of lung cancer, and serve as an important diagnostic marker for the treatment of lung cancer.31–37 For example, circNOL10 increased the expression of transcription factor sex comb on midleg-like 1 (SCML1) by inhibiting ubiquitination and regulating the humanin (HN) polypeptide family through affecting multiple signaling pathways. This ultimately inhibited proliferation and induced cell cycle arrest in lung cancer cells.38 Recent study revealed that circPTPRA suppressed the epithelial-mesenchymal transition (EMT) and metastasis of NSCLC cells by sponging miR-96-5p, thereby regulating the expression of downstream tumor suppressor ras association domain family 8 (RASSF8) gene.39 In line with this, other circRNAs, such as circP4HB, circMTO1, circ_0026134, circ-FOXM1, circ_003645, circ_0006427, circABCB10, circ-BANP, circFADS2, circ_103809, circMAN2B2, circ_0012673 and circ_0020123 showed similar roles that were served as sponge of miRNAs to regulate the occurrence and development of lung cancer.40–54 Emerging evidences demonstrated that cancer-associated chromosomal translocations and encoding fusion gene could generate circRNA, contributing to tumorigenesis.55 For instance, SLC34A2-ROS1 fusion gene could produce circRNA F-circSR, which promoted cell migration of NSCLC cells.56 CircRNA F-circEA-2a, and circRNAF-circEA deriving from oncogenic fusion gene echinoderm microtubule-associated protein-like 4-anaplastic lymphoma kinase (EML4-ALK), enhanced the cell migration and invasion in NSCLC, and might act as a novel liquid biopsy biomarker in NSCLC.57,58 Moreover, some circRNAs regulated cell pathogenesis, development and prognosis of lung cancer by affecting the cell cycle-related signaling pathway. Circ_0007766 induced the proliferation and migration of lung cancer cells through regulating and modulating the cyclin D1/cyclin E1/CDK4 regulatory axis.59 Also, others were involved in the influencing the expression of apoptosis-related protein, circVANGL1 contributed to proliferation, migration, invasion, and progression of NSCLC via competing endogenous RNA (ceRNA), becoming a sink for miR-195 thereby modulating the expression of Bcl-2 in NSCLC cells.60 Overall, circRNA have been involved in the pathogenesis, development and prognosis of lung cancer, and provided potential biomarker and prospective targets for lung cancer treatment.
Breast Cancer
Breast cancer (BC) is one of the leading causes of cancer-related mortality and the second most common cancer in females. Recently, an increasing number of circRNAs have been identified and correlated with clinical-pathological characteristics in the progression of BC. CircRNAs also participated in the biological functions and progression of BC, such as tumorigenesis, proliferation, apoptosis, cell cycle, vascularization, invasion, migration and metastasis.61–65 For example, circ_001569 was identified to be associated with clinical-pathological features and prognosis in BC patients, and knockdown of circ_001569 remarkably inhibited the activation of PI3-K/Akt signal pathway,66 Moreover, circRNA-MTO1 (also known as circRNA-007874) could significantly suppress cell viability and reverse monastrol resistance in BC. Mechanistic studies showed that circRNA-MTO1 reduced the Eg5 protein expression but not mRNA level through preventing TNFα receptor-associated factor (TRAF) 4 from activating Eg5 translation.67 CircRNAs were also widely involved in the occurrence and development of BC by acting as “miRNA sponge.” For instance, circTADA2A-E6 could sponge miR-203a-3p to reduce the expression of miR-203a-3p; thereby restoring the expression of suppressor of cytokine signaling 3 (SOCS3), a target gene of miR-203a-3p, resulting in suppressed the progression and metastasis of BC.68 In addition, circKIF4A promoted proliferation and migration of triple-negative breast cancer (TNBC), cells by directly sponging miR-375 to relieve the suppression of KIF4A target gene.69 Moreover, more circRNAs, such as circPLK1, circ-UBE2D2, circBMPR2, circ_0025202, circ_0103552, circ_0072309, circ_001783, involved in the occurrence and development of BC by acting as miRNA sponge have been reported.70–76 Together, circRNA is broadly expressed in BC tissues and cells with variable levels associated with clinical pathogenesis of BC, and could be used as a potential biomarker and therapeutic target in the treatment of BC.
Colorectal Cancer
Colorectal cancer (CRC) is the third most common malignant cancer and the fourth leading cause of cancer death around the world.77 In recent years, circRNA is considered as an important regulator for the tumorigenesis and progression in CRC. CircRNA was reported to be associated with the development of CRC, and acted as potential biomarkers and therapeutic target for the diagnosis and treatment of CRC.78 Recently, circRNA expression profiles in CRC patients were performed through high-throughput RNA sequencing (RNA-seq) and total of 448 significantly dysregulated circRNAs were identified. Among those, 394 were up-regulated and 54 were down-regulated, all of which were involved in cell proliferation, migration, invasion and apoptosis in CRC.79 circRNA-0000523 activated the activity of Wnt/β-catenin signaling pathway to regulate the proliferation and apoptosis of CRC cells by sponging miR-31.80 Also, circ_0009361 acted as the sponge of miR-582 to enhance the expression of adenomatous polyposis coli 2 (APC2) and blocked the Wnt/β-catenin signaling; resulting in suppressing cell growth and metastasis of CRC.81 Furthermore, many other circRNAs, such as circVAPA, circ_0136666, circRNA_103809, circRNA_100290, circ_0026344, circHIPK3, and circ_001569 were also acted as sponge of miRNAs to regulate the tumorigenesis and progression of CRC have been reported.82–88 In addition, circRNA was also associated with chemoradiation resistance (CRR) of CRC. One study showed that among 71 circRNAs expressed in 5-FU chemo-resistant CRC cells by microarray analysis, 47 circRNAs were increased and 24 circRNAs were decreased significantly. The study provided a useful database for further understanding of CRR and presented potential targets to reverse CRR in CRC.89 To this end, circRNA play an important role in the occurrence and development of CRC, and could also be involved in diagnosis and treatment of CRC.
Esophageal Squamous Cell Carcinoma
Esophageal squamous cell carcinoma (ESCC) is the globally predominant aggressive malignancies of the gastrointestinal tract.90,91 CircRNAs have been studied to serve as biomarkers of diagnosis and treatment for ESCC and been involved in the regulation of the cell proliferation, migration, invasion and metastasis of ESCC.92,93 Increasing number of studies have shown that circRNA modulated the occurrence and progression of ESCC via acting as “miRNA sponge.” For example, circ-TTC17, deriving from tetratricopeptide repeat domain 17 (TTC17) gene, showed to promote proliferation and migration of ESCC cells by serving as “miRNA sponge.” The bioinformatics analysis observed a network of circ-TTC17 with its targeted miRNA interactions and corresponding mRNAs, and found that a total of 20 miRNAs were predicted to have binding sites with circ_TTC17 suggesting that circ-TTC17 might regulate progress of ESCC by acting as a sponge for miRNAs.94 Moreover, circ-PRKCI promoted cell migration and proliferation through enhancing the expression of AKT serine/threonine kinase 3 by sponging miR-3680-3p in ESCC cells.95 Circ_0006168 could regulate the mammalian target of rapamycin (mTOR) expression by sponging miR-100 to facilitate ESCC cell proliferation, migration and invasion. Thus, circ_0006168 has been considered to be a promising prognostic biomarker and effective therapeutic target for ESCC patients.96 In line with this, other studies also found that circRNAs, such as circ-DLG1, circular RNA ciRS-7, circ_0000337, could interact with miRNAs by acting as sponge or competing endogenous RNA in the progression of ESCC.97–99 Together, these findings suggested that circRNA were involved in the carcinogenesis and progression of ESCC, and could be a promising diagnostic biomarker and potential therapeutic target in patients with ESCC.
Gastric Cancer
Gastric cancer (GC) is one of the most common malignant tumors in the digestive system and most GC is found at an advanced stage, which poses a great challenge to the treatment of this malignancy.100,101 An increasing number of studies have suggested that circRNA play critical roles and act as potential biomarker for the diagnosis and treatment of GC.102–104 However, the functions and underlying mechanisms of circRNAs in GC remain to be further studied. Likewise, “miRNA sponge” is also the main mechanism of circRNAs to participate in the progression of GC. For instance, circYAP1 was identified to suppress cell proliferation and invasion of GC by sponging miR-367-5p, then inhibited the expression of p27Kip1.105 Also, circNRIP1 was found to sponge miR-149-5p to further regulate AKT/mTOR signaling axis and effected the cell proliferation, migration and invasion in GC.106 Likewise, several other circRNAs such as circ_00610, circ-NOTCH1, circ-DCAF6, circ_0008035, circ_0001368, circPSMC3, cir-NF1, circ-SFMBT2, circFAT1(e2), circPDSS1, circ_0027599, and circ-0081143, could also serve as “miRNA sponge” to modulate other gene expressions in GC.107–118
In addition, circRNA could also perform the biofunctions by serving as protein scaffolding in the progression pathogenesis of GC. For example, circ-DONSON was identified to promote cell proliferation, migration and invasion while inhibiting cell apoptosis in GC. Mechanistically, circ-DONSON could significantly recruit the NURF complex by acting as a protein scaffolding, to regulate a transcription factor Sex-determining region Y (SRY)-related high-mobility group box 4 (SOX4) promoter activity and stimulate transcription.119 Moreover, circAGO2, deriving from Argonaute 2 (AGO2), the core component of miRNA-induced silencing complex, physically interacted with human antigen R (HuR) protein to activate its activities and enrichment on the 3ʹ-UTR of target gene, which significantly reduced the binding activation of AGO2 and thereby overcoming the effect of AGO2/miRNA-mediated gene silencing that was associated with the progression of GC.120 Overall, circRNA played an important role in the tumorigenesis and progression through multiple mechanisms, and unveiled significant potential for the prevention and treatment of GC.
Hepatocellular Carcinoma
Hepatocellular carcinoma (HCC) is one of the most common cancers and the second cause of cancer mortality worldwide; nearly two-thirds of all patients with HCC are diagnosed at advanced stages.121–123 Growing evidences indicated that the expression alterations of circRNAs have a significant impact on biological characteristics of HCC. Recent study showed that circ-ZEB1.33 was a potential biomarker for the prognosis of HCC patients. They found that the expression of circ-ZEB1.33 was related to different TMN stages in HCC patients. Moreover, there were significant interactions between circ-ZEB1.33 and miR-200a-3p, as well as cyclin-dependent kinase 6 (CDK6) indicating that circ-ZEB1.33-miR-200a-3p-CDK6 regulatory pathway played a critical role in the progression of HCC.124 Increasing number of studies have focused circRNA on their functions as efficient miRNA sponges in HCC as well. For example, circHIAT1 could act the miR-3171 sponge to further regulate the expression of PTEN that was a target of miR-3171, thereby inhibiting cell growth of HCC.125 Likewise, cirRNAs, such as circ_0101432 sponge miR-1258/miR-622,126 circZNF652 sponge miR-203/miR-502-5p,127 circSETD3 sponge miR-421,128 circ_103809 sponge miR-620,129 circ_0000267 sponge miR-646,130 circ_0008450 sponge miR-548p,131 circDAMTS13 sponge miR-484,132 circ_0078710 sponge miR-31,133 circ_101280 sponge miR-375,134 circSLC3A2 sponge miR-490-3p,135 and circTRIM33-12 sponge miR-191136 have also been shown. All of these circRNAs were involved in the occurrence and progression of HCC by acting as “miRNA sponge.” In addition, some circRNAs could serve as immune-associated biomarker to regulate the tumourigenesis and metastasis. For example, circARSP91 regulated the progression of HCC through enhancing the activation of natural killer (NK) cells and increasing the susceptibility of HCC cells to NK cell cytotoxicity associated with cell immune surveillance.137 Overall, circRNAs played key roles in tumorigenesis and development, epigenetic regulation, drug resistance, and could be considered as immune-associated biomarker and therapeutic target in HCC.138–144 Regardless, more in-depth study of circRNA biofunctions in HCC is greatly desired.
Bladder Cancer
Bladder cancer is the 9th most common cancer around the world with an estimated 165,000 deaths per year.145,146 Increased number of studies indicated that circRNAs were involved in the occurrence and development of bladder cancer.147–150 For instance, circUVRAG derived from the exon from the UV radiation resistance-associated gene (UVRAG) was highly increased in bladder cancer cells, and regulated the aggressive biological phenotype through targeting the miR-223/fibroblast growth factor receptor 2 (FGFR2) signaling pathway. Downregulation of circUVRAG promoted miR-223, but suppressed FGFR2 expressions.151 Circ-BPTF was also increased in bladder cancer tissues compared with noncancerous ones, and promoted the progression and recurrence of bladder cancer by regulating the miR-31-5p/RAB27A signaling pathway.152 Moreover, circLPAR1 inhibited the activity of miR-762 by directly binding to miR-762 thereby regulating invasion and metastasis of muscle-invasive bladder cancer cells.153 circRNA-cTFRC regulated cell invasion and proliferation through acting as a ceRNA for miR-107 to affect the expression of TFRC expression in bladder cancer cells.154 CircUBXN7 showed to inhibit the proliferation and invasion of bladder cancer cells through binding to miR-1247-3p to elevate the expression of β-1, 4-Galactosyltransferase III (B4GALT3), which is the direct target gene of miR-1247-3p.155 Thus, circRNAs were involved in regulation of bladder cancer progression by serving as ceRNA. Interestingly, circRNAs could be also involved in the bladder oncogenesis and metastasis through regulating self-renewal function of cancer stem cells (CSCs). For example, knockdown of circGprc5a, a circRNA with peptide-coding potential and functions through a peptide-dependent manner, impaired the self-renewal and metastasis of bladder CSCs.156 Together, data demonstrated that circRNA may provide a potential biomarker and therapeutic target for the management of bladder cancer.
Pancreatic Cancer
Pancreatic cancer is one of the most common malignancy and the fourth leading cause of cancer-related death worldwide with low 5-year overall survival rate of less than 7%.157,158 An increasing number of studies showed that circRNAs were associated with the occurrence and progression of pancreatic cancer.157,159 Circ-LDLRAD3 was reported to be increased in both cells and tumor tissues. High expression of circ-LDLRAD3 was significantly associated with venous invasion, lymphatic invasion and metastasis, indicating that circ-LDLRAD3 might be a critical biomarker in the diagnosis and treatment of pancreatic cancer.160 Circ_0006215 was also regulated the progression of pancreatic cancer cells through the circ_0006215/miR-378a-3p/serpina family A member 4 (SERPINA4) signaling pathway.161 Moreover, circRNA_100782 regulated the miR-124/IL6/STAT3 pathway. Knockdown of circRNA_100782 significantly modulated miR-124 expression, and reduced miR-124 target genes interleukin-6 receptor (IL-6R) and STAT3 expressions in pancreatic cancer cells.162 Similarly, knockdown of circZMYM2 significantly repressed the tumorigenesis through sponging miR-335-5p, followed by affecting the expression of histone lysine demethylases jumonji domain-containing 2c (JMJD2C), which is the target gene of miR-335-3p in pancreatic cancer cells.163 In line with this, the results of bioinformatics analysis showed that miR-26b-3p, miR-125a-3p, miR-330-5p and miR-382-5p had binding sites for circRHOT1 suggesting that circRHOT1 might act as “miRNA sponge” to promote proliferation and invasion in pancreatic cancer cells.164 In addition, exosomal circRNAs also contributed to tumor invasion and metastasis. For example, exosomal circ-PDE8A stimulated cell growth and invasive ability by acting as ceRNA to sponge miR-338 and regulated miR-338/metastasis-associated in colon cancer-1 (MACC1)/MET or hepatocyte growth factor (HGF) receptor regulatory axis in pancreatic cancer cells.165 Moreover, exosomal circ-IRAS significantly promoted invasion and metastasis of pancreatic cancer cells by down-regulating miR-122 and ZO-1 levels, and up-regulating RhoA and RhoA-GTP levels, F-actin expression, regulating endothelial monolayer permeability.166 We believed that circRNAs could provide a new promising biomarker for diagnosis and therapeutic target for the treatment of pancreatic cancer.
Papillary Thyroid Carcinoma
Thyroid cancer is one of the most common malignant endocrine tumors, with an incidence of 1–2% of all types of cancer, Despite of a good and overall prognosis, papillary thyroid cancer (PTC), which accounts for 75% of thyroid cancer, could still affect the quality of life of PTC patients.167,168 A large number of circRNAs showed promising as potential prognostic biomarkers for the PTC patients, and played a critical role in the pathogenesis and progression of PTC.169–171 For example, knockdown of circBACH2 inhibited the cell proliferation, migration and invasion of PTC cells in vitro and suppressed the growth of PTC xenografts in vivo. Mechanistically, circBACH2 directly interacted with miR-139-5p and relieved inhibition of its target gene LIM-domain only protein 4 (LMO4). Therefore, circBACH2/miR-139-5p/LMO4 regulatory axis could be a promising treatment strategy for PTC patients.172 Likewise, low expression of circRAPGEF5 plays an important role in suppressing the aggressive biological behaviors of PTC by sponging miR-198, this subsequently downregulated the expression of fibroblast growth factor receptor 1 (FGFR1), a target gene of miR-198.173 Moreover, overexpression of circ_0025033 promoted proliferation and invasion of PTC by directly sponging miR-1231 and miR-1304, therefore, circ_0025033/miR-1231/miR-1304 signaling pathway was considered to be a new regulatory mechanism in PTC initiation and progression.174 Recent study found that knockdown of circRNA_102171 inhibited PTC progression. CircRNA_102171 could interact with catenin beta interacting protein 1 (CTNNBIP1) and block its association with the β-catenin/TCF complex to further activate the activity of Wnt/β-catenin pathway in PTC cells.175 In a similar way, circ-ITCH was also correlative with Wnt/β-catenin pathway. Bioinformatics analysis and luciferase reporter assays showed that circ-ITCH could sponge miR-22-3p to increase the expression of CBL, an E3 ligase of nuclear β-catenin. This led to suppress activation of the Wnt/β-catenin pathway and consequently inhibited the progression of PTC.176 Moreover, circZFR was negative correlated with clinical severity of PTC patients. Knockdown of circZFR significantly activated C8orf4 (chromosome 8 open reading frame 4), an activator of Wnt signaling pathway via sponging miR-1261 thereby inhibiting proliferation, migration and invasion of PTC cells.177 In addition, circ_0058124 acted as a ceRNA to directly regulate the expression of miRNA-218-5p and its target gene NUMB, and consequently inhibited the activation of the NOTCH3/GATA zinc finger domain-containing 2A (GATAD2A) signaling axis. This let to promote cell proliferation, tumorigenicity, invasion, and metastasis of PTC, thus highlighting a novel therapeutic target for intervening PTC.178 Growing evidences have also shown that circRNA not only play an important role in carcinogenesis and development of PTC, but also have great diagnostic and prognostic value for PTC. One study showed the expression of circRNAs in PTC tissues and adjacent noncancerous tissues, and assess the diagnostic value of circRNAs through analyzing the correlation between cirRNAs and aggressive clinic-pathologic characteristics of PTC indicated that circ_0137287 had a potential diagnostic value in predicting severity of malignancy, extra thyroidal extension and lymph node metastasis, and may act as a novel biomarker for PTC.179 Taken together, circRNA might play an important role in the progression and pathogenesis and be considered as potential biomarkers of PTC.
Osteosarcoma
Osteosarcoma is a malignant bone tumor that has the highest morbidity in adolescent and childhood with 60% of patient aged under 25 years; however, there is a second peak of incidence in later life with 30% of patients being over 40 years of age.180–182 Several studies have indicated the correlations between circRNA and occurrence and progression of osteosarcoma.183–185 CircFAT1, deriving from exon 2 of FAT atypical cadherin 1 (FAT1) gene, significantly inhibited the cell migration, invasion and tumorigenesis of osteosarcoma by sponging miR-375 to enhance the expression of yes-associated protein 1 (YAP1) protein.186 Similarly, knockdown of circCDR1as significantly suppressed tumor growth of osteosarcoma through directly targeting miR-7 and subsequently reduced EGFR, Cyclin E1 (CCNE1), phosphatidylinositol-4, 5-bisphosphate 3-kinase catalytic subunit delta (PI3KCD) and RAF1 proto-oncogene serine/threonine-protein kinase (RAF1) expressions.187 As we know, transcription factor CREB3 is a driver gene in osteosarcoma. CircTADA2A could upregulate the expression of CREB3 by sponging miR-203-3p thereby significantly promoting the progression and metastasis in osteosarcoma cells.188 In addition, circ_0081001 was highly expressed in the osteosarcoma tissues and cells, which may be a potential biomarker for diagnosis and therapeutic target of osteosarcoma. Moreover, serum circ_0081001 might be a better diagnostic and independent prognostic factor than alkaline phosphatase (ALP) and lactate dehydrogenase (LDH) in osteosarcoma patients.189 More importantly, some of circRNAs are closely related to chem-resistance of osteosarcoma. For example, upregulation of circ_001569 not only promoted cell proliferation, but also enhanced the cisplatin resistance of osteosarcoma cells through activating the Wnt/β-catenin signaling pathway.190 Thus, the abnormal expression of circRNAs is an important factor in regulating the occurrence and development of osteosarcoma. We believed that circRNA could be served as the valuable biomarker for the prevention, diagnosis and treatment of osteosarcoma in the future.
Glioblastoma
Glioblastoma is the most common and fatal primary malignant brain tumor. The published data have shown a strong association between circRNA and glioblastoma. The results from microarray analysis showed that most circRNAs were dysregulated in glioblastoma. circ_0001946 was downregulated in glioblastoma and overexpression of circ_0001946 significantly inhibited cell proliferation, migration, invasion and induced apoptosis through upregulating the expression of cerebellar degeneration-related auto-antigen 1(CDR1) by suppressing miR-671-5p expression in glioblastoma cell.191 Obviously, circRNA could also act as “miRNA sponge” to regulate glioblastoma progression. CircMMP9 could regulate the expression of eukaryotic initiation factor 4A3 (eIF4A3) to further accelerate proliferation, migration and invasion via sponging miR-124 in glioblastoma cells.192 Moreover, circNT5E, deriving from ecto-5ʹ-nucleotidase (NT5E) gene and regulating by adenosine deaminase, RNA-specific B2 (ADARB2), also acted as the sponge of miR-422a and reduced its expression, thereby promoting glioblastoma tumorigenesis.193 Furthermore, circ_0029426 served as the sponge of miR-197 to promote cell proliferation, migration and invasion, inhibited cell apoptosis of glioblastoma cells as well.194 Taken together, acting as “miRNA sponge” is one of the most important functions and mechanisms for circRNA to modulation of miRNA and downstream target gene. Recently, comparative results of circRNAs expression profiles showed that 254 circRNAs were up-regulated and 361 circRNAs were down-regulated in IDH-wt glioblastoma compared with the adjacent normal brain tissues. Gene Ontology (GO) analysis revealed that differentially expressions of circRNAs were correlated with cell division, DNA damage repair, cytoskeleton, and protein ubiquitination.195 Their results suggested that differential expressions of circRNAs might serve as biomarkers for prognosis and treatment targets for IDH-wt glioblastoma.195 In addition, there was evidence that endogenous circRNA was involved in gene translation. CircRNA containing an ORF could translate a functional protein through driving by Internal Ribosome Entry Site (IRES) elements.196 For example, circ-SHPRH could produce a 17 kDa protein, the circular form of the SNF2 histone linker PHD RING helicase (SHPRH) gene encoded a novel protein that we termed SHPRH-146aa, which was a tumor suppressor in human glioblastoma. Excessive expressed SHPRH-146aa reduced malignant behavior and tumorigenicity in U251 and U373 glioblastoma cells by protecting SHPRH from degradation through the ubiquitin proteasome. An increased patient survival was observed with elevated levels of SHPRH-146aa in glioblastoma patients.196,197 Collectively, these results showed that circRNA was a group of important regulatory factor and related to the occurrence and progression of glioblastoma. Thus, circRNA could serve as potential and valuable biomarker for diagnosis and treatment for glioblastoma patients in the future.
Ovarian Cancer
Ovarian cancer is the leading cause of death from gynecological malignancies worldwide. The overall 5-year survival rate was particularly low for patients with advanced stages. In recent years, numerous studies focused on differentially expressed circRNAs and their function in this malignancy indicating that circRNAs may act as potentially novel biomarkers or therapeutic agents in this cancer type.198 Increasing number of circRNAs have been reported to be involved in the progression and tumorigenesis of ovarian cancer. One study revealed that circPUM1 promoted cell proliferation, migration, invasion, and metastasis through increased the expression of nuclear factor kappa B (NF-κB) and matrix metallopeptidase 2 (MMP2) by sponging miR-615-5p and miR-6753-5p in ovarian cancer cells.199 CircWHSC1 increased proliferation, migration and invasion, and inhibited apoptosis by sponging miR-145 and miR-1182 thereby increasing the expression of downstream targets mucin 1 (MUC1) and human telomerase reverse transcriptase (hTERT) in ovarian cancer cells.200 Similarly, scores of circRNAs performed their biological functions through acting as sponge of miRNAs in ovarian cancer, such as circUBAP2 sponge miR-144, circCDR1 sponge miR-135b-5p, circ-CSPP1 sponge miR-1236-3p, circ_0051240 sponge miR-637, circEPSTI1 sponge miR-942, circ-ITCH sponge miR-10a, circ_0061140 sponge miR-370, and circGFRA1 sponge miR-449a.201–208 In addition, study showed that circSETDB1 expression levels were closely associated with advanced clinical stage and lymph node metastasis of high-grade serous ovarian cancer patients. Patients with higher levels of circSETDB1 had a shorter progression-free survival time. Thus, circSETDB1 might be a promising biomarker for the treatment and relapse in high-grade serous ovarian cancer.209 Moreover, upregulation of circ-FAM53B accelerated the proliferation, migration, and invasion of ovarian cancer via regulating the miR-646/vesicle associated membrane protein 2 (VAMP2) and miR-647/mouse double minute 2 (MDM2) signaling regulatory pathways.210 Also, circPLEKHM3 could inhibit cell growth, migration and EMT via miR-9/BRCA1/DnaJ/Hsp40 homolog, subfamily B, member 6 (DNAJB6)/Kruppel-like factor 4 (KLF4)/AKT1 regulatory axis in ovarian cancer suggesting that circPLEKHM3 might act as a prognostic indicator and therapeutic target in ovarian cancer patients.211 In addition, circ-SMAD7 enhanced cell metastasis, proliferation and progression of ovarian cancer via suppressing the expression of Krüppel-like factor 6.212 Overall, the differentially expressed circRNAs may participate in the pathogenesis of ovarian cancer, and may be novel diagnostic and prognostic biomarkers for ovarian cancer although more studies are still needed to be evaluated.
Prostate Cancer
Prostate cancer (PCa) is one of the most common cancers and the third leading cause of deaths with high mortality and morbidity, especially for elderly men around the world.213,214 CircRNAs play important roles in the regulation of cell proliferation, apoptosis, angiogenesis and metastasis in a series of cancers including prostate cancer. Most of them could be used for the promising biomarkers and therapeutic target for the treatment of prostate cancer.215 One recent study have found that circITCH was significantly down-regulated in PCa cells and tissues, and inhibited the malignant phenotype of PCa via increasing the expression of homeobox protein B13 (HOXB13) through sponging miR-17-5p.216 Likewise, circRNA-UCK2 inhibited cell proliferation and invasion via increasing tet methylcytosine dioxygenase 1 (TET1) expression by sponging miR-767-5p in prostate cancer.217 Similarly, circFOXO3 sponged miR-29a-3p, circABCC4 sponged miR-1182, circHIPK3 sponged miRNA-338-3p and miR-193a-3p, and circAMOTL1L sponge dmiR-193a-5p were also reported in other studies.218–222 This may be one the main mechanisms of circRNAs to function as miRNA sponges in many cancers including prostate cancer. Moreover, circ_KATNAL1 significantly inhibited cell proliferation, invasion, migration of prostate cancer through the miR-145-3p/Wnt1 inducible signaling pathway protein 1 (WISP1) pathway, which might be a new mechanism for the progression of prostate cancer.223 More interestingly, recent study revealed that some of circRNAs could perform their functions by cooperating with their host genes. For example, X-linked inhibitor of apoptosis protein (XIAP), a host gene for circRNA0005276, showed to interact with circ0005276 to mediate the progression of prostate cancer through activating the transcription of XIAP via interacting with FUS binding protein.224 Thus, circRNAs are important regulators in gene expression and play a crucial role in prostate cancer, however, the detailed mechanisms for the tumorigenesis and progression should be more explored in the future.
Myeloid Leukemia
There are two main types of myeloid leukemia, acute myeloid leukemia (AML) and chronic myeloid leukemia (CML). AML is one of the most common myeloid malignancy in adults, characterized by the proliferation of abnormal and immature myeloid blasts in the bone marrow and presented great biological and clinical heterogeneity.225 CircRNAs were served as potential biomarkers for the diagnosis and treatment of AML because of their stability against exo-nuclear degradation, diversity of action modes, tissue specificity and richness in body fluids.225,226 Through high-throughput sequencing and bioinformatics analysis, 1824 circRNAs were detected as differentially expressed in AML cells.227 In addition, a total of 273 circRNAs were increased and 296 were decreased in pediatric AML. Among them, circ-0004136 was significantly increased and promoted cell proliferation by acting as a sponge of miR-142.228 Similarly, the following circRNAs also function as a miRNA sponge in acute myeloid leukemia, such as circ_0009910 sponging miR-20a-5p, circ_100290 sponging to miR-203 and circ-ANAPC7 sponging to miR-181.229–231 Furthermore, recent studies observed that circ-Foxo3 could compete with Foxo3 for binding to some miRNAs and then regulated the expression of Foxo3.232 Circ-Foxo3 and Foxo3 were frequently decreased in AML and positively associated with each other. Circ-Foxo3 might be a promising biomarker for the prognosis and treatment of AML.233 In addition, some circRNAs might be related to drug resistance in acute myeloid leukemia. For example, silence of circPAN3 significantly restored drug sensitivity to ADM in the two ADM-resistant cell lines, and overexpression of circPAN3 had the opposite effect. The results suggested that circPAN3 might facilitate AML drug resistance through regulating the AMPK/mTOR signaling pathway.234 In chronic myeloid leukemia (CML), recent study found that expressions of circHIPK3 and circRNA_100053 were significantly increased compared with healthy controls. Induced circHIPK3 expression predicted a poor outcome of CML patients, and circ_100053 might be associated with imatinib resistance in CML.235,236 In addition, circ_0080145 was found to be up-regulated in CML, and silence of circ_0080145 significantly inhibited cell proliferation of CML by sponging miR-29b.237 Taken together, circRNAs were distributed broadly in myeloid leukemia, and abnormal expressions of circRNAs were closely related to the progression and tumorigenesis of myeloid leukemia including AML and CML. Nevertheless, further studies are still required to determine the potential roles of circRNAs in diagnostic biomarker and therapeutic targets.
circRNAs in Cancer Stem Cells
Cancer stem cells (CSCs), a small proportion of cells that possess self-renewal and tumor-initiating capabilities, are considered to be responsible for metastatic dissemination and therapeutic failure. Several lines of evidence have suggested that circRNAs might contribute to the stemness of cancer.238 For example, around 27 dysregulated circRNA were observed through high-throughput sequencing to screen the circRNA expression profiles in breast CSCs (BCSCs) and matched non-BCSCs. Among these, expression of circVRK1 was reduced and was able to inhibit the self-renewal capacity of BCSCs, thereby displaying an inhibiting role in the stemness of BCSCs. Breast cancer cells with silenced circVRK1 demonstrated an enhanced capacity to form mammospheres and colonies, and an increased expression of CSC-related markers and core pluripotency genes (OCT4, SOX2, NANOG), indicating that circVRK1 was involved in suppressing the stemness of BCSCs.239 MiR-153-5p was one of the targets of circVRK1 and was involved in stemness maintenance of breast cancer cells via reducing the expression of KLF5. Thus, circVRK1 was negatively correlated with stemness of BCSCs through the miR-153-5p/krüppel-like factor 5 (KLF5) regulatory pathways.240 Stem cell plasticity and identity are also controlled by master regulatory genes and complex circuits involving circRNAs as well. One study showed that compared to differentiated mesodermal derivatives, circFOXP1 levels were enriched in mesenchymal stem cell (MSC) and silencing of circFOXP1 dramatically impaired MSC differentiation in vitro and in vivo. A direct interaction between circFOXP1 and miR-17-3p/miR-127-5p resulted in the modulation of the epidermal growth factor receptor (EGFR) and noncanonical Wnt pathways suggesting the regulatory role for circFOXP1 as a gatekeeper of pivotal stem cell molecular networks.241 In addition, the underlying correlation between circRNAs and cancer stem cells (CSCs) has been reported in HCC. For example, the absence of circZKSCAN1 endowed several malignant properties including cancer stemness and closely correlated with poor overall and recurrence-free survival in HCC. Bioinformatics analysis and RNA immunoprecipitation-sequencing (RIP-seq) experiments revealed that circZKSCAN1 showed inhibitory role by competitively binding RBP fragile X mental retardation protein (FMRP), thereby blocking the binding between FMRP and the downstream target gene cell cycle and apoptosis regulator 1 (CCAR1) mRNA, and subsequently retarded the transcriptional activity of Wnt signaling resulting in suppressing cell stemness in HCC cells.242 CD133+CD44+ cancer stem cells (TDP cells) previously isolated from laryngeal squamous cell carcinoma (LSCC) cells showed strong malignancy and tumorigenicity. These TDP cells were shown to highly express the stem-cell markers SOX2 and OCT4. Hg19_circ_0005033 was one of the upregulated circRNAs in TDP cells promoted the proliferation, migration, invasion, and resistance to chemotherapy of TDP cells.243 The expression of stem cell marker Kruppel-like factor-4 (KLF-4), which has been reported as the target of miR7, increased significantly in ciRS-7 transfected ESCC cells. Knockdown of KLF-4 also attenuated over expression of ciRS-7 induced cell invasion.98 Overall, the potential regulatory mechanism of circRNAs in CSC phenotypes and potential clinical applications in CSC-targeted therapy, including functioning as new biomarkers, acting as vaccines and breaking the therapeutic resistance of CSCs have been summarized.244 Researches regarding the regulatory roles of circRNAs on CSCs are still in the initial stages although the increased numbers of studies have demonstrated that the aberrant expression of circRNAs play a key role in the regulation and progression of cancers and CSCs. Thus, the practical application of circRNAs in clinic arena still remains to be determined. Future studies are needed to explore how circRNAs change in the CSC environment, among others.
Discussion and Prospective
CircRNAs have attracted increasing attention over the last decade. With the rapid development of biotechnology, bioinformatics analysis and publicly available high-throughput RNA-Seq data from the ENCODE consortium, a large numbers of circRNAs have been identified in recent years. CircRNAs can be detectable in body fluids, such as blood and saliva, urine, and breast milk including membrane-bound vesicles, such as exosome, and has widely involved in a variety of cancer-related physiological and pathology processes, including cancer initiation, progression and metastasis, drug resistance and played an important role in the diagnostic and prognostic biomarker and the therapeutic target in human cancer.245–247 It has become increasingly clear that circRNAs regulate gene expression through various actions and play diverse roles in many fields of human cancer biology. Recently, investigating the presence and expression levels of exosomal circRNAs could allow us to discriminate cancer patients from healthy individuals, identifying new potential exosome-based cancer biomarkers.248 Exosomal circRNAs are a novel frontier in cancer research and exploring the mysterious connection of exosome and circRNA may provide a vital hint to understand the biological functions of exosomal circRNAs. New studies show that exosomal circRNAs originating from tumor cells or other cells can transfer biological information to the specific cells to achieve the efficient transmission of phenotypical changes and thereby promoting cancer metastasis. Taking advantage of the stability and high specificity of exosomal circRNAs, these molecules might serve as promising cancer biomarkers with early detection and powerful prediction for patients to receive the most suitable therapy and might have potential for monitoring cancer progression or recurrence, and even to successfully develop therapeutic methods for the treatment of cancer although gaps in our current understanding of the connection of circRNAs with exosome still remain, such as the mechanism by which exosomal circRNAs travel in bodily fluids and the roles of exosomal circRNAs in cancer. Upon complete elucidation of exosomal circRNA functionality and molecular mechanisms relevant to human cancer, avenues of new insight will be opened, providing novel therapeutic approaches in malignant tumors.11
CircRNAs may also play a key role in the development of drug resistance. Recently, multiple studies have highlighted the key roles of ncRNAs in chemoresistance of cancer, such as HCC,142,249 lung cancer,250,251 gastric cancer,252,253 breast cancer,254,255 multiple myeloma,256 acute myeloid leukemia,234 prostate cancer,257 bladder cancer,258 among others. The up-to-date information regarding the role of circRNAs in the resistance of tumors to chemotherapy has been recently summarized with multiple mechanisms, such as modulating various regulatory pathways and processes including the ceRNA regulatory network axis, EMT process, regulation of ABC transporters, apoptosis, autophagy, and CSCs, among others although many physiological processes and biological signaling pathways through which circRNAs are involved in drug resistance still remained unknown.259 Thus, more mechanisms of action of chemoresistance-related circRNAs need to be explored in the future.
Importantly, the unique cellular stability and function of circRNAs to sponge miRNA and proteins may also indicate that circRNA is a promising vehicle for targeted drug delivery.4 So far, there has been no preclinical data demonstrating that circRNAs alone have been used as targets or therapeutic vectors for cancer treatment, but this direction will likely show promising in the future. The unique cellular stability and capacity of circRNA to sponge miRNA and protein may place circRNA as a promising vehicle for the delivery of cancer therapeutics. It is reasonable to believe that circRNAs will bring a new revolution for the diagnosis and treatment of human cancer in the near future.
However, there are also a number of challenges that need to be addressed. First of all, the expression level of most circRNAs are relative low in human cancer, therefore, we will require more advanced and sensitive technologies and tools to detect the molecular function of specific circRNA in the future.10 Secondly, owing to the fact that the majority of circRNA sequence is shared with the mRNA generated from the host gene. Hence, there are some troublesome technical problems need to be solved, such as circRNA quantification and validation, as well as overexpression and silencing strategies.10,260 Thirdly, the names of many circRNAs have not yet been standardized. As a result, many independent studies cannot be unified and generalized, which were not conducive to the sustainability and refer ability of circRNA research.25 Finally, the study of circRNAs in cancer is still in its infancy, and the functional role and mechanism of circRNA in distinct human cancers remains unclear. The current knowledge of circRNAs in tumorigenesis as well as their potential in diagnostic and prognostic biomarkers and possible therapeutic targets still remained to be elucidated. Thus, the in-depth underlying mechanism of circRNA in cancer biology needs to be explored further. It also speculated that the aberrant expression of circRNAs observed in cancer might also be explained by genetic and/or epigenetic changes of genes involved in their biogenesis. By addressing these issues and challenges with the advanced technology, improved experimental approaches and further research, we believe that circRNA could become a medically valuable diagnostic tool and an effective biological target for various cancers in the near future. Therefore, revealing cancer pathogenesis mechanisms and seeking novel potential diagnostic biomarkers or therapeutic targets will be popular topics in the future.261 Future detection of circRNA should also be explored the utilities of some new technologies, such as Oxford Nanopore sequencing, which can potentially provide information on the entire circRNA and could be an important addition to the mammalian transcriptomics toolbox262 and the NanoString nCounter Analysis System, which can quantify RNA molecules quantitative data output without amplification and reverse transcription.4,263 In conclusion, this study describes major features of circRNAs, summarizes the biological functions and mechanisms of circRNA associated with the occurrence, growth, progression, metastasis, drug resistance of human cancers. CircRNA can be used as potential diagnostic, prognostic biomarker and therapeutic target for personalized therapeutic for human cancer.
Abbreviations
circRNA, Circular RNA; MDM2, mouse double minute 2; RBPs, RNA-binding proteins; 3ʹUTRs, 3ʹ-untranslated regions; ILF2/3, interleukin enhancer-binding factor 2 and 3 complex; ITGA7, integrin alpha 7; RREB1, RAS-responsive element-binding protein 1; NSCLC, non-small cell lung cancer; NOVA1, NOVA alternative splicing regulator 1; snRNA, U1 small nuclear RNA; ORF, open reading frame; IRES, internal ribosome entry site; STAT3, signal transducer and activator of transcription 3; PRKAA, protein kinase AMP-activated catalytic subunit alpha 2; AMPKα, AMP-activated protein kinase; RBPs, RNA-binding proteins; RBM3, RNA-binding protein 3; SCD, stearoyl-CoA desaturase; RPCs, RNA-protein complexes; CDK2, cyclin-dependent kinase 2; SCML1, sex comb on midleg-like 1; HN, humanin; EMT, epithelial mesenchymal transitioning; RASSF8, ras association domain family 8; EML4-ALK, echinoderm microtubule-associated protein-like 4-anaplastic lymphoma kinase; ceRNA, endogenous RNA; BC, breast cancer; TRAF, TNFα receptor associated factor; TNBC, triple-negative breast cancer; CRC, colorectal cancer; RNA-seq, RNA sequencing; APC2, adenomatous polyposis coli 2; CRR, chemoradiation resistance; ESCC, Esophageal squamous cell carcinoma; TTC17, tetratricopeptide repeat domain 17; mTOR, mammalian target of rapamycin; GC, gastric cancer; SRY sex-determining region Y; SOX4, SRY-related high-mobility group box 4; AGO2, argonaute 2; HuR, human antigen R; HCC, hepatocellular carcinoma; CDK6, cyclin-dependent kinase 6; NK, natural killer; UVRAG, UV radiation resistance associated gene; FGFR2, fibroblast growth factor receptor 2; B4GALT3, β-1, 4-Galactosyltransferase III; CSCs, cancer stem cells; SERPINA4, serpina family A member 4; IL-6R, interleukin-6 receptor; JMJD2C, jumonji domain containing 2c; MACC1, metastasis-associated in colon cancer-1; HGF, hepatocyte growth factor; PTC, papillary thyroid cancer; LMO4, LIM-domain only protein 4; FGFR1, fibroblast growth factor receptor 1; CTNNBIP1, catenin beta interacting protein 1; C8orf4, chromosome 8 open reading frame 4; GATAD2A, GATA zinc finger domain containing 2A; FAT1, FAT atypical cadherin 1; YAP1, yes-associated protein 1; CCNE1, Cyclin E1; PI3KCD, phosphatidylinositol-4, 5-bisphosphate 3-kinase catalytic subunit delta; RAF1, RAF1 proto-oncogene serine/threonine-protein kinase; ALP, alkaline phosphatase; LDH, lactate dehydrogenase; CDR1, cerebellar degeneration-related auto-antigen 1; ADARB2, adenosine deaminase, RNA-specific B2; GO, Gene Ontology; SHPRH, SNF2 histone linker PHD RING helicase; SOCS3, suppressor of cytokine signaling 3; RIP-seq, RNA immunoprecipitation-sequencing; CCAR1, cell cycle and apoptosis regulator 1; FMRP, fragile X mental retardation protein; EGFR, epidermal growth factor receptor; MSC, mesenchymal stem cell; KLF5, krüppel-like factor 5; BCSCs, breast CSCs; KLF-4, Kruppel-like factor-4; ecRNAs, exonic circRNAs; ciRNAs, intronic circRNAs; eIciRNAs, exon-intron circRNAs; LSCC, laryngeal squamous cell carcinoma; TDP cells, CD133+CD44+ cancer stem cells; NF-κB, nuclear factor kappa B; MMP2, matrix metallopeptidase 2; MUC1, mucin 1; hTERT, human telomerase reverse transcriptase; VAMP2, vesicle-associated membrane protein 2; MDM2, miR-647mouse double minute 2; DNAJB6, DnaJ/Hsp40 homolog, subfamily B, member 6; PCa, prostate cancer; HOXB13, homeobox protein B13; TET1, tet methylcytosine dioxygenase 1; WISP1, Wnt1 inducible signaling pathway protein 1; XIAP, X-linked inhibitor of apoptosis protein.
Acknowledgments
This work was supported in part by the grants from the National Natural Scientific Foundation of China (Nos. 81703551, 81871863) and the Major Program of National Natural Science Foundation of Guangdong (No. 2018B030311061).
Author Contributions
All authors made substantial contributions to conception and design, acquisition of data, or analysis and interpretation of data; took part in drafting the article or revising it critically for important intellectual content; gave final approval of the version to be published; and agree to be accountable for all aspects of the work.
Disclosure
The authors report no conflicts of interest in this work.
References
1. Chen G, Wang Q, Yang Q, et al. Circular RNAs hsa_circ_0032462, hsa_circ_0028173, hsa_circ_0005909 are predicted to promote CADM1 expression by functioning as miRNAs sponge in human osteosarcoma. PLoS One. 2018;13:e0202896. doi:10.1371/journal.pone.0202896
2. Liu X, Abraham JM, Cheng Y, et al. Synthetic circular RNA functions as a miR-21 sponge to suppress gastric carcinoma cell proliferation. Mol Ther Nucleic Acids. 2018;13:312–321. doi:10.1016/j.omtn.2018.09.010
3. Du WW, Fang L, Yang W, et al. Induction of tumor apoptosis through a circular RNA enhancing Foxo3 activity. Cell Death Differ. 2017;24:357–370. doi:10.1038/cdd.2016.133
4. Kristensen LS, Hansen TB, Veno MT, Kjems J. Circular RNAs in cancer: opportunities and challenges in the field. Oncogene. 2018;37:555–565. doi:10.1038/onc.2017.361
5. Li Z, Huang C, Bao C, et al. Exon-intron circular RNAs regulate transcription in the nucleus. Nat Struct Mol Biol. 2015;22:256–264. doi:10.1038/nsmb.2959
6. Liu F, Zhang J, Qin L, et al. Circular RNA EIF6 (Hsa_circ_0060060) sponges miR-144-3p to promote the cisplatin-resistance of human thyroid carcinoma cells by autophagy regulation. Aging (Albany NY). 2018;10:3806–3820. doi:10.18632/aging.v10i12
7. Sanger HL, Klotz G, Riesner D, Gross HJ, Kleinschmidt AK. Viroids are single-stranded covalently closed circular RNA molecules existing as highly base-paired rod-like structures. Proc Natl Acad Sci U S A. 1976;73:3852–3856. doi:10.1073/pnas.73.11.3852
8. Chen LL. The biogenesis and emerging roles of circular RNAs. Nat Rev Mol Cell Biol. 2016;17:205–211. doi:10.1038/nrm.2015.32
9. Pamudurti NR, Bartok O, Jens M, et al. Translation of circRNAs. Mol Cell. 2017;66:9–21 e7. doi:10.1016/j.molcel.2017.02.021
10. Patop IL, Kadener S. circRNAs in cancer. Curr Opin Genet Dev. 2018;48:121–127. doi:10.1016/j.gde.2017.11.007
11. Shi X, Wang B, Feng X, Xu Y, Lu K, Sun M. circRNAs and exosomes: a mysterious frontier for human cancer. Mol Ther Nucleic Acids. 2019;19:384–392. doi:10.1016/j.omtn.2019.11.023
12. Shang Q, Yang Z, Jia R, Ge S. The novel roles of circRNAs in human cancer. Mol Cancer. 2019;18:6. doi:10.1186/s12943-018-0934-6
13. Hansen TB, Jensen TI, Clausen BH, et al. Natural RNA circles function as efficient microRNA sponges. Nature. 2013;495:384–388. doi:10.1038/nature11993
14. Xie R, Tang J, Zhu X, Jiang H. Silencing of hsa_circ_0004771 inhibits proliferation and induces apoptosis in breast cancer through activation of miR-653 by targeting ZEB2 signaling pathway. Biosci Rep. 2019;39. doi:10.1042/BSR20181919
15. Yang R, Xing L, Zheng X, Sun Y, Wang X, Chen J. The circRNA circAGFG1 acts as a sponge of miR-195-5p to promote triple-negative breast cancer progression through regulating CCNE1 expression. Mol Cancer. 2019;18:4. doi:10.1186/s12943-018-0933-7
16. Guarnerio J, Zhang Y, Cheloni G, et al. Intragenic antagonistic roles of protein and circRNA in tumorigenesis. Cell Res. 2019;29:628–640. doi:10.1038/s41422-019-0192-1
17. Li X, Wang J, Zhang C, et al. Circular RNA circITGA7 inhibits colorectal cancer growth and metastasis by modulating the Ras pathway and upregulating transcription of its host gene ITGA7. J Pathol. 2018;246:166–179. doi:10.1002/path.2018.246.issue-2
18. Qin M, Wei G, Sun X. Circ-UBR5: an exonic circular RNA and novel small nuclear RNA involved in RNA splicing. Biochem Biophys Res Commun. 2018;503:1027–1034. doi:10.1016/j.bbrc.2018.06.112
19. Li YJ, Lei YH, Yao N, et al. Autophagy and multidrug resistance in cancer. Chin J Cancer. 2017;36:52. doi:10.1186/s40880-017-0219-2
20. Mowers EE, Sharifi MN, Macleod KF. Autophagy in cancer metastasis. Oncogene. 2017;36:1619–1630. doi:10.1038/onc.2016.333
21. Chen X, Mao R, Su W, et al. Circular RNA circHIPK3 modulates autophagy via MIR124-3p-STAT3-PRKAA/AMPKalpha signaling in STK11 mutant lung cancer. Autophagy. 2019;1–13. doi:10.1080/15548627.2019.1634945
22. Chi G, Xu D, Zhang B, Yang F. Matrine induces apoptosis and autophagy of glioma cell line U251 by regulation of circRNA-104075/BCL-9. Chem Biol Interact. 2019;308:198–205. doi:10.1016/j.cbi.2019.05.030
23. Zang J, Lu D, Xu A. The interaction of circRNAs and RNA binding proteins: an important part of circRNA maintenance and function. J Neurosci Res. 2020;98(1):87–97.
24. Hentze MW, Preiss T. Circular RNAs: splicing’s enigma variations. EMBO J. 2013;32:923–925. doi:10.1038/emboj.2013.53
25. Xu S, Zhou L, Ponnusamy M, et al. A comprehensive review of circRNA: from purification and identification to disease marker potential. PeerJ. 2018;6:e5503. doi:10.7717/peerj.5503
26. Dong W, Dai ZH, Liu FC, et al. The RNA-binding protein RBM3 promotes cell proliferation in hepatocellular carcinoma by regulating circular RNA SCD-circRNA 2 production. EBioMedicine. 2019. doi:10.1016/j.ebiom.2019.06.030
27. Conn SJ, Pillman KA, Toubia J, et al. The RNA binding protein quaking regulates formation of circRNAs. Cell. 2015;160:1125–1134. doi:10.1016/j.cell.2015.02.014
28. Khan MA, Reckman YJ, Aufiero S, et al. RBM20 regulates circular RNA production from the titin gene. Circ Res. 2016;119:996–1003. doi:10.1161/CIRCRESAHA.116.309568
29. Du WW, Yang W, Liu E, Yang Z, Dhaliwal P, Yang BB. Foxo3 circular RNA retards cell cycle progression via forming ternary complexes with p21 and CDK2. Nucleic Acids Res. 2016;44:2846–2858. doi:10.1093/nar/gkw027
30. Ji X, Bosse Y, Landi MT, et al. Identification of susceptibility pathways for the role of chromosome 15q25.1 in modifying lung cancer risk. Nat Commun. 2018;9:3221. doi:10.1038/s41467-018-05074-y
31. Chen Y, Wei S, Wang X, Zhu X, Han S. Progress in research on the role of circular RNAs in lung cancer. World J Surg Oncol. 2018;16:215. doi:10.1186/s12957-018-1515-2
32. Liu XX, Yang YE, Liu X, et al. A two-circular RNA signature as a noninvasive diagnostic biomarker for lung adenocarcinoma. J Transl Med. 2019;17:50. doi:10.1186/s12967-019-1800-z
33. Ma Y, Zhang X, Wang YZ, Tian H, Xu S. Research progress of circular RNAs in lung cancer. Cancer Biol Ther. 2019;20:123–129. doi:10.1080/15384047.2018.1523848
34. Cheng XY, Shen H. Circular RNA in lung cancer research: biogenesis, functions and roles. Zhongguo Fei Ai Za Zhi. 2018;21:50–56. doi:10.3779/j.issn.1009-3419.2018.01.07
35. Ding X, Zhang S, Li X, et al. Profiling expression of coding genes, long noncoding RNA, and circular RNA in lung adenocarcinoma by ribosomal RNA-depleted RNA sequencing. FEBS Open Bio. 2018;8:544–555. doi:10.1002/feb4.2018.8.issue-4
36. Hu W, Bi ZY, Chen ZL, et al. Emerging landscape of circular RNAs in lung cancer. Cancer Lett. 2018;427:18–27. doi:10.1016/j.canlet.2018.04.006
37. Zhao J, Li L, Wang Q, Han H, Zhan Q, Xu M. CircRNA expression profile in early-stage lung adenocarcinoma patients. Cell Physiol Biochem. 2017;44:2138–2146. doi:10.1159/000485953
38. Nan A, Chen L, Zhang N, et al. Circular RNA circNOL10 inhibits lung cancer development by promoting SCLM1-mediated transcriptional regulation of the humanin polypeptide family. Adv Sci (Weinh). 2019;6:1800654. doi:10.1002/advs.201800654
39. Wei S, Zheng Y, Jiang Y, et al. The circRNA circPTPRA suppresses epithelial-mesenchymal transitioning and metastasis of NSCLC cells by sponging miR-96-5p. EBioMedicine. 2019. doi:10.1016/j.ebiom.2019.05.032
40. Chen D, Ma W, Ke Z, Xie F. CircRNA hsa_circ_100395 regulates miR-1228/TCF21 pathway to inhibit lung cancer progression. Cell Cycle. 2018;17:2080–2090. doi:10.1080/15384101.2018.1515553
41. Han J, Zhao G, Ma X, et al. CircRNA circ-BANP-mediated miR-503/LARP1 signaling contributes to lung cancer progression. Biochem Biophys Res Commun. 2018;503:2429–2435. doi:10.1016/j.bbrc.2018.06.172
42. Li Y, Hu J, Li L, et al. Upregulated circular RNA circ_0016760 indicates unfavorable prognosis in NSCLC and promotes cell progression through miR-1287/GAGE1 axis. Biochem Biophys Res Commun. 2018;503:2089–2094. doi:10.1016/j.bbrc.2018.07.164
43. Liu W, Ma W, Yuan Y, Zhang Y, Sun S. Circular RNA hsa_circRNA_103809 promotes lung cancer progression via facilitating ZNF121-dependent MYC expression by sequestering miR-4302. Biochem Biophys Res Commun. 2018;500:846–851. doi:10.1016/j.bbrc.2018.04.172
44. Tian X, Zhang L, Jiao Y, Chen J, Shan Y, Yang W. CircABCB10 promotes nonsmall cell lung cancer cell proliferation and migration by regulating the miR-1252/FOXR2 axis. J Cell Biochem. 2019;120:3765–3772. doi:10.1002/jcb.v120.3
45. Yao Y, Hua Q, Zhou Y. CircRNA has_circ_0006427 suppresses the progression of lung adenocarcinoma by regulating miR-6783-3p/DKK1 axis and inactivating Wnt/beta-catenin signaling pathway. Biochem Biophys Res Commun. 2019;508:37–45. doi:10.1016/j.bbrc.2018.11.079
46. Zhao F, Han Y, Liu Z, Zhao Z, Li Z, Jia K. circFADS2 regulates lung cancer cells proliferation and invasion via acting as a sponge of miR-498. Biosci Rep. 2018;38. doi:10.1042/BSR20180570
47. Chang H, Qu J, Wang J, Liang X, Sun W. Circular RNA circ_0026134 regulates non-small cell lung cancer cell proliferation and invasion via sponging miR-1256 and miR-1287. Biomed Pharmacother. 2019;112:108743. doi:10.1016/j.biopha.2019.108743
48. Liu G, Shi H, Deng L, et al. Circular RNA circ-FOXM1 facilitates cell progression as ceRNA to target PPDPF and MACC1 by sponging miR-1304-5p in non-small cell lung cancer. Biochem Biophys Res Commun. 2019;513:207–212. doi:10.1016/j.bbrc.2019.03.213
49. Wang T, Wang X, Du Q, et al. The circRNA circP4HB promotes NSCLC aggressiveness and metastasis by sponging miR-133a-5p. Biochem Biophys Res Commun. 2019;513:904–911. doi:10.1016/j.bbrc.2019.04.108
50. Zhang B, Chen M, Jiang N, Shi K, Qian R. A regulatory circuit of circ-MTO1/miR-17/QKI-5 inhibits the proliferation of lung adenocarcinoma. Cancer Biol Ther. 2019;20(8):1–9.
51. An J, Shi H, Zhang N, Song S. Elevation of circular RNA circ_0003645 forecasts unfavorable prognosis and facilitates cell progression via miR-1179/TMEM14A pathway in non-small cell lung cancer. Biochem Biophys Res Commun. 2019;511:921–925. doi:10.1016/j.bbrc.2019.03.011
52. Ma X, Yang X, Bao W, et al. Circular RNA circMAN2B2 facilitates lung cancer cell proliferation and invasion via miR-1275/FOXK1 axis. Biochem Biophys Res Commun. 2018;498:1009–1015. doi:10.1016/j.bbrc.2018.03.105
53. Wang X, Zhu X, Zhang H, et al. Increased circular RNA hsa_circ_0012673 acts as a sponge of miR-22 to promote lung adenocarcinoma proliferation. Biochem Biophys Res Commun. 2018;496:1069–1075. doi:10.1016/j.bbrc.2018.01.126
54. Wan J, Hao L, Zheng X, Li Z. Circular RNA circ_0020123 promotes non-small cell lung cancer progression by acting as a ceRNA for miR-488-3p to regulate ADAM9 expression. Biochem Biophys Res Commun. 2019;515:303–309. doi:10.1016/j.bbrc.2019.05.158
55. Guarnerio J, Bezzi M, Jeong JC, et al. Oncogenic role of fusion-circRNAs derived from cancer-associated chromosomal translocations. Cell. 2016;166:1055–1056. doi:10.1016/j.cell.2016.07.035
56. Wu K, Liao X, Gong Y, et al. Circular RNA F-circSR derived from SLC34A2-ROS1 fusion gene promotes cell migration in non-small cell lung cancer. Mol Cancer. 2019;18:98. doi:10.1186/s12943-019-1028-9
57. Tan S, Sun D, Pu W, et al. Circular RNA F-circEA-2a derived from EML4-ALK fusion gene promotes cell migration and invasion in non-small cell lung cancer. Mol Cancer. 2018;17:138. doi:10.1186/s12943-018-0887-9
58. Tan S, Gou Q, Pu W, et al. Circular RNA F-circEA produced from EML4-ALK fusion gene as a novel liquid biopsy biomarker for non-small cell lung cancer. Cell Res. 2018;28:693–695. doi:10.1038/s41422-018-0033-7
59. Zhang S, Xia W, Dong G, Xu W, Li M, Xu L. Cyclic RNA molecule circ_0007766 promotes the proliferation of lung adenocarcinoma cells by up-regulating the expression of Cyclin D1/CyclinE1/CDK4]. Zhongguo Fei Ai Za Zhi. 2019;22:271–279. doi:10.3779/j.issn.1009-3419.2019.05.03
60. Wang L, Ma H, Kong W, Liu B, Zhang X. Up-regulated circular RNA VANGL1 contributes to progression of non-small cell lung cancer through inhibition of miR-195 and activation of Bcl-2. Biosci Rep. 2019;39(6).
61. Zhou SY, Chen W, Yang SJ, et al. The emerging role of circular RNAs in breast cancer. Biosci Rep. 2019;39. doi:10.1042/BSR20190621
62. Wang H, Xiao Y, Wu L, Ma D. Comprehensive circular RNA profiling reveals the regulatory role of the circRNA-000911/miR-449a pathway in breast carcinogenesis. Int J Oncol. 2018;52:743–754. doi:10.3892/ijo.2018.4265
63. Nair AA, Niu N, Tang X, et al. Circular RNAs and their associations with breast cancer subtypes. Oncotarget. 2016;7:80967–80979. doi:10.18632/oncotarget.13134
64. Wang X, Fang L. Advances in circular RNAs and their roles in breast cancer. J Exp Clin Cancer Res. 2018;37:206. doi:10.1186/s13046-018-0870-8
65. Zeng K, He B, Yang BB, et al. The pro-metastasis effect of circANKS1B in breast cancer. Mol Cancer. 2018;17:160. doi:10.1186/s12943-018-0914-x
66. Xu JH, Wang Y, Xu D. Hsa_circ_001569 is an unfavorable prognostic factor and promotes cell proliferation and metastasis by modulating PI3K-AKT pathway in breast cancer. Cancer Biomark. 2019;25:193–201. doi:10.3233/CBM-182293
67. Liu Y, Dong Y, Zhao L, Su L, Luo J. Circular RNAMTO1 suppresses breast cancer cell viability and reverses monastrol resistance through regulating the TRAF4/Eg5 axis. Int J Oncol. 2018;53:1752–1762. doi:10.3892/ijo.2018.4485
68. Xu JZ, Shao CC, Wang XJ, et al. circTADA2As suppress breast cancer progression and metastasis via targeting miR-203a-3p/SOCS3 axis. Cell Death Dis. 2019;10:175. doi:10.1038/s41419-019-1382-y
69. Tang H, Huang X, Wang J, et al. circKIF4A acts as a prognostic factor and mediator to regulate the progression of triple-negative breast cancer. Mol Cancer. 2019;18:23. doi:10.1186/s12943-019-0946-x
70. Kong Y, Yang L, Wei W, et al. CircPLK1 sponges miR-296-5p to facilitate triple-negative breast cancer progression. Epigenomics. 2019;11:1163–1176. doi:10.2217/epi-2019-0093
71. Wang Y, Li J, Du C, et al. Upregulated circular RNA circ-UBE2D2 predicts poor prognosis and promotes breast cancer progression by sponging miR-1236 and miR-1287. Transl Oncol. 2019;12:1305–1313. doi:10.1016/j.tranon.2019.05.016
72. Liang Y, Song X, Li Y, et al. Targeting the circBMPR2/miR-553/USP4 axis as a potent therapeutic approach for breast cancer. Mol Ther Nucleic Acids. 2019;17:347–361. doi:10.1016/j.omtn.2019.05.005
73. Sang Y, Chen B, Song X, et al. circRNA_0025202 regulates tamoxifen sensitivity and tumor progression via regulating the miR-182-5p/FOXO3a axis in breast cancer. Mol Ther. 2019;27:1638–1652. doi:10.1016/j.ymthe.2019.05.011
74. Yang L, Song C, Chen Y, Jing G, Sun J. Circular RNA circ_0103552 forecasts dismal prognosis and promotes breast cancer cell proliferation and invasion by sponging miR-1236. J Cell Biochem. 2019;120:15553–15560. doi:10.1002/jcb.v120.9
75. Yan L, Zheng M, Wang H. Circular RNA hsa_circ_0072309 inhibits proliferation and invasion of breast cancer cells via targeting miR-492. Cancer Manag Res. 2019;11:1033–1041. doi:10.2147/CMAR.S186857
76. Liu Z, Zhou Y, Liang G, et al. Circular RNA hsa_circ_001783 regulates breast cancer progression via sponging miR-200c-3p. Cell Death Dis. 2019;10:55. doi:10.1038/s41419-018-1287-1
77. Taborda MI, Ramirez S, Bernal G. Circular RNAs in colorectal cancer: possible roles in regulation of cancer cells. World J Gastrointest Oncol. 2017;9:62–69. doi:10.4251/wjgo.v9.i2.62
78. Zhang P, Zuo Z, Shang W, et al. Identification of differentially expressed circular RNAs in human colorectal cancer. Tumour Biol. 2017;39:1010428317694546.
79. Li XN, Wang ZJ, Ye CX, Zhao BC, Li ZL, Yang Y. RNA sequencing reveals the expression profiles of circRNA and indicates that circDDX17 acts as a tumor suppressor in colorectal cancer. J Exp Clin Cancer Res. 2018;37:325. doi:10.1186/s13046-018-1006-x
80. Jin Y, Yu LL, Zhang B, Liu CF, Chen Y. Circular RNA hsa_circ_0000523 regulates the proliferation and apoptosis of colorectal cancer cells as miRNA sponge. Braz J Med Biol Res. 2018;51:e7811. doi:10.1590/1414-431x20187811
81. Geng Y, Zheng X, Hu W, et al. Hsa_circ_0009361 acts as the sponge of miR-582 to suppress colorectal cancer progression by regulating APC2 expression. Clin Sci (Lond). 2019;133:1197–1213. doi:10.1042/CS20190286
82. Li XN, Wang ZJ, Ye CX, Zhao BC, Huang XX, Yang L. Circular RNA circVAPA is up-regulated and exerts oncogenic properties by sponging miR-101 in colorectal cancer. Biomed Pharmacother. 2019;112:108611. doi:10.1016/j.biopha.2019.108611
83. Jin C, Wang A, Liu L, Wang G, Li G. Hsa_circ_0136666 promotes the proliferation and invasion of colorectal cancer through miR-136/SH2B1 axis. J Cell Physiol. 2019;234:7247–7256. doi:10.1002/jcp.27482
84. Bian L, Zhi X, Ma L, et al. Hsa_circRNA_103809 regulated the cell proliferation and migration in colorectal cancer via miR-532-3p/FOXO4 axis. Biochem Biophys Res Commun. 2018;505:346–352. doi:10.1016/j.bbrc.2018.09.073
85. Fang G, Ye BL, Hu BR, Ruan XJ, Shi YX. CircRNA_100290 promotes colorectal cancer progression through miR-516b-induced downregulation of FZD4 expression and Wnt/beta-catenin signaling. Biochem Biophys Res Commun. 2018;504:184–189. doi:10.1016/j.bbrc.2018.08.152
86. Yuan Y, Liu W, Zhang Y, Zhang Y, Sun S. CircRNA circ_0026344 as a prognostic biomarker suppresses colorectal cancer progression via microRNA-21 and microRNA-31. Biochem Biophys Res Commun. 2018;503:870–875. doi:10.1016/j.bbrc.2018.06.089
87. Zeng K, Chen X, Xu M, et al. CircHIPK3 promotes colorectal cancer growth and metastasis by sponging miR-7. Cell Death Dis. 2018;9:417. doi:10.1038/s41419-018-0454-8
88. Xie H, Ren X, Xin S, et al. Emerging roles of circRNA_001569 targeting miR-145 in the proliferation and invasion of colorectal cancer. Oncotarget. 2016;7:26680–26691. doi:10.18632/oncotarget.8589
89. Xiong W, Ai YQ, Li YF, et al. Microarray analysis of circular RNA expression profile associated with 5-fluorouracil-based chemoradiation resistance in colorectal cancer cells. Biomed Res Int. 2017;2017:8421614. doi:10.1155/2017/8421614
90. Kang M, Li Y, Zhu S, Zhang S, Guo S, Li P. MicroRNA-193b acts as a tumor suppressor gene in human esophageal squamous cell carcinoma via target regulation of KRAS. Oncol Lett. 2019;17:3965–3973. doi:10.3892/ol.2019.10039
91. Wen J, Hu Y, Liu Q, et al. miR-424 coordinates multilayered regulation of cell cycle progression to promote esophageal squamous cell carcinoma cell proliferation. EBioMedicine. 2018;37:110–124. doi:10.1016/j.ebiom.2018.10.043
92. Fan L, Cao Q, Liu J, Zhang J, Li B. Circular RNA profiling and its potential for esophageal squamous cell cancer diagnosis and prognosis. Mol Cancer. 2019;18:16. doi:10.1186/s12943-018-0936-4
93. Wang Y, Lu T, Wang Q, Liu J, Jiao W. Circular RNAs: crucial regulators in the human body (review). Oncol Rep. 2018;40:3119–3135. doi:10.3892/or.2018.6733
94. Wang Q, Zhang Q, Sun H, et al. Circ-TTC17 promotes proliferation and migration of esophageal squamous cell carcinoma. Dig Dis Sci. 2019;64:751–758. doi:10.1007/s10620-018-5382-z
95. Shi N, Shan B, Gu B, Song Y, Chu H, Qian L. Circular RNA circ-PRKCI functions as a competitive endogenous RNA to regulate AKT3 expression by sponging miR-3680-3p in esophageal squamous cell carcinoma. J Cell Biochem. 2019;120:10021–10030. doi:10.1002/jcb.v120.6
96. Shi Y, Guo Z, Fang N, et al. hsa_circ_0006168 sponges miR-100 and regulates mTOR to promote the proliferation, migration and invasion of esophageal squamous cell carcinoma. Biomed Pharmacother. 2019;117:109151. doi:10.1016/j.biopha.2019.109151
97. Rong J, Wang Q, Zhang Y, et al. Circ-DLG1 promotes the proliferation of esophageal squamous cell carcinoma. Onco Targets Ther. 2018;11:6723–6730. doi:10.2147/OTT
98. Huang H, Wei L, Qin T, Yang N, Li Z, Xu Z. Circular RNA ciRS-7 triggers the migration and invasion of esophageal squamous cell carcinoma via miR-7/KLF4 and NF-kappaB signals. Cancer Biol Ther. 2019;20:73–80. doi:10.1080/15384047.2018.1507254
99. Song H, Xu D, Shi P, et al. Upregulated circ RNA hsa_circ_0000337 promotes cell proliferation, migration, and invasion of esophageal squamous cell carcinoma. Cancer Manag Res. 2019;11:1997–2006. doi:10.2147/CMAR.S195546
100. Song Z, Wu Y, Yang J, Yang D, Fang X. Progress in the treatment of advanced gastric cancer. Tumour Biol. 2017;39:1010428317714626. doi:10.1177/1010428317714626
101. Chevallay M, Jung M, Morel P, Monig S. Gastric cancer: management and multidisciplinary treatment. Rev Med Suisse. 2018;14:2221–2225.
102. Tian M, Chen R, Li T, Xiao B. Reduced expression of circRNA hsa_circ_0003159 in gastric cancer and its clinical significance. J Clin Lab Anal. 2018;32. doi:10.1002/jcla.22281
103. Chen S, Li T, Zhao Q, Xiao B, Guo J. Using circular RNA hsa_circ_0000190 as a new biomarker in the diagnosis of gastric cancer. Clin Chim Acta. 2017;466:167–171. doi:10.1016/j.cca.2017.01.025
104. Jiang F, Hong F, Shah MW, Shen X. Circular RNAs as diagnostic biomarkers in gastric cancer: a meta-analysis review. Pathol Res Pract. 2019;215:152419. doi:10.1016/j.prp.2019.04.011
105. Liu H, Liu Y, Bian Z, et al. Circular RNA YAP1 inhibits the proliferation and invasion of gastric cancer cells by regulating the miR-367-5p/p27 (Kip1) axis. Mol Cancer. 2018;17:151. doi:10.1186/s12943-018-0902-1
106. Zhang X, Wang S, Wang H, et al. Circular RNA circNRIP1 acts as a microRNA-149-5p sponge to promote gastric cancer progression via the AKT1/mTOR pathway. Mol Cancer. 2019;18:20. doi:10.1186/s12943-018-0935-5
107. Liang M, Huang G, Liu Z, et al. Elevated levels of hsa_circ_006100 in gastric cancer promote cell growth and metastasis via miR-195/GPRC5A signalling. Cell Prolif. 2019;52(5):e12661.
108. Guan EC, Xu XG, Xue FX. circ-NOTCH1 acts as a sponge of miR-637 and affects the expression of its target gene apelin to regulate gastric cancer cell growth. Biochem Cell Biol. 2019. doi:10.1139/bcb-2019-0079
109. Wu L, Liu D, Yang Y. Enhanced expression of circular RNA circ-DCAF6 predicts adverse prognosis and promotes cell progression via sponging miR-1231 and miR-1256 in gastric cancer. Exp Mol Pathol. 2019;110:104273. doi:10.1016/j.yexmp.2019.104273
110. Huang S, Zhang X, Guan B, et al. A novel circular RNA hsa_circ_0008035 contributes to gastric cancer tumorigenesis through targeting the miR-375/YBX1 axis. Am J Transl Res. 2019;11:2455–2462.
111. Lu J, Zhang PY, Li P, et al. Circular RNA hsa_circ_0001368 suppresses the progression of gastric cancer by regulating miR-6506-5p/FOXO3 axis. Biochem Biophys Res Commun. 2019;512:29–33. doi:10.1016/j.bbrc.2019.02.111
112. Rong D, Lu C, Zhang B, et al. CircPSMC3 suppresses the proliferation and metastasis of gastric cancer by acting as a competitive endogenous RNA through sponging miR-296-5p. Mol Cancer. 2019;18:25. doi:10.1186/s12943-019-0958-6
113. Wang Z, Ma K, Pitts S, et al. Novel circular RNA NF1 acts as a molecular sponge, promoting gastric cancer by absorbing miR-16. Endocr Relat Cancer. 2019;26(3):265–277.
114. Sun H, Xi P, Sun Z, et al. Circ-SFMBT2 promotes the proliferation of gastric cancer cells through sponging miR-182-5p to enhance CREB1 expression. Cancer Manag Res. 2018;10:5725–5734. doi:10.2147/CMAR
115. Fang J, Hong H, Xue X, et al. A novel circular RNA, circFAT1(e2), inhibits gastric cancer progression by targeting miR-548g in the cytoplasm and interacting with YBX1 in the nucleus. Cancer Lett. 2019;442:222–232. doi:10.1016/j.canlet.2018.10.040
116. Ouyang Y, Li Y, Huang Y, et al. CircRNA circPDSS1 promotes the gastric cancer progression by sponging miR-186-5p and modulating NEK2. J Cell Physiol. 2019;234:10458–10469. doi:10.1002/jcp.27714
117. Wang L, Shen J, Jiang Y. Circ_0027599/PHDLA1 suppresses gastric cancer progression by sponging miR-101-3p.1. Cell Biosci. 2018;8:58. doi:10.1186/s13578-018-0252-0
118. Xue M, Li G, Fang X, Wang L, Jin Y, Zhou Q. hsa_circ_0081143 promotes cisplatin resistance in gastric cancer by targeting miR-646/CDK6 pathway. Cancer Cell Int. 2019;19:25. doi:10.1186/s12935-019-0737-x
119. Ding L, Zhao Y, Dang S, et al. Circular RNA circ-DONSON facilitates gastric cancer growth and invasion via NURF complex dependent activation of transcription factor SOX4. Mol Cancer. 2019;18:45. doi:10.1186/s12943-019-1006-2
120. Chen Y, Yang F, Fang E, et al. Circular RNA circAGO2 drives cancer progression through facilitating HuR-repressed functions of AGO2-miRNA complexes. Cell Death Differ. 2019;26:1346–1364. doi:10.1038/s41418-018-0220-6
121. Boyvat F. Interventional radiologic treatment of hepatocellular carcinoma. Exp Clin Transplant. 2017;15:25–30. doi:10.6002/ect.TOND16.L8
122. Chen S, Cao Q, Wen W, Wang H. Targeted therapy for hepatocellular carcinoma: challenges and opportunities. Cancer Lett. 2019;460:1–9.
123. Grandhi MS, Kim AK, Ronnekleiv-Kelly SM, Kamel IR, Ghasebeh MA, Pawlik TM. Hepatocellular carcinoma: from diagnosis to treatment. Surg Oncol. 2016;25:74–85. doi:10.1016/j.suronc.2016.03.002
124. Gong Y, Mao J, Wu D, et al. Circ-ZEB1.33 promotes the proliferation of human HCC by sponging miR-200a-3p and upregulating CDK6. Cancer Cell Int. 2018;18:116.
125. Wang Z, Zhao Y, Wang Y, Jin C. Circular RNA circHIAT1 inhibits cell growth in hepatocellular carcinoma by regulating miR-3171/PTEN axis. Biomed Pharmacother. 2019;116:108932. doi:10.1016/j.biopha.2019.108932
126. Zou H, Xu X, Luo L, et al. Hsa_circ_0101432 promotes the development of hepatocellular carcinoma (HCC) by adsorbing miR-1258 and miR-622. Cell Cycle. 2019;18:2398–2413. doi:10.1080/15384101.2019.1618120
127. Guo J, Duan H, Li Y, Yang L, Yuan L. A novel circular RNA circ-ZNF652 promotes hepatocellular carcinoma metastasis through inducing snail-mediated epithelial-mesenchymal transition by sponging miR-203/miR-502-5p. Biochem Biophys Res Commun. 2019;513:812–819. doi:10.1016/j.bbrc.2019.03.214
128. Xu L, Feng X, Hao X, et al. CircSETD3 (Hsa_circ_0000567) acts as a sponge for microRNA-421 inhibiting hepatocellular carcinoma growth. J Exp Clin Cancer Res. 2019;38:98. doi:10.1186/s13046-019-1041-2
129. Li X, Shen M. Circular RNA hsa_circ_103809 suppresses hepatocellular carcinoma proliferation and invasion by sponging miR-620. Eur Rev Med Pharmacol Sci. 2019;23:555–566. doi:10.26355/eurrev_201902_16868
130. Pan H, Tang L, Jiang H, et al. Enhanced expression of circ_0000267 in hepatocellular carcinoma indicates poor prognosis and facilitates cell progression by sponging miR-646. J Cell Biochem. 2019. doi:10.1002/jcb.28411
131. Zhang J, Chang Y, Xu L, Qin L. Elevated expression of circular RNA circ_0008450 predicts dismal prognosis in hepatocellular carcinoma and regulates cell proliferation, apoptosis, and invasion via sponging miR-548p. J Cell Biochem. 2019;120:9487–9494. doi:10.1002/jcb.v120.6
132. Qiu L, Huang Y, Li Z, et al. Circular RNA profiling identifies circADAMTS13 as a miR-484 sponge which suppresses cell proliferation in hepatocellular carcinoma. Mol Oncol. 2019;13:441–455. doi:10.1002/1878-0261.12424
133. Xie B, Zhao Z, Liu Q, Wang X, Ma Z, Li H. CircRNA has_circ_0078710 acts as the sponge of microRNA-31 involved in hepatocellular carcinoma progression. Gene. 2019;683:253–261. doi:10.1016/j.gene.2018.10.043
134. Cao S, Wang G, Wang J, Li C, Zhang L. Hsa_circ_101280 promotes hepatocellular carcinoma by regulating miR-375/JAK2. Immunol Cell Biol. 2019;97:218–228. doi:10.1111/imcb.2019.97.issue-2
135. Wang H, Chen W, Jin M, et al. CircSLC3A2 functions as an oncogenic factor in hepatocellular carcinoma by sponging miR-490-3p and regulating PPM1F expression. Mol Cancer. 2018;17:165. doi:10.1186/s12943-018-0909-7
136. Zhang PF, Wei CY, Huang XY, et al. Circular RNA circTRIM33-12 acts as the sponge of MicroRNA-191 to suppress hepatocellular carcinoma progression. Mol Cancer. 2019;18:105. doi:10.1186/s12943-019-1031-1
137. Ma Y, Zhang C, Zhang B, Yu H, Yu Q. circRNA of AR-suppressed PABPC1 91 bp enhances the cytotoxicity of natural killer cells against hepatocellular carcinoma via upregulating UL16 binding protein 1. Oncol Lett. 2019;17:388–397. doi:10.3892/ol.2018.9606
138. Wang M, Yu F, Li P. Circular RNAs: characteristics, function and clinical significance in hepatocellular carcinoma. Cancers (Basel). 2018;10:258. doi:10.3390/cancers10080258
139. Fu L, Jiang Z, Li T, Hu Y, Guo J. Circular RNAs in hepatocellular carcinoma: functions and implications. Cancer Med. 2018. doi:10.1002/cam4.1574
140. Wang S, Cui S, Zhao W, et al. Screening and bioinformatics analysis of circular RNA expression profiles in hepatitis B-related hepatocellular carcinoma. Cancer Biomark. 2018;22:631–640. doi:10.3233/CBM-170910
141. Su Y, Xu C, Liu Y, Hu Y, Wu H. Circular RNA hsa_circ_0001649 inhibits hepatocellular carcinoma progression via multiple miRNAs sponge. Aging (Albany NY). 2019. doi:10.18632/aging.101988
142. Ding B, Lou W, Xu L, Fan W. Non-coding RNA in drug resistance of hepatocellular carcinoma. Biosci Rep. 2018;38. doi:10.1042/BSR20180915
143. Yao R, Zou H, Liao W. Prospect of circular RNA in hepatocellular carcinoma: a novel potential biomarker and therapeutic target. Front Oncol. 2018;8:332. doi:10.3389/fonc.2018.00332
144. Qiu LP, Wu YH, Yu XF, Tang Q, Chen L, Chen KP. The emerging role of circular RNAs in hepatocellular carcinoma. J Cancer. 2018;9:1548–1559. doi:10.7150/jca.24566
145. Ferlay J, Soerjomataram I, Dikshit R, et al. Cancer incidence and mortality worldwide: sources, methods and major patterns in GLOBOCAN 2012. Int J Cancer. 2015;136:E359–E386. doi:10.1002/ijc.29210
146. Lodewijk I, Duenas M, Rubio C, et al. Liquid biopsy biomarkers in bladder cancer: a current need for patient diagnosis and monitoring. Int J Mol Sci. 2018;19:2514. doi:10.3390/ijms19092514
147. Geng Y, Jiang J, Wu C. Function and clinical significance of circRNAs in solid tumors. J Hematol Oncol. 2018;11:98. doi:10.1186/s13045-018-0643-z
148. Liu H, Bi J, Dong W, et al. Invasion-related circular RNA circFNDC3B inhibits bladder cancer progression through the miR-1178-3p/G3BP2/SRC/FAK axis. Mol Cancer. 2018;17:161. doi:10.1186/s12943-018-0908-8
149. Wu Z, Huang W, Wang X, et al. Circular RNA CEP128 acts as a sponge of miR-145-5p in promoting the bladder cancer progression via regulating SOX11. Mol Med. 2018;24:40. doi:10.1186/s10020-018-0039-0
150. Xie F, Li Y, Wang M, et al. Circular RNA BCRC-3 suppresses bladder cancer proliferation through miR-182-5p/p27 axis. Mol Cancer. 2018;17:144. doi:10.1186/s12943-018-0892-z
151. Yang C, Wu S, Wu X, Zhou X, Jin S, Jiang H. Silencing circular RNA UVRAG inhibits bladder cancer growth and metastasis by targeting the microRNA-223/fibroblast growth factor receptor 2 axis. Cancer Sci. 2019;110:99–106. doi:10.1111/cas.2019.110.issue-1
152. Bi J, Liu H, Cai Z, et al. Circ-BPTF promotes bladder cancer progression and recurrence through the miR-31-5p/RAB27A axis. Aging (Albany NY). 2018;10:1964–1976. doi:10.18632/aging.v10i8
153. Lin G, Sheng H, Xie H, et al. circLPAR1 is a novel biomarker of prognosis for muscle-invasive bladder cancer with invasion and metastasis by miR-762. Oncol Lett. 2019;17:3537–3547. doi:10.3892/ol.2019.9970
154. Su H, Tao T, Yang Z, et al. Circular RNA cTFRC acts as the sponge of MicroRNA-107 to promote bladder carcinoma progression. Mol Cancer. 2019;18:27. doi:10.1186/s12943-019-0951-0
155. Liu H, Chen D, Bi J, et al. Circular RNA circUBXN7 represses cell growth and invasion by sponging miR-1247-3p to enhance B4GALT3 expression in bladder cancer. Aging (Albany NY). 2018;10:2606–2623. doi:10.18632/aging.v10i10
156. Gu C, Zhou N, Wang Z, et al. circGprc5a promoted bladder oncogenesis and metastasis through Gprc5a-targeting peptide. Mol Ther Nucleic Acids. 2018;13:633–641. doi:10.1016/j.omtn.2018.10.008
157. Guo S, Xu X, Ouyang Y, et al. Microarray expression profile analysis of circular RNAs in pancreatic cancer. Mol Med Rep. 2018;17:7661–7671. doi:10.3892/mmr.2018.8827
158. Chen W, Zheng R, Zeng H, Zhang S. The incidence and mortality of major cancers in China, 2012. Chin J Cancer. 2016;35:73. doi:10.1186/s40880-016-0137-8
159. Wei DM, Jiang MT, Lin P, et al. Potential ceRNA networks involved in autophagy suppression of pancreatic cancer caused by chloroquine diphosphate: a study based on differentially expressed circRNAs, lncRNAs, miRNAs and mRNAs. Int J Oncol. 2019;54:600–626. doi:10.3892/ijo.2018.4660
160. Yang F, Liu DY, Guo JT, et al. Circular RNA circ-LDLRAD3 as a biomarker in diagnosis of pancreatic cancer. World J Gastroenterol. 2017;23:8345–8354. doi:10.3748/wjg.v23.i47.8345
161. Zhu P, Ge N, Liu D, et al. Preliminary investigation of the function of hsa_circ_0006215 in pancreatic cancer. Oncol Lett. 2018;16:603–611. doi:10.3892/ol.2018.8652
162. Chen G, Shi Y, Zhang Y, Sun J. CircRNA_100782 regulates pancreatic carcinoma proliferation through the IL6-STAT3 pathway. Onco Targets Ther. 2017;10:5783–5794. doi:10.2147/OTT
163. An Y, Cai H, Zhang Y, et al. circZMYM2 competed endogenously with miR-335-5p to regulate JMJD2C in pancreatic cancer. Cell Physiol Biochem. 2018;51:2224–2236. doi:10.1159/000495868
164. Qu S, Hao X, Song W, et al. Circular RNA circRHOT1 is upregulated and promotes cell proliferation and invasion in pancreatic cancer. Epigenomics. 2019;11:53–63. doi:10.2217/epi-2018-0051
165. Li Z, Yanfang W, Li J, et al. Tumor-released exosomal circular RNA PDE8A promotes invasive growth via the miR-338/MACC1/MET pathway in pancreatic cancer. Cancer Lett. 2018;432:237–250. doi:10.1016/j.canlet.2018.04.035
166. Li J, Li Z, Jiang P, et al. Circular RNA IARS (circ-IARS) secreted by pancreatic cancer cells and located within exosomes regulates endothelial monolayer permeability to promote tumor metastasis. J Exp Clin Cancer Res. 2018;37:177. doi:10.1186/s13046-018-0822-3
167. Trabzonlu L, Paksoy N. Cytomorphological analysis of thyroid nodules diagnosed as follicular variant of papillary thyroid carcinoma: a fine needle aspiration study of diagnostic clues in 42 cases and the impact of using bethesda system in reporting-an institutional experience. Endocr Pathol. 2018;29:351–356. doi:10.1007/s12022-018-9550-7
168. Yang C, Wei Y, Yu L, Xiao Y. Identification of altered circular RNA expression in serum exosomes from patients with papillary thyroid carcinoma by high-throughput sequencing. Med Sci Monit. 2019;25:2785–2791. doi:10.12659/MSM.915658
169. Lan X, Xu J, Chen C, et al. The landscape of circular RNA expression profiles in papillary thyroid carcinoma based on RNA sequencing. Cell Physiol Biochem. 2018;47:1122–1132. doi:10.1159/000490188
170. Peng N, Shi L, Zhang Q, Hu Y, Wang N, Ye H. Microarray profiling of circular RNAs in human papillary thyroid carcinoma. PLoS One. 2017;12:e0170287. doi:10.1371/journal.pone.0170287
171. Ren H, Liu Z, Liu S, et al. Profile and clinical implication of circular RNAs in human papillary thyroid carcinoma. PeerJ. 2018;6:e5363. doi:10.7717/peerj.5363
172. Cai X, Zhao Z, Dong J, et al. Circular RNA circBACH2 plays a role in papillary thyroid carcinoma by sponging miR-139-5p and regulating LMO4 expression. Cell Death Dis. 2019;10:184. doi:10.1038/s41419-019-1439-y
173. Liu W, Zhao J, Jin M, Zhou M. circRAPGEF5 contributes to papillary thyroid proliferation and metastasis by regulation miR-198/FGFR1. Mol Ther Nucleic Acids. 2019;14:609–616. doi:10.1016/j.omtn.2019.01.003
174. Pan Y, Xu T, Liu Y, Li W, Zhang W. Upregulated circular RNA circ_0025033 promotes papillary thyroid cancer cell proliferation and invasion via sponging miR-1231 and miR-1304. Biochem Biophys Res Commun. 2019;510:334–338. doi:10.1016/j.bbrc.2019.01.108
175. Bi W, Huang J, Nie C, et al. CircRNA circRNA_102171 promotes papillary thyroid cancer progression through modulating CTNNBIP1-dependent activation of beta-catenin pathway. J Exp Clin Cancer Res. 2018;37:275. doi:10.1186/s13046-018-0936-7
176. Wang M, Chen B, Ru Z, Cong L. CircRNA circ-ITCH suppresses papillary thyroid cancer progression through miR-22-3p/CBL/beta-catenin pathway. Biochem Biophys Res Commun. 2018;504:283–288. doi:10.1016/j.bbrc.2018.08.175
177. Wei H, Pan L, Tao D, Li R. Circular RNA circZFR contributes to papillary thyroid cancer cell proliferation and invasion by sponging miR-1261 and facilitating C8orf4 expression. Biochem Biophys Res Commun. 2018;503:56–61. doi:10.1016/j.bbrc.2018.05.174
178. Yao Y, Chen X, Yang H, et al. Hsa_circ_0058124 promotes papillary thyroid cancer tumorigenesis and invasiveness through the NOTCH3/GATAD2A axis. J Exp Clin Cancer Res. 2019;38:318. doi:10.1186/s13046-019-1321-x
179. Lan X, Cao J, Xu J, et al. Decreased expression of hsa_circ_0137287 predicts aggressive clinicopathologic characteristics in papillary thyroid carcinoma. J Clin Lab Anal. 2018;32:e22573. doi:10.1002/jcla.2018.32.issue-8
180. El-Naggar AM, Clarkson PW, Negri GL, et al. HACE1 is a potential tumor suppressor in osteosarcoma. Cell Death Dis. 2019;10:21. doi:10.1038/s41419-018-1276-4
181. Zhao GS, Gao ZR, Zhang Q, et al. TSSC3 promotes autophagy via inactivating the Src-mediated PI3K/Akt/mTOR pathway to suppress tumorigenesis and metastasis in osteosarcoma, and predicts a favorable prognosis. J Exp Clin Cancer Res. 2018;37:188. doi:10.1186/s13046-018-0856-6
182. Zuo D, Shogren KL, Zang J, et al. Inhibition of STAT3 blocks protein synthesis and tumor metastasis in osteosarcoma cells. J Exp Clin Cancer Res. 2018;37:244. doi:10.1186/s13046-018-0914-0
183. Wang C, Ren M, Zhao X, Wang A, Wang J. Emerging roles of circular RNAs in osteosarcoma. Med Sci Monit. 2018;24:7043–7050. doi:10.12659/MSM.912092
184. Xi Y, Fowdur M, Liu Y, Wu H, He M, Zhao J. Differential expression and bioinformatics analysis of circRNA in osteosarcoma. Biosci Rep. 2019;39. doi:10.1042/BSR20181514
185. Zhu K, Niu L, Wang J, et al. Circular RNA hsa_circ_0000885 levels are increased in tissue and serum samples from patients with osteosarcoma. Med Sci Monit. 2019;25:1499–1505. doi:10.12659/MSM.914899
186. Liu G, Huang K, Jie Z, et al. CircFAT1 sponges miR-375 to promote the expression of yes-associated protein 1 in osteosarcoma cells. Mol Cancer. 2018;17:170. doi:10.1186/s12943-018-0917-7
187. Xu B, Yang T, Wang Z, Zhang Y, Liu S, Shen M. CircRNA CDR1as/miR-7 signals promote tumor growth of osteosarcoma with a potential therapeutic and diagnostic value. Cancer Manag Res. 2018;10:4871–4880. doi:10.2147/CMAR
188. Wu Y, Xie Z, Chen J, et al. Circular RNA circTADA2A promotes osteosarcoma progression and metastasis by sponging miR-203a-3p and regulating CREB3 expression. Mol Cancer. 2019;18:73. doi:10.1186/s12943-019-1007-1
189. Kun-Peng Z, Chun-Lin Z, Jian-Ping H, Lei Z. A novel circulating hsa_circ_0081001 act as a potential biomarker for diagnosis and prognosis of osteosarcoma. Int J Biol Sci. 2018;14:1513–1520. doi:10.7150/ijbs.27523
190. Zhang H, Yan J, Lang X, Zhuang Y. Expression of circ_001569 is upregulated in osteosarcoma and promotes cell proliferation and cisplatin resistance by activating the Wnt/beta-catenin signaling pathway. Oncol Lett. 2018;16:5856–5862. doi:10.3892/ol.2018.9410
191. Li X, Diao H. Circular RNA circ_0001946 acts as a competing endogenous RNA to inhibit glioblastoma progression by modulating miR-671-5p and CDR1. J Cell Physiol. 2019;234:13807–13819. doi:10.1002/jcp.28061
192. Wang R, Zhang S, Chen X, et al. EIF4A3-induced circular RNA MMP9 (circMMP9) acts as a sponge of miR-124 and promotes glioblastoma multiforme cell tumorigenesis. Mol Cancer. 2018;17:166. doi:10.1186/s12943-018-0911-0
193. Wang R, Zhang S, Chen X, et al. CircNT5E acts as a sponge of miR-422a to promote glioblastoma tumorigenesis. Cancer Res. 2018;78:4812–4825. doi:10.1158/0008-5472.CAN-18-0532
194. Zhang G, Sun W, Zhu L, Feng Y, Wu L, Li T. Overexpressed circ_0029426 in glioblastoma forecasts unfavorable prognosis and promotes cell progression by sponging miR-197. J Cell Biochem. 2019;120:10295–10302. doi:10.1002/jcb.v120.6
195. Wang HX, Huang QL, Shen JY, et al. Expression profile of circular RNAs in IDH-wild type glioblastoma tissues. Clin Neurol Neurosurg. 2018;171:168–173. doi:10.1016/j.clineuro.2018.06.020
196. Begum S, Yiu A, Stebbing J, Castellano L. Novel tumour suppressive protein encoded by circular RNA, circ-SHPRH, in glioblastomas. Oncogene. 2018;37:4055–4057. doi:10.1038/s41388-018-0230-3
197. Zhang M, Huang N, Yang X, et al. A novel protein encoded by the circular form of the SHPRH gene suppresses glioma tumorigenesis. Oncogene. 2018;37:1805–1814. doi:10.1038/s41388-017-0019-9
198. Shabaninejad Z, Vafadar A, Movahedpour A, et al. Circular RNAs in cancer: new insights into functions and implications in ovarian cancer. J Ovarian Res. 2019;12:84. doi:10.1186/s13048-019-0558-5
199. Guan X, Zong ZH, Liu Y, Chen S, Wang LL, Zhao Y. circPUM1 promotes tumorigenesis and progression of ovarian cancer by sponging miR-615-5p and miR-6753-5p. Mol Ther Nucleic Acids. 2019;18:882–892. doi:10.1016/j.omtn.2019.09.032
200. Zong ZH, Du YP, Guan X, Chen S, Zhao Y. CircWHSC1 promotes ovarian cancer progression by regulating MUC1 and hTERT through sponging miR-145 and miR-1182. J Exp Clin Cancer Res. 2019;38:437. doi:10.1186/s13046-019-1437-z
201. Sheng M, Wei N, Yang HY, Yan M, Zhao QX, Jing LJ. CircRNA UBAP2 promotes the progression of ovarian cancer by sponging microRNA-144. Eur Rev Med Pharmacol Sci. 2019;23:7283–7294. doi:10.26355/eurrev_201909_18833
202. Chen H, Mao M, Jiang J, Zhu D, Li P. Circular RNA CDR1as acts as a sponge of miR-135b-5p to suppress ovarian cancer progression. Onco Targets Ther. 2019;12:3869–3879. doi:10.2147/OTT.S207938
203. Li QH, Liu Y, Chen S, et al. circ-CSPP1 promotes proliferation, invasion and migration of ovarian cancer cells by acting as a miR-1236-3p sponge. Biomed Pharmacother. 2019;114:108832. doi:10.1016/j.biopha.2019.108832
204. Zhang M, Xia B, Xu Y, et al. (hsa_circ_0051240) promotes cell proliferation, migration and invasion in ovarian cancer through miR-637/KLK4 axis. Artif Cells Nanomed Biotechnol. 2019;47:1224–1233. doi:10.1080/21691401.2019.1593999
205. Xie J, Wang S, Li G, et al. circEPSTI1 regulates ovarian cancer progression via decoying miR-942. J Cell Mol Med. 2019;23:3597–3602. doi:10.1111/jcmm.2019.23.issue-5
206. Luo L, Gao YQ, Sun XF. Circular RNA ITCH suppresses proliferation and promotes apoptosis in human epithelial ovarian cancer cells by sponging miR-10a-alpha. Eur Rev Med Pharmacol Sci. 2018;22:8119–8126. doi:10.26355/eurrev_201812_16503
207. Chen Q, Zhang J, He Y, Wang Y. hsa_circ_0061140 knockdown reverses FOXM1-mediated cell growth and metastasis in ovarian cancer through miR-370 sponge activity. Mol Ther Nucleic Acids. 2018;13:55–63. doi:10.1016/j.omtn.2018.08.010
208. Liu J, Yu F, Wang S, et al. circGFRA1 promotes ovarian cancer progression by sponging miR-449a. J Cancer. 2019;10:3908–3913. doi:10.7150/jca.31615
209. Wang W, Wang J, Zhang X, Liu G. Serum circSETDB1 is a promising biomarker for predicting response to platinum-taxane-combined chemotherapy and relapse in high-grade serous ovarian cancer. Onco Targets Ther. 2019;12:7451–7457. doi:10.2147/OTT.S220700
210. Sun D, Liu J, Zhou L. Upregulation of circular RNA circFAM53B predicts adverse prognosis and accelerates the progression of ovarian cancer via the miR646/VAMP2 and miR647/MDM2 signaling pathways. Oncol Rep. 2019;42:2728–2737. doi:10.3892/or.2019.7366
211. Zhang L, Zhou Q, Qiu Q, et al. CircPLEKHM3 acts as a tumor suppressor through regulation of the miR-9/BRCA1/DNAJB6/KLF4/AKT1 axis in ovarian cancer. Mol Cancer. 2019;18:144. doi:10.1186/s12943-019-1080-5
212. Zhao Y, Qin XP, Lang YP, Kou D, Shao ZW. Circular RNA circ-SMAD7 promoted ovarian cancer cell proliferation and metastasis by suppressing KLF6. Eur Rev Med Pharmacol Sci. 2019;23:5603–5610. doi:10.26355/eurrev_201907_18294
213. Zhang C, Xiong J, Yang Q, et al. Profiling and bioinformatics analyses of differential circular RNA expression in prostate cancer cells. Future Sci OA. 2018;4:FSOA340. doi:10.4155/fsoa-2018-0046
214. Xia Q, Ding T, Zhang G, et al. Circular RNA expression profiling identifies prostate cancer- specific circRNAs in prostate cancer. Cell Physiol Biochem. 2018;50:1903–1915. doi:10.1159/000494870
215. Yan Z, Xiao Y, Chen Y, Luo G. Screening and identification of epithelial-to-mesenchymal transition-related circRNA and miRNA in prostate cancer. Pathol Res Pract. 2020;216(2):152784.
216. Wang X, Wang R, Wu Z, Bai P. Circular RNA ITCH suppressed prostate cancer progression by increasing HOXB13 expression via spongy miR-17-5p. Cancer Cell Int. 2019;19:328. doi:10.1186/s12935-019-0994-8
217. Xiang Z, Xu C, Wu G, Liu B, Wu D. CircRNA-UCK2 increased TET1 inhibits proliferation and invasion of prostate cancer cells via sponge MiRNA-767-5p. Open Med (Wars). 2019;14:833–842. doi:10.1515/med-2019-0097
218. Kong Z, Wan X, Lu Y, et al. Circular RNA circFOXO3 promotes prostate cancer progression through sponging miR-29a-3p. J Cell Mol Med. 2020;24:799–813. doi:10.1111/jcmm.v24.1
219. Huang C, Deng H, Wang Y, et al. Circular RNA circABCC4 as the ceRNA of miR-1182 facilitates prostate cancer progression by promoting FOXP4 expression. J Cell Mol Med. 2019;23:6112–6119. doi:10.1111/jcmm.v23.9
220. Cai C, Zhi Y, Wang K, et al. CircHIPK3 overexpression accelerates the proliferation and invasion of prostate cancer cells through regulating miRNA-338-3p. Onco Targets Ther. 2019;12:3363–3372. doi:10.2147/OTT.S196931
221. Chen D, Lu X, Yang F, Xing N. Circular RNA circHIPK3 promotes cell proliferation and invasion of prostate cancer by sponging miR-193a-3p and regulating MCL1 expression. Cancer Manag Res. 2019;11:1415–1423. doi:10.2147/CMAR.S190669
222. Yang Z, Qu CB, Zhang Y, et al. Dysregulation of p53-RBM25-mediated circAMOTL1L biogenesis contributes to prostate cancer progression through the circAMOTL1L-miR-193a-5p-Pcdha pathway. Oncogene. 2019;38:2516–2532. doi:10.1038/s41388-018-0602-8
223. Zheng Y, Chen CJ, Lin ZY, et al. Circ_KATNAL1 regulates prostate cancer cell growth and invasion through miR-145-3p/WISP1 pathway. Biochem Cell Biol. 2019. doi:10.1139/bcb-2019-0211
224. Feng Y, Yang Y, Zhao X, et al. Circular RNA circ0005276 promotes the proliferation and migration of prostate cancer cells by interacting with FUS to transcriptionally activate XIAP. Cell Death Dis. 2019;10:792. doi:10.1038/s41419-019-2028-9
225. Jamal M, Song T, Chen B, et al. Recent progress on circular RNA research in acute myeloid leukemia. Front Oncol. 2019;9:1108. doi:10.3389/fonc.2019.01108
226. Liu Y, Cheng Z, Pang Y, et al. Role of microRNAs, circRNAs and long noncoding RNAs in acute myeloid leukemia. J Hematol Oncol. 2019;12:51. doi:10.1186/s13045-019-0734-5
227. Li M, Meng F, Lu Q. Expression profile screening and bioinformatics analysis of circrna, lncRNA and mRNA in acute myeloid leukemia drug-resistance cells. Turk J Haematol. 2019. doi:10.4274/tjh
228. Yuan DM, Ma J, Fang WB. Identification of non-coding RNA regulatory networks in pediatric acute myeloid leukemia reveals circ-0004136 could promote cell proliferation by sponging miR-142. Eur Rev Med Pharmacol Sci. 2019;23:9251–9258. doi:10.26355/eurrev_201911_19417
229. Ping L, Jian-Jun C, Chu-Shu L, Guang-Hua L, Ming Z. Silencing of circ_0009910 inhibits acute myeloid leukemia cell growth through increasing miR-20a-5p. Blood Cells Mol Dis. 2019;75:41–47. doi:10.1016/j.bcmd.2018.12.006
230. Fan H, Li Y, Liu C, Liu Y, Bai J, Li W. Circular RNA-100290 promotes cell proliferation and inhibits apoptosis in acute myeloid leukemia cells via sponging miR-203. Biochem Biophys Res Commun. 2018;507:178–184. doi:10.1016/j.bbrc.2018.11.002
231. Chen H, Liu T, Liu J, et al. Circ-ANAPC7 is upregulated in acute myeloid leukemia and appears to target the MiR-181 family. Cell Physiol Biochem. 2018;47:1998–2007. doi:10.1159/000491468
232. Yang W, Du WW, Li X, Yee AJ, Yang BB. Foxo3 activity promoted by non-coding effects of circular RNA and Foxo3 pseudogene in the inhibition of tumor growth and angiogenesis. Oncogene. 2016;35:3919–3931. doi:10.1038/onc.2015.460
233. Zhou J, Zhou LY, Tang X, et al. Circ-Foxo3 is positively associated with the Foxo3 gene and leads to better prognosis of acute myeloid leukemia patients. BMC Cancer. 2019;19:930. doi:10.1186/s12885-019-5967-8
234. Shang J, Chen WM, Liu S, et al. CircPAN3 contributes to drug resistance in acute myeloid leukemia through regulation of autophagy. Leuk Res. 2019;85:106198. doi:10.1016/j.leukres.2019.106198
235. Feng XQ, Nie SM, Huang JX, et al. Circular RNA circHIPK3 serves as a prognostic marker to promote chronic myeloid leukemia progression. Neoplasma. 2019.
236. Ping L, Jian-Jun C, Chu-Shu L, Guang-Hua L, Ming Z. High circ_100053 predicts a poor outcome for chronic myeloid leukemia and is involved in imatinib resistance. Oncol Res. 2019. doi:10.3727/096504018X15412701483326
237. Liu J, Kong F, Lou S, Yang D, Gu L. Global identification of circular RNAs in chronic myeloid leukemia reveals hsa_circ_0080145 regulates cell proliferation by sponging miR-29b. Biochem Biophys Res Commun. 2018;504:660–665. doi:10.1016/j.bbrc.2018.08.154
238. Su M, Xiao Y, Ma J, et al. Circular RNAs in cancer: emerging functions in hallmarks, stemness, resistance and roles as potential biomarkers. Mol Cancer. 2019;18:90. doi:10.1186/s12943-019-1002-6
239. Yan N, Xu H, Zhang J, et al. Circular RNA profile indicates circular RNA VRK1 is negatively related with breast cancer stem cells. Oncotarget. 2017;8:95704–95718. doi:10.18632/oncotarget.v8i56
240. Liu R, Shi P, Nie Z, et al. Mifepristone suppresses basal triple-negative breast cancer stem cells by down-regulating KLF5 expression. Theranostics. 2016;6:533–544. doi:10.7150/thno.14315
241. Cherubini A, Barilani M, Rossi RL, et al. FOXP1 circular RNA sustains mesenchymal stem cell identity via microRNA inhibition. Nucleic Acids Res. 2019;47:5325–5340. doi:10.1093/nar/gkz199
242. Zhu YJ, Zheng B, Luo GJ, et al. Circular RNAs negatively regulate cancer stem cells by physically binding FMRP against CCAR1 complex in hepatocellular carcinoma. Theranostics. 2019;9:3526–3540. doi:10.7150/thno.32796
243. Wu Y, Zhang Y, Niu M, et al. Whole-transcriptome analysis of CD133+CD144+ cancer stem cells derived from human laryngeal squamous cell carcinoma cells. Cell Physiol Biochem. 2018;47:1696–1710. doi:10.1159/000490992
244. Feng Z, Meng S, Zhou H, et al. Functions and potential applications of circular RNAs in cancer stem cells. Front Oncol. 2019;9:500. doi:10.3389/fonc.2019.00500
245. Shi P, Sun J, He B, et al. Profiles of differentially expressed circRNAs in esophageal and breast cancer. Cancer Manag Res. 2018;10:2207–2221. doi:10.2147/CMAR
246. Wu J, Qi X, Liu L, et al. Emerging epigenetic regulation of circular RNAs in human cancer. Mol Ther Nucleic Acids. 2019;16:589–596. doi:10.1016/j.omtn.2019.04.011
247. Liu W, Zhang J, Zou C, et al. Microarray expression profile and functional analysis of circular RNAs in osteosarcoma. Cell Physiol Biochem. 2017;43:969–985. doi:10.1159/000481650
248. Fanale D, Taverna S, Russo A, Bazan V. Circular RNA in exosomes. Adv Exp Med Biol. 2018;1087:109–117.
249. Chen H, Liu S, Li M, Huang P, Li X. circ_0003418 inhibits tumorigenesis and cisplatin chemoresistance through Wnt/beta-Catenin pathway in hepatocellular carcinoma. Onco Targets Ther. 2019;12:9539–9549. doi:10.2147/OTT.S229507
250. Di X, Jin X, Li R, Zhao M, Wang K. CircRNAs and lung cancer: biomarkers and master regulators. Life Sci. 2019;220:177–185. doi:10.1016/j.lfs.2019.01.055
251. Yu W, Peng W, Sha H, Li J. Hsa_circ_0003998 promotes chemoresistance via modulation of miR-326 in lung adenocarcinoma cells. Oncol Res. 2019;27:623–628. doi:10.3727/096504018X15420734828058
252. Xu QY, Xie MJ, Huang J, Wang ZW. Effect of circ MTHFD2 on resistance to pemetrexed in gastric cancer through regulating expression of miR-124. Eur Rev Med Pharmacol Sci. 2019;23:10290–10299. doi:10.26355/eurrev_201912_19667
253. Huang X, Li Z, Zhang Q, et al. Circular RNA AKT3 upregulates PIK3R1 to enhance cisplatin resistance in gastric cancer via miR-198 suppression. Mol Cancer. 2019;18:71. doi:10.1186/s12943-019-0969-3
254. Yang W, Gu J, Wang X, et al. Inhibition of circular RNA CDR1as increases chemosensitivity of 5-FU-resistant BC cells through up-regulating miR-7. J Cell Mol Med. 2019;23:3166–3177. doi:10.1111/jcmm.2019.23.issue-5
255. Chi BJ, Zhao DM, Liu L, et al. Downregulation of hsa_circ_0000285 serves as a prognostic biomarker for bladder cancer and is involved in cisplatin resistance. Neoplasma. 2019;66:197–202. doi:10.4149/neo_2018_180318N185
256. Gao M, Li C, Xiao H, et al. hsa_circ_0007841: a novel potential biomarker and drug resistance for multiple myeloma. Front Oncol. 2019;9:1261. doi:10.3389/fonc.2019.01261
257. Greene J, Baird AM, Casey O, et al. Circular RNAs are differentially expressed in prostate cancer and are potentially associated with resistance to enzalutamide. Sci Rep. 2019;9:10739. doi:10.1038/s41598-019-47189-2
258. Su Y, Yang W, Jiang N, et al. Hypoxia-elevated circELP3 contributes to bladder cancer progression and cisplatin resistance. Int J Biol Sci. 2019;15:441–452. doi:10.7150/ijbs.26826
259. Hua X, Sun Y, Chen J, et al. Circular RNAs in drug resistant tumors. Biomed Pharmacother. 2019;118:109233. doi:10.1016/j.biopha.2019.109233
260. Szabo L, Salzman J. Detecting circular RNAs: bioinformatic and experimental challenges. Nat Rev Genet. 2016;17:679–692. doi:10.1038/nrg.2016.114
261. Wang Y, Liu J, Ma J, et al. Exosomal circRNAs: biogenesis, effect and application in human diseases. Mol Cancer. 2019;18:116. doi:10.1186/s12943-019-1041-z
262. Soneson C, Yao Y, Bratus-neuenschwander A, Patrignani A, Robinson MD, Hussain S. A comprehensive examination of Nanopore native RNA sequencing for characterization of complex transcriptomes. Nat Commun. 2019;10:3359. doi:10.1038/s41467-019-11272-z
263. Warren S. Simultaneous, multiplexed detection of RNA and protein on the NanoString((R)) nCounter((R)) Platform. Methods Mol Biol. 2018;1783:105–120.
 © 2020 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.
© 2020 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.