Back to Journals » Hepatic Medicine: Evidence and Research » Volume 13
Binding of the SARS-CoV-2 Spike Protein to the Asialoglycoprotein Receptor on Human Primary Hepatocytes and Immortalized Hepatocyte-Like Cells by Confocal Analysis
Authors Collins DP , Steer CJ
Received 14 January 2021
Accepted for publication 20 March 2021
Published 14 April 2021 Volume 2021:13 Pages 37—44
DOI https://doi.org/10.2147/HMER.S301979
Checked for plagiarism Yes
Review by Single anonymous peer review
Peer reviewer comments 2
Editor who approved publication: Dr Gerry Lake-Bakaar
Daniel P Collins,1 Clifford J Steer2,3
1CMDG, LLC, Saint Paul, MN, USA; 2Department of Medicine, University of Minnesota Medical School, Minneapolis, MN, USA; 3Department of Genetics, Cell Biology and Development, University of Minnesota Medical School, Minneapolis, MN, USA
Correspondence: Daniel P Collins Email [email protected]
Background: The SARS-CoV-2 virus may have direct or indirect effects on other human organs beyond the respiratory system and including the liver, via binding of the spike protein. This study investigated the potential direct interactions with the liver by comparing the binding of SARS-CoV-2 spike proteins to human AT2-like cells, primary human hepatocytes and immortalized hepatocyte-like hybrid cells. Receptors with binding specificity for SARS-CoV-2 spike protein on AT2 cells and hepatocytes were identified.
Methods: The specific binding of biotinylated spike and spike 1 proteins to undifferentiated human E12 MLPC (E12), E12 differentiated alveolar type 2 (AT2) cells, primary human hepatocytes (PHH) and E12 human hepatocyte-like hybrid cells (HLC) was studied by confocal microscopy. We investigated the expression of ACE-2, binding of biotinylated spike protein, biotinylated spike 1 and inhibition of binding by unlabeled spike protein, two neutralizing antibodies and an antibody directed against the hepatocyte asialoglycoprotein receptor 1 (ASGr1).
Results: E12 MLPC did not express ACE-2 and did not bind either of spike or spike 1 proteins. AT2-like cells expressed ACE-2 and bound both spike and spike 1. Both PHH and HLC did not express ACE-2 and did not bind spike 1 protein. However, both PHH and HLC actively bound the spike protein. Biotinylated spike protein binding was inhibited by unlabeled spike but not spike 1 protein on PHH and HLC. Two commercial neutralizing antibodies blocked the binding of the spike to PHH and HLC but only one blocked binding to AT2. An antibody to the hepatocyte ASGr1 blocked the binding of the spike protein to PHH and HLC.
Conclusion: The absence of ACE-2 receptors and inhibition of spike binding by an antibody to the ASGr1 on both PHH and HLC suggested that the spike protein interacts with the ASGr1. The differential antibody blocking of spike binding to AT2, PHH and HLC indicated that neutralizing activity of SARS-CoV-2 binding might involve additional mechanisms beyond RBD binding to ACE-2.
Keywords: asialoglycoprotein receptor, E12 MLPC, SARS-CoV-2, AT2, human hepatocytes, spike proteins
Introduction
SARS-CoV-2 is the virus responsible for the COVID-19 pandemic and its damaging effects on both health and economics worldwide. Transmission and pathology of the virus appears to be mediated through the respiratory system via interaction of the viral spike protein with the ACE-2 receptor1–3 differentially presented on various cells within the respiratory system.4,5 Expression of ACE-2 has been reported on lung alveolar epithelial cells, enterocytes of the small intestine, circulatory endothelial cells, arterial smooth muscle cells,6 adipose tissue, bone marrow, duodenum, endometrium, heart, kidney, testis, and thyroid7 suggesting a potential direct effect of COVID-19 on those tissues. Additionally, infection with COVID-19 has been associated with significant liver injuries and altered liver function tests.8
Alterations in liver function have been attributed to secondary effects of cytokine cascade, hypoxia, underlying liver disease,9–11 or infection of ACE-2 positive cholangiocytes.12 There have also been reports of coronavirus particles in hepatocytes without a defined mechanism for infection.13
In a recent report studying the receptome of spike binding, ACE-2 was confirmed as the primary receptor for the spike protein via the binding domain (RBD) on the spike 1 portion of the molecule and the N-terminal-domain as the sites critical for virus–host interaction.14 Additionally, the report described binding of the spike protein with ectopically expressed ASGR1 and KREMEN1 in transfected non-liver cells. The results strongly suggested the existence of additional entry points into cells for the SARS-CoV-2 virus via the spike protein. Differences in primary infection sites and clinical manifestations of SARS-CoV and SARS-CoV-2, both utilizing ACE-2 as the primary site of cellular infection, suggested that other cellular receptors may be involved in SARS-CoV-2 host interactions.14
E12 TERT-immortalized multi-lineage progenitor cells (MLPC) derived from human umbilical cord blood have been differentiated into immortalized AT2-like cells (AT2) (manuscript submitted) and fused directly with primary human hepatocytes to create immortalized hepatocyte-like cells (HLC).15 The resultant E12 AT2-like cells expressed the characteristics of small airway epithelial cells associated with alveolar type 2 cells and not alveolar type 1 cells. The E12/PHH fusion cells (HLC) expressed the characteristics of fully mature and highly differentiated hepatocytes.15
This report studied the interactions of the SARS-CoV-2 spike protein with potential receptors on human cord blood-derived MLPC differentiated AT2, HLC and primary human hepatocytes (PHH) by confocal analysis. The characteristics of spike protein binding were examined using biotinylated spike proteins and blockade of binding by un-labeled spike proteins, spike protein-directed neutralizing antibodies and an antibody directed against the hepatocyte surface membrane asialoglycoprotein receptor 1 (ASGr1). The results suggested that binding and inhibition analyses can be used to assess the potential mechanisms of viral host cell interactions with a myriad of different target cells in the body, but also to assess therapeutics designed to inhibit that binding.
Immortalized AT2 and HLC could provide accurate and reproducible tools to study the differential virus–host interactions between these targets of COVID-19 infection and aid in the development of therapeutics designed to inhibit binding and infection by the SARS-CoV-2 virus. In addition, the potential binding of spike protein to the ASGr1 on hepatocytes suggested a mechanism of viral entry via the clathrin-coated pit receptor-mediated pathway and direct injury to the liver.16,17
Methods and Materials
E12 Multi-Lineage Progenitor Cells (MLPC)
MLPC are multi-potent non-hematopoietic stem cells isolated from human umbilical cord blood.15 Umbilical cord blood was collected as part of an FDA submission to market PrepaCyte-CB, a product to de-bulk cord blood for cryo-banking and transplantation. IRB approval of the studies was conducted by the University of Minnesota, the Saint Louis Cord Blood Bank and by Quorum Review Protocol #800, March 3, 2005. The cord blood samples were collected by the American Red Cross Cord Blood Program (Saint Paul, Minnesota) and Ridgeview Medical Center (Waconia, MN). Donations were collected with donor consent for research use only.
Briefly, isolated leukocytes were incubated overnight in MSCGM (PT-4105, Lonza, Walkerville, MD) after which non-adherent cells were removed. Cells were cultured in MSCGM until 80–90% of cells had a fibroblastic morphology. These cells were transfected with the gene for TERT, as previously described15 and were cloned by limited dilution. The E12 clone was selected for both immortality and differentiating potential. The E12 MLPC, expanded and cryopreserved for over 14 years, were used as undifferentiated control cells and as the source of cells for the development of the AT2-like cells and the fusion partner in the development of MLPC/hepatocyte hybrid cells.15 For confocal analysis, E12 cells (106/mL in MSCGM, 200 μL per well) were plated in non-coated 16 well chamber slides (Nalge, Nunc International, Rochester, NY) and allowed to attach overnight before use in the analysis.
Alveolar Type 2-Like Cells (AT2)
AT2-like cells were developed from the differentiation of E12 MLPC. Briefly, E12 cells (3 x 105 cells/mL) in MSCGM were added to non-coated tissue culture vessels and allowed to attach overnight. Medium was then exchanged with SAGM (SAGM, Lonza, Walkerville, MD, cat # 3118) and allowed to culture for 8–14 days with 3 medium changes per week. Upon achieving 70% confluence, cells were harvested by treatment with Tryp-LE (12605–028, Life Technologies, Grand Island, NY) allowed to dissociate from the culture vessel and used for confocal analysis, as a positive control for binding spike proteins and ACE-2 expression. Cells (106/mL in SAGM, 200 μL per well) were plated in non-coated 16 well chamber slides and allowed to adhere overnight prior to confocal analysis.
Hepatocyte-Like Cells (HLC)
Hepatocyte-like fusion cells were created by the fusion of E12 MLPC with primary human hepatocytes, as previously described.15 Equal numbers of E12 MLPC and primary hepatocytes were fused using 50% polyethylene glycol in RMPI + 0.01% EDTA. Resultant cells were plated into collagen-coated 75 cm2 tissue culture flasks and were cultured for 7 days in RPMI + 20% FBS. After 7 days, non-fused PHH were no longer viable and did not contribute to the HLC cell lines. HLC were examined for hepatocyte-specific markers including albumin and urea production. HLC were demonstrated to express markers and production consistent with fully mature and well-differentiated hepatocytes. HLC (106/mL in hepatocyte expansion medium, 200 μL per well) were plated in collagen-coated 16 well chamber slides and were allowed to adhere overnight prior to confocal analysis. Hepatocyte expansion medium consisted of Williams Medium E supplemented with 2% fatty acid-free BSA (Sigma, A7030), 1% ITS solution (Lonza, 17–838Z), 5mM hydrocortisone 21-hemisuccinate (Sigma, H2270) and glutamax (35050, Gibco) supplemented with FGF basic (20 ng/mL) (233-FB), FGF-4 (20 ng/mL) (7460-F4), HFG (40 ng/mL) (294-HG), SCF (40 ng/mL) (255-SC), Oncostatin M (20 ng/mL) (295-OM), BMP-4 (20 ng/mL) (314-BP), EGF (40 ng/mL) (236-EG) and IL-1β (20 ng/mL)(201-LB) all from R&D Systems (Minneapolis, MN).
Primary Human Hepatocytes (PHH)
Cryo-preserved primary human hepatocytes and media were obtained from Zenotech (Kansas City, KS). Cells were thawed with OptiThaw medium and enumerated with OptiCount medium in a standard hemacytometer. Hepatocytes were diluted to a final concentration of 106 cells/mL of OptiPlate medium and were plated in collagen-coated 16 well chamber slides at 200 μL per well. After 4 hours of plating, the medium was changed to OptiCulture medium to allow overnight attachment and spread of cells prior to confocal analysis.
Confocal Immunofluorescent Analysis
Cells were prepared for staining with antibodies and binding of spike proteins by fixing the cells in 1% formaldehyde for 1 hour. Cells were then washed x 2 with PermaCyte permeabilization medium (WBP-1000, CMDG, St. Paul, MN). All staining took place in the presence of PermaCyte. Cells were incubated with an unlabeled primary antibody (100 ng) for 30 minutes at room temperature. ACE-2 (labeled with alexa 594, FAB9332T), albumin (MAB1456) and asialoglycoprotein receptor 1 (MAB4394) antibodies were obtained from R&D Systems (Minneapolis, MN). Unbound antibody was removed by washing with PermaCyte and the cells were counterstained with a secondary antibody specific for mouse (A-11005) antibody labelled with Alexa 594 dye (Life Technologies, (Eugene, OR)). Marker expression was confirmed by positive staining when compared to cells stained with antibody isotype controls (QTC1000, CMDG, St. Paul, MN). The nuclei of the cells were visualized by staining with DAPI.
The binding of SARS-CoV-2 spike and spike 1 proteins was analyzed by confocal microscopy using biotinylated spike proteins. Cells were prepared as described above. Cells were labelled with 250 ng of either biotinylated spike (RBD) (SPD-C8E9, ACROBiosystems, Newark, DE) or biotinylated spike 1 protein (SIN-C82E8, ACROBiosystems) for 30 minutes. Unbound spike proteins were removed by washing cells twice with PermaCyte medium. Bound spike proteins were visualized by secondary staining with streptavidin-alexa 594 (S11227 Life Technologies). Cells were counterstained with DAPI to visualize the nuclei.
Specificity of biotinylated spike proteins binding to the cells was confirmed by blockade of binding by a 5 molar excess of unlabeled spike protein. Cells were prepared as per the confocal analysis of antibody binding. Cells were incubated with 1.25 μg of unlabeled spike protein (ACROBiosystems, SPD-S52H6) or spike 1 protein (ACROBiosystems, S1N-C52H3) for 1 hour. Without washing the unbound unlabeled spike protein, biotinylated spike and spike 1 proteins were added to the cells and incubated for 30 minutes. Cells were washed twice with PermaCyte medium to remove any unbound proteins. Bound biotinylated spike proteins were observed by secondary labeling with streptavidin-alexa 594. Cells were counterstained with DAPI to visualize the nucleus.
The effects of antibodies on the binding of the spike proteins to the cells were examined using two commercially available neutralizing antibodies obtained from ACROBiosystems (SAD-S35) and Novatein Biosystems (PR-nCOV-mABS1, Boston, MA) and the ASGr1-specific antibody (R&D Systems). One μg of either neutralizing antibody was preincubated with the spike protein for one hour prior to the addition of the mixture to the cells prepared as described for binding of the spike proteins. The ASGr1 antibody (300 ng) was preincubated with cells prior to the addition of the spike protein. Visualization of the binding of biotinylated spike protein was accomplished by secondary staining with streptavidin-alexa 594. The nuclei of the cells were visualized with DAPI.
Cells were analyzed on the Olympus Fluoview 1000 confocal microscope. The confocal images in Figures 1–4 are representative of at least 3 studies done on different days.
Results
Binding of Spike Proteins to AT2-Like Cells
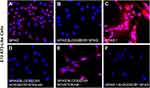
In a parallel study that surveyed the differentiation of E12 MLPC to AT2-like cells, it was demonstrated that AT2-like cells were positive for markers associated with AT2 cells (surfactant protein C, ACE2, TM4SF1, HT2-280), negative for markers associated with AT1 cells (AGER, caveolin 1 and aquaporin) and positive for markers not unique to AT2 cells but known to be expressed on AT2 cells (CK19, CD26 and EpCAM). These results were identical to primary small airway epithelial cells. Both cell types were also shown to bind spike and spike 1 proteins. Biotinylated spike proteins could be blocked by unlabeled spike protein (RBD). Pre-incubation with neutralizing antibodies prevented the binding of biotinylated spike protein by the ACROBiosystems neutralizing antibody, but not the Novatein antibody. Expressions of ACE-2, spike protein binding and inhibition were repeated for this study to confirm the involvement of the ACE-2 receptor (Figure 1).
Expression of ACE-2, ASGr1 and Albumin
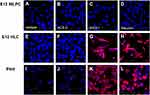
The expressions of ACE-2, ASGr1 and albumin in control E12 MLPC, HLC and PHH were studied by antibody staining. E12 MLPC were shown to be negative for ACE-2, ASGr1 and albumin expression. In contrast, ASGr1 and albumin were shown to be strongly expressed by both HLC and PHH. ACE-2 was not detectible in either cell type (Figure 2).
Binding of Spike Protein
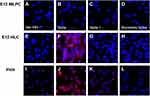
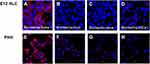
The ability of E12 MLPC, HLC and PHH to bind spike and spike 1 proteins was studied using biotinylated spike proteins. E12 MLPC were unable to bind either spike or spike 1 proteins. HLC and PHH were able to bind spike protein but not spike 1 protein. The binding of biotinylated spike protein could be blocked by pre-incubation with unlabeled spike protein (Figure 3). The binding of biotinylated spike protein could not be blocked by spike 1, but could be blocked by ACROBiosystems and Novatein neutralizing antibodies and also an antibody directed against the ASGr1 (Figure 4).
Discussion
It is critical to elucidate the mechanisms of virus/host interactions of the SARS-CoV-2 for the development of therapeutics designed to inhibit the binding and internalization of the virus to a myriad of cell types. The overarching strategy for the development of vaccines or therapeutics has involved the interaction between the S1 portion of the viral spike protein and the ACE-2 cellular receptor found in the respiratory tract and in various other tissues.4–7 Differences in transmission, pathology and organ involvement between SARS-CoV and SARS-CoV-2 (both dependent upon ACE-2 binding) suggested that additional receptors may contribute to the attachment and internalization of the SARS-CoV-2 virus14 in both the respiratory lungs and other organ systems.
The potential infection of tissues that are ACE-2 negative has spurred the search for additional receptor interactions of the spike protein. Some of these potential spike protein receptor targets include neuropilin-1,18 ASGR1 and KREMEN1.14 The observation of SARS-CoV-2 particles in hepatocytes8 and the robust expression of ASGr1 receptors and neuropilin-1 on hepatocytes suggested that altered liver function associated with COVID-19 infection may be directly caused by infection with the virus and mediated by binding to one or both receptors.
We investigated the potential virus: receptor interactions via the spike protein using fluorescent confocal microscopy and biotinylated spike (RBD) and spike 1 proteins. In a parallel study, E12 MLPC were differentiated to AT2-like cells (manuscript submitted). These cells expressed markers associated with AT2 cells (surfactant protein C, ACE2, TM4SF1, HT2-280), negative for markers associated with AT1 cells (AGER, caveolin 1 and aquaporin) and positive for markers not unique to AT2 cells but known to be expressed on AT2 cells (CK19, CD26 and EpCAM). These results were identical to primary small airway epithelial cells. The binding of biotinylated spike proteins and specific blocking by unlabeled protein and neutralizing antibodies confirmed that the primary interaction of spike protein with AT2-like cells and primary small airway epithelial cells was via the S1 portion of the protein with the ACE-2 receptor. We repeated those studies in support of our findings with the HLC and PHH. The differential inhibition of spike protein binding with two different antibodies suggested that viral neutralization could result from mechanisms other than direct inhibition of S1 (RBD) binding to ACE-2.
The characteristics of SARS-CoV-2 interactions with hepatocytes were studied by observing the binding of biotinylated spike (RBD) and spike 1 proteins to HLC and PHH using the undifferentiated E12 as a known negative control. It was observed that HLC and PHH were both negative for ACE-2, precluding that as a potential site of viral binding. This was confirmed by the inability of the cells to bind S1 protein. The binding of biotinylated spike protein and blockade by unlabeled spike protein on HLC and PHH suggested that the binding was specific, and via a mechanism distinct from ACE-2. The complete inhibition of spike binding by an antibody directed against the ASGr1 is strongly suggestive that ASGr1 is a binding site for the spike protein on hepatocytes. Interestingly, blockade of spike binding by both neutralizing antibodies on HLC and PHH was distinct from AT2 cells where inhibition occurred solely with the antibody that was directed against the RBD. This is suggestive of neutralizing activity that can occur outside the RBD.
Utilization of multiple cell types to study the interactions of spike protein binding will help identify additional receptor pathways for infection with COVID-19. They could also provide a powerful tool to aid in the development of therapeutics against multiple sites on the spike protein or receptors of the host cells. With the existent emergence of new variants and mutations of the SARS-CoV-2 exhibiting enhanced transmissibility, it is especially important to expand our repertoire of cellular models to investigate the effects of the mutations on the binding characteristics of the virus to host cells. The availability of immortalized cells with the stable characteristics of human alveolar type 2 cells and mature well-differentiated hepatocytes could provide an accurate and reproducible tool to effectively study the various virus–host interactions via spike proteins by providing potential viral receptors that are segregated according to cell type. We believe that AT2 and HLC provide such a tool.
Disclosure
Dr Daniel P Collins reports personal fees from BioE, LLC, during the conduct of the study. In addition, Dr Daniel P Collins has a patent “Composition for an in vitro culture medium to maintain and expand stem cell-derived hepatocyte-like cells” pending, as well as, a patent “Methods to develop immortalized hybrid hepatocyte-like cells”, also pending. The authors report no other conflicts of interest in this work.
References
1. Shang J, Wan Y, Luo C, et al. Cell entry mechanisms of SARS-CoV-2. Proc Natl Acad Sci USA. 2020;117(21):11727–11734. doi:10.1073/pnas.2003138117
2. Lan J, Ge J, Yu J, et al. Structure of the SARS-CoV-2 spike receptor-binding domain bound to the ACE2 receptor. Nature. 2020;581(7807):215–220. doi:10.1038/s41586-020-2180-5
3. Premkumar L, Segovia-Chumbez B, Jadi R, et al. The receptor-binding domain of the viral spike protein is an immunodominant and highly specific target of antibodies in SARS-CoV-2 patients. Sci Immunol. 2020;5(48):eabc8413. doi:10.1126/sciimmunol.abc8413
4. Hikmet F, Méar L, Edvinsson Å, et al. The protein expression profile of ACE2 in human tissues. Mol Sys Biol. 2020;16(7):e9610. doi:10.15252/msb.20209610
5. Sungnak W, Huang N, Bécavin C, et al. SARS-CoV-2 entry factors are highly expressed in nasal epithelial cells together with innate immune genes. Nat Med. 2020;26(5):681–687. doi:10.1038/s41591-020-0868-6
6. Hamming I, Timens W, Bulthuis MLC, Lely AT, Navis GJ, van Goor H. Tissue distribution of ACE2 protein, the functional receptor for SARS corona virus. A first step in understanding SARS pathogenesis. J Pathol. 2004;203(2):631–637. doi:10.1002/path.1570
7. Wang D, Eraslan B, Weiland T, et al. A deep proteome and transcriptome abundance atlas of 29 healthy human tissues. Mol Syst Biol. 2019;15(2):e8503. doi:10.15252/msb.20188503
8. Wang Y, Liu S, Liu H, et al. SARS-CoV-2 infection of the liver directly contributes to hepatic impairment in patients with COVID-19. J Hepatol. 2020;73(4):807–816. doi:10.1016/jhep.2020.05.002
9. Desai N, Neyaz A, Szabolcs A, et al. Temporal and spatial heterogeneity of host response to SARS-CoV pulmonary infection. Nat Comm. 2020;11(1):6319. doi:10.1038/s41467-020-20139-7
10. Mehta P, McAuley DF, Brown M, et al. COVID-19: consider cytokine storm syndromes and immunosuppression. Lancet. 2020;395(10229):1033–1034. doi:10.1016/S0140-6736(20)30628-0
11. Chu H, Chan JF, Wang Y, et al. Comparative replication and immune activation profiles of SARS-CoV-2 and SARS-CoV in human lungs: an ex vivo study with implications for the pathogenesis of COVID-19. Clin Infect Dis. 2020;71(6):1400–1409. doi:10.1093/cid/ciaa410
12. Chai X, Hu L, Zhang Y, et al. Specific ACE2 expression in cholangiocytes may cause liver damage after 2019-nCoV infection. bioRxiv. 2020. doi:10.1107/2020.02.03.931/766
13. Lozano-Sepulveda SA, Galan-Huerta K, Martínez-Acuña N, Arellanos-Soto D, Rivas-Estilla AM. SARS-CoV-2 another kind of liver aggressor, how does it do that? Ann Hepatol. 2020;19(6):592–596. doi:10.1016/j.aohep.2020.08.062
14. Gu Y, Cao J, Zhang X, et al. Interaction network of SARS-CoV-2 with host receptome through spike protein. bioRxiv. 2020. doi:10.1101/2020.09.09.28/508
15. Collins DP, Hapke JH, Aravalli RN, Steer CJ. Development of immortalized human hepatocyte-like hybrid cells by fusion of multi-lineage progenitor cells with primary hepatocytes. PLoS One. 2020;15(6):e234002. doi:10.1371/journal.pone.0234002
16. D’Souza AA, Devarajan PV. Asialoglycoprotein receptor mediated hepatocyte targeting – strategies and applications. J Control Release. 2015;203:126–139. doi:10.1016/j.jconrel.2015.02.022
17. Huang X, Leroux JC, Castagner B. Well-defined multivalent ligands for hepatocytes targeting via asialoglycoprotein receptor. Bioconjug Chem. 2017;28(2):283–295. doi:10.1021/acs.bioconjchem.6b00651
18. Daly JL, Simonetti B, Klein K, et al. Neuropilin-1 is a host factor for SARS-CoV-2 infection. Science. 2020;370:861–865. doi:10.1126/science.abd3072
 © 2021 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.
© 2021 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.