Back to Journals » Infection and Drug Resistance » Volume 12
Association of pvl gene with incomplete hemolytic phenotype in clinical Staphylococcus aureus
Authors Gao M , Sang R , Wang G, Xu Y
Received 5 December 2018
Accepted for publication 9 May 2019
Published 14 June 2019 Volume 2019:12 Pages 1649—1656
DOI https://doi.org/10.2147/IDR.S197167
Checked for plagiarism Yes
Review by Single anonymous peer review
Peer reviewer comments 4
Editor who approved publication: Dr Joachim Wink
Menglu Gao,1 Ruirui Sang,1 Gang Wang,1 Yuanhong Xu1,2
1Department of Clinical Laboratory, The First Affiliated Hospital of Anhui Medical University, Hefei, 230000, People’s Republic of China; 2Provincial Laboratories of Pathogen Biology and Zoonoses, Anhui Medical University, Anhui, Hefei 230000, People’s Republic of China
Purpose: This experiment aimed to evaluate the correlation between the hemolytic phenotype of Staphylococcus aureus and pvl gene in terms of characteristics of antibiotic resistance.
Materials and methods: Two-hundred and eleven strains of hospital-acquired S. aureus and their bacterial susceptibility to 20 antibiotics were determined by MicroScan WalkAway96. All strains were cultured on Columbia sheep blood agar plates for 24 hours and then underwent ten passages for investigation of their hemolytic phenotypes. S. aureus produced incomplete β-hemolytic phenotype, termed as S. aureus strains with incomplete hemolytic phenotype (SIHP). The pvl gene was identified by PCR amplification followed by DNA sequencing. Statistical analyses of the data were performed using SPSS version 16.0 software.
Results: Fifty-two (24.64%) strains were confirmed to maintain the incomplete hemolytic phenotype of S. aureus (SIHP). Meanwhile, 15 (7.11%) of 211 strains were found to carry the pvl gene, and eight of the 15 strains were SIHP. Compared with S. aureus strains with complete hemolytic phenotype (SCHP), SIHP showed higher susceptibility to seven of the 20 antibiotics (oxacillin, ciprofloxacin, gentamicin, ceftriaxone, cefoxitin, levofloxacin, and moxifloxacin) (P<0.05). The pvl-positive bacteria had a higher rate of resistance to four antibiotics (rifampin, ciprofloxacin, levofloxacin, and moxifloxacin) in comparison with the pvl-negative strains (P<0.05).
Conclusion: SIHP had a high frequency of pvl gene. The pvl-positive isolates showed less resistance to rifampin, ciprofloxacin, levofloxacin, and moxifloxacin. Additionally, the majority of SIHP isolates (61.54%) were methicillin-resistant S. aureus. SIHP strains had significantly higher antibiotic resistance to cefoxitin when compared with SCHP, while SCHP strains had a high rate of antibiotic resistance to ciprofloxacin, gentamicin, ceftriaxone, levofloxacin, and moxifloxacin. The results may help to provide medical advice for selection of antibiotics for patients with SIHP-associated infections.
Keywords: S. aureus, incomplete hemolytic phenotype, pvl, antimicrobial drug resistance, virulence factor
Introduction
Staphylococcus aureus is one of the most common pathogens isolated from both healthy individuals and patients worldwide, which can produce a variety of secreted toxins and cause various infections in humans.1–3 PVL is one of the toxins of synergohymenotropic exotoxin produced by S. aureus and belongs to the pore-forming toxin family.4–6 PVL shows strong lytic activity against host defense cells such as human polymorphonuclear neutrophils,7 monocytes, macrophages, and rabbit neutrophils but not murine neutrophils in vitro.8–10 Pore formation requires the presence of the two components of the toxin, LukS-PV and LukF-PV. Previous studies demonstrated that PVL aggravated many infections,11 such as skin and soft tissue infection,1 necrotizing pneumonia, bone joint infections, and even bacteremia.12–14
It has been found that the pvl gene can spread from strain to strain by bacteriophages and plasmids.15,16 As a result of the pvl gene's transmission mechanism, the prevalence of pvl-positive strains has gradually increased.17
The prevalence of the pvl gene has been less common in methicillin-susceptible S. aureus isolates than in methicillin-resistant S. aureus (MRSA).18 The pvl gene locus represents a stable genetic marker of community-acquired MRSA (CA-MRSA) strains.19 Studies have shown that PVL had a high proportion in CA-MRSA.20 The PVL genes have also been reported to be present in 77% of the CA-MRSA isolates, but only 4% of HA-MRSA isolates were pvl-positive.21 Although the toxin may be a highly linked epidemiological marker for CA-MRSA strains, PVL is not the major virulence determinant of CA-MRSA.22
Incomplete hemolytic phenotype has an opaque, darker hemolytic ring in contrast with the transparent hemolytic phenotype of S. aureus strains with complete hemolytic phenotype (SCHP) .23 Recently, Zhang et al proposed that a number of S. aureus strains with incomplete hemolytic phenotype (SIHP) strains belong to the class of S. aureus. The authors demonstrated that SIHP strains were MRSA which highly expressed β-hemolysin and carried the tst gene. Because toxic shock syndrome toxin (expressed by tst) causes toxic shock syndrome,24,25 SIHP possesses high virulence potential.23 Here we determined the prevalence of SIHP, antimicrobial resistance, and PVL genes carried in S. aureus, and compared the diversity of these factors between SIHP and SCHP strains.
Materials and methods
Bacterial strains
After excluding strains from repeated sources, a total of 211 strains were selected randomly and isolated from patients in the First Affiliated Hospital of Anhui Medical University from January 2016 to December 2017. Constituent ratios of departments where strains were isolated are shown in Table 1. The specimens included 30 samples of blood (14.22%), 70 of secreta (33.18%), 45 of sputa (21.33%), nine of shunt fluids (4.27%), 25 from wounds (11.85%), eight from interstitial fluids (3.79%), and 24 others (11.37%). The Medical Ethics Committee of the First Affiliated Hospital of Anhui Medical University approved this study (no Quick-PJ2018-07–29) and all isolates were collected with written informed consent from the patients, conducted in accordance with the Declaration of Helsinki.
 | Table 1 Constituent ratios of departments where strains were isolated |
Antimicrobial sensitivity testing
The bacteria were cultured on Columbia sheep blood agar plates at 35°C in an atmosphere containing 5% CO2 (v/v) for 24 hours. According to the manufacturer’s instructions, 1–2 single colonies were added to Mueller-Hinton broth (MHB), followed by proper adjustment of bacteria concentration. Based on the broth microdilution method and the Clinical and Laboratory Standards Institute criteria, bacterial identification and antimicrobial susceptibility testing were performed using MicroScan WalkAway 96 (Beckman Coulter, Brea, CA, USA).26 The bacterial isolates were subjected to 20 antibiotics for sensitivity testing. Antibiotics included: tetracycline, beta-lactams (oxacillin, ceftriaxone, cefoxitin), fluoroquinolones (levofloxacin, ciprofloxacin, moxifloxacin), gentamicin, ampicillin, amoxicillin, daptomycin, clindamycin, nitrofurantoin, linezolid, penicillin, rifampin, sulfamethoxazole–trimethoprim, quinupristin-dalfopristin>, erythromycin, and vancomycin.
Detection of hemolytic phenotype
All the strains were identified as S. aureus by matrix-assisted laser desorption ionization time-of-flight mass spectrometry (MALDI-TOF MS VITEK V3.0, BioMérieux, Craponne, France). The strain ATCC25923 with the complete hemolytic phenotype was taken as SCHP reference. All the isolates were cultured on Columbia sheep blood agar plates from different companies (OXOID, UK; BioMérieux; BD Biosciences, San Jose, CA, USA; TianDa, People's Republic of China) at 35°C in an atmosphere containing 5% CO2 (v/v) for 24 hours, and 1–2 single colonies from one agar plate were taken to the next Columbia sheep blood agar plates from the same company and cultured for 24 hours, and then underwent ten serial passages. The hemolytic phenomenon was observed. Strains with incomplete hemolytic phenotype on all four blood agar plates and their phenotypes stable to ten passages were identified as SIHP and otherwise, as SCHP. Several SIHP colonies gathered and formed a grid-like hemolytic phenotype.
Detection of pvl genes
The strains were inoculated on Columbia blood agar plates and cultured at 35°C in an atmosphere containing 5% CO2 for 24 hours. Four bacterial colonies were selected and suspended in 8 mL of MHB and cultured at 37°C with shaking at 200 rpm for 12 hours. One mL of MHB bacterial suspension was moved to a sterile centrifuge tube with 200 μL lysostaphin (Sangon, Shanghai, People's Republic of China), boiled at 37°C for 1 hour. After that, DNA were extracted and purified using DNA Extraction Kit (TianGen, Beijing, People's Republic of China) according to the manufacturer’s instructions. The DNA were stored at −20°C and prepared for PCR detection.
PCR was run as follows: pre-denaturation at 93°C for 3 minutes, denaturation at 93°C for 30 seconds, annealing at 55°C for 30 seconds, extension at 72°C for 1 minute. PCR products were sequenced and analyzed by BLAST for the expected sequences of peptides. Primers for pvl gene (PVL-F, 5′–ATCATTAGGTAAAATGTCTGGACATGATCCA–3′; and PVL-R, 5′–GCATCAAGTGTATTGGATAGCAAAAGC–3′) were used to generate an internal control with an amplicon size of 433 bp.27
Statistical analysis
Chi-squared test and Fisher’s exact test were used for the statistical analysis and P<0.05 was considered statistically significant. All the data were analyzed by using SPSS version 16.0 software (SPSS Inc., Chicago, IL, USA) and Figures 3–7 were made by GraphPad Prism version 5.0 (GraphPad Software, Inc., La Jolla, CA, USA).
Results
Hemolytic phenotype of SIHP strains on sheep agar blood plates
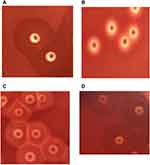
One-hundred and fifty-nine strains cultured on blood agar plates for 24 hours presented complete hemolytic rings (ß-hemolytic phenotype) (Figure 1B). The incomplete hemolytic phenotype formed by a single colony of bacteria looked like a donut in which a small, lighter circular hemolytic ring located at the center and the outside was wrapped in an opaque, darker hemolytic ring (Figure 1A). When the two colonies grew close, a transparent linear hemolysis zone was formed at the junction of the two colonies on the hemolytic ring. Several SIHP colonies aggregated, forming a grid-like hemolytic phenotype (Figure 1C). In addition, after ten serial passages, the incomplete hemolytic phenotype still remained (Figure 1D). Fifty-two of 211 hospital-acquired S. aureus were identified as SIHP strains.
Testing for the pvl gene
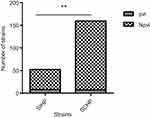
Among the 211 clinical isolates, 15 (7.11%) yielded positive pvl gene amplification (Figure 2). Eight (53.33%) of the 15 pvl-positive isolates belonged to SIHP strains and seven (3.57%) of 196 pvl-negative isolates were SCHP strains. The prevalence of pvl gene in SIHP was significantly higher than that in SCHP (P<0.05) (Figure 3).
Drug resistance of S. aureus
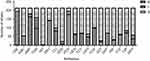
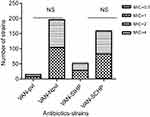
Antimicrobial characteristics of 211 S. aureus isolates are demonstrated in Figure 4. It shows that S. aureus strains presented high resistance to oxacillin (45.02%), cefoxitin (46.45%), ampicillin (76.30%), penicillin (82.46%), and erythromycin (65.88%) when compared with the other antibiotics tested such as tetracycline (31.28%), ceftriaxone (21.80%), levofloxacin (27.01%), ciprofloxacin (32.70%), moxifloxacin (17.06%), gentamicin (28.44%), amoxicillin (24.64%), daptomycin (0.47%), clindamycin (37.91%), nitrofurantoin (0.95%), linezolid (0.95%), rifampin (13.74%), sulfamethoxazole–trimethoprim (9.00%), and quinupristin-dalfopristin (2.84%). Among 211 S. aureus isolates, the minimum inhibitory concentrations (MICs) of vancomycin were found to be MIC =0.5 (two strains), MIC =1.0 (109 strains), MIC 2.0 (99 strains), and MIC =4.0 (one strain). The MICs of S. aureus are shown in Figure 5. No statistically significant difference in MICs was noted between SIHP and SCHP strains. Similarly, no difference was seen between pvl-positive and -negative isolates.
Antimicrobial characteristics of pvl-positive and -negative strains
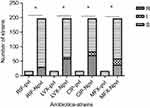
In comparison with the sensitivity of pvl-negative strains to quinolones (levofloxacin 29.08%, ciprofloxacin 35.20%, and moxifloxacin 18.37%) and rifampin (14.29%), the sensitivity of pvl-positive S. aureus to quinolones (levofloxacin 0%, ciprofloxacin 0%, and moxifloxacin 0%) and rifampin (6.67%) was significantly elevated (P<0.05) (Figure 6). No statistically significant difference in antibacterial sensitivity was noted in the remaining 16 antibiotics between pvl-positive and -negative strains (P>0.05).
Antimicrobial characteristics of SIHP and SCHP strains
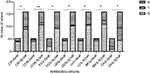
Compared with SCHP strains, SIHP strains had a remarkably higher susceptibility to levofloxacin, ciprofloxacin, and moxifloxacin. Additionally, SIHP strains were more susceptible to gentamicin and ceftriaxone than SCHP strains, but more resistant to cefoxitin than SCHP strains. There were 61.54% of SIHP strains but only 39.62% of SCHP strains found to be MRSA (P>0.05). All statistical differences between SIHP and SCHP strains are shown in Figure 7.
Discussion
Bacteriophages are responsible for spreading pvl gene among bacterial strains.15 Recently, phage-therapy as a therapeutic strategy in some infections has been proposed.28 Bacteriophages are able to completely prevent biofilm formation through this strain and eliminate the staphylococcal biofilms within 4 hours.29 In in vitro inhibitory assays, phage κ lysed a range of clinically isolated MRSA strains successfully.30 We speculate that bacteriophages might inhibit the resistance of pvl-positive strains to quinolones and rifampin.
The pvl gene prevalence of SIHP strains is significantly higher than that of SCHP strains. Moreover, SIHP strains have high susceptibility to quinolones and the susceptibility pattern is similar to pvl-positive strains. We believe that bacteriophages might inhibit the resistance of pvl-positive strains to quinolones, and the correlation between pvl gene and incomplete hemolytic phenotype may be responsible for quinolones sensitivity of SIHP strains.
MRSA occurs more frequently in SIHP strains than SCHP ones, our results are similar to those in the study of Zhang et al.23 They demonstrated that SIHP isolates carried the tst gene which expressed toxic shock syndrome toxin and caused toxic shock syndrome. In the present study, we noted that the pvl gene was frequently carried by SIHP strains and they were more resistant to cefoxitin. Therefore, we suggest that SIHP might be a class of S. aureus with potentially high virulence, that commonly carries virulence factors and presents relatively high resistance to oxacillin and cefoxitin.
SIHP strains are more susceptible to gentamicin and ceftriaxone, and more resistant to oxacillin and cefoxitin than SCHP strains. There may be a series of common regulatory genes that regulate antibiotic sensitivity and hemolytic phenotype. The mechanism of susceptibility, however, remains unclear.
As described previously, SIHP strains with tst and pvl gene cause toxic shock syndrome and aggravate many infections.11,24 They have high susceptibility to quinolones, gentamicin, and ceftriaxone as well. Unfortunately, identification of antibiotic susceptibility from the isolation and culture of bacteria is time-consuming. Therefore, for patients with severe S. aureus infection, selection of antibiotics will be preferentially based on clinical experience. These results may assist when providing medical advice on the appropriate use of antibiotics when the S. aureus isolated from patients has been defined as incomplete hemolytic phenotype. The relationship between the pvl gene and drug resistance provides microbiologists with a way to explore how the PVL gene reduces drug resistance in S. aureus. Our study, however, may not reflect a general situation of hospital-acquired S. aureus in central China, due to the limitation of district and the number of S. aureus strains collected. Further approaches would be needed to clarify the significance of PVL gene in formation of incomplete hemolytic ring and increased antibiotic resistance of S. aureus.
Conclusion
Taken together, the frequency of pvl gene is likely associated with the incomplete hemolytic ring of S. aureus strains. pvl-positive S. aureus shows a high sensitivity to quinolones and rifampin. SIHP strains are susceptible to quinolones, ceftriaxone, and gentamicin but have a higher resistance to oxacillin and cefoxitin when compared with SCHP strains, which may be helpful for correct selection of antibiotics in patients with SIHP-associated infections.
Ethical approval
The Medical Ethics Committee of the First Affiliated Hospital of Anhui Medical University approved this study (no Quick-PJ2018-07-29) and all isolates were collected with written informed consent from the patients, in accordance with the Declaration of Helsinki.
Data availability
All data generated or analyzed during this study are included in this published article.
Acknowledgments
We are grateful to the Department of Anhui Provincial Hospital for the materials used in this study. We especially thank Dr JL Shen from the Key Laboratory of Parasitology and Zoonoses, Anhui for his technical assistance and for access to the laboratory facilities. This work was supported by Anhui Natural Science Foundation (9021138203).
Author contributions
All authors contributed to data analysis, drafting and revising the article, gave final approval of the version to be published, and agree to be accountable for all aspects of the work.
Disclosure
The authors report no conflicts of interest in this work.
References
1. Cribier B, Prevost G, Couppie P, Finck-Barbançon V, Grosshans E, Piémont Y. Staphylococcus aureus leukocidin: a new virulence factor in cutaneous infections? An epidemiological and experimental study. Dermatology. 1992;185:175–180. doi:10.1159/000247443
2. Lina G, Piemont Y, Godail-Gamot F, et al. Involvement of Panton-Valentine leukocidin-producing Staphylococcus aureus in primary skin infections and pneumonia. Clin Infect Dis. 1999;29:1128–1132. doi:10.1086/313461
3. Diep BA, Gillet Y, Etienne J, Lina G, Vandenesch F. Panton-Valentine leucocidin and pneumonia. Lancet Infect Dis. 2013;13:566. doi:10.1016/S1473-3099(13)70102-6
4. Prevost G, Cribier B, Couppie P, et al. Panton-Valentine leucocidin and gamma-hemolysin from Staphylococcus aureus ATCC 49775 are encoded by distinct genetic loci and have different biological activities. Infect Immun. 1995;63:4121–4129.
5. Woodin AM. Fractionation of a leucocidin from Staphylococcus aureus. Biochem J. 1959;73:225–237. doi:10.1042/bj0730225
6. Pedelacq JD, Maveyraud L, Prevost G, et al. The structure of a Staphylococcus aureus leucocidin component (LukF-PV) reveals the fold of the water-soluble species of a family of transmembrane pore-forming toxins. Structure. 1999;7:277–287.
7. Finck-Barbancon V, Duportail G, Meunier O, Colin DA. Pore formation by a two-component leukocidin from Staphylococcus aureus within the membrane of human polymorphonuclear leukocytes. Biochim Biophys Acta. 1993;1182:275–282.
8. Gladstone GP, Van Heyningen WE. Staphylococcal leucocidins. Br J Exp Pathol. 1957;38:123–137.
9. Diep BA, Chan L, Tattevin P, et al. Polymorphonuclear leukocytes mediate Staphylococcus aureus Panton-Valentine leukocidin-induced lung inflammation and injury. Proc Natl Acad Sci U S A. 2010;107:5587–5592. doi:10.1073/pnas.0912403107
10. Hongo I, Baba T, Oishi K, Morimoto Y, Ito T, Hiramatsu K. Phenol-soluble modulin alpha 3 enhances the human neutrophil lysis mediated by Panton-Valentine leukocidin. J Infect Dis. 2009;200:715–723. doi:10.1086/605332
11. Tong SY, Davis JS, Eichenberger E, Holland TL, Fowler VG. Staphylococcus aureus infections: epidemiology, pathophysiology, clinical manifestations, and management. Clin Microbiol Rev. 2015;28:603–661. doi:10.1128/CMR.00134-14
12. Kreienbuehl L, Charbonney E, Eggimann P. Community-acquired necrotizing pneumonia due to methicillin-sensitive Staphylococcus aureus secreting Panton-Valentine leukocidin: a review of case reports. Ann Intensive Care. 2011;1:52. doi:10.1186/2110-5820-1-52
13. Flammier S, Rasigade JP, Badiou C, et al. Human monocyte-derived osteoclasts are targeted by Staphylococcal pore-forming toxins and superantigens. PLoS One. 2016;11:e0150693. doi:10.1371/journal.pone.0150693
14. Shallcross LJ, Fragaszy E, Johnson AM, Hayward AC. The role of the Panton-Valentine leucocidin toxin in staphylococcal disease: a systematic review and meta-analysis. Lancet Infect Dis. 2013;13:43–54. doi:10.1016/S1473-3099(12)70238-4
15. Xia G, Wolz C. Phages of Staphylococcus aureus and their impact on host evolution. Infect Genet Evol. 2014;21:593–601. doi:10.1016/j.meegid.2013.04.022
16. Boyle-Vavra S, Daum RS. Community-acquired methicillin-resistant Staphylococcus aureus: the role of Panton-Valentine leukocidin. Lab Invest. 2007;87:3–9. doi:10.1038/labinvest.3700501
17. Klein S, Menz MD, Zanger P, Heeg K, Nurjadi D. Increase in the prevalence of Panton-Valentine leukocidin and clonal shift in community-onset methicillin-resistant Staphylococcus aureus causing skin and soft-tissue infections in the Rhine-Neckar Region, Germany, 2012-2016. Int J Antimicrob Agents. 2019;53:261–267. doi:10.1016/j.ijantimicag.2018.10.026
18. David MZ, Daum RS. Community-associated methicillin-resistant Staphylococcus aureus: epidemiology and clinical consequences of an emerging epidemic. Clin Microbiol Rev. 2010;23:616–687. doi:10.1128/CMR.00081-09
19. Vandenesch F, Naimi T, Enright MC, et al. Community-acquired methicillin-resistant Staphylococcus aureus carrying Panton-Valentine leukocidin genes: worldwide emergence. Emerg Infect Dis. 2003;9:978–984. doi:10.3201/eid0908.030089
20. Nakagawa S, Taneike I, Mimura D, et al. Gene sequences and specific detection for Panton-Valentine leukocidin. Biochem Biophys Res Commun. 2005;328:995–1002. doi:10.1016/j.bbrc.2005.01.054
21. Naimi TS, LeDell KH, Como-Sabetti K, et al. Comparison of community- and health care-associated methicillin-resistant Staphylococcus aureus infection. Jama. 2003;290:2976–2984. doi:10.1001/jama.290.22.2976
22. Voyich JM, Otto M, Mathema B, et al. Is Panton-Valentine leukocidin the major virulence determinant in community-associated methicillin-resistant Staphylococcus aureus disease? J Infect Dis. 2006;194:1761–1770. doi:10.1086/509506
23. Zhang H, Zheng Y, Gao H, et al. Identification and characterization of Staphylococcus aureus strains with an incomplete hemolytic phenotype. Front Cell Infect Microbiol. 2016;6:146. doi:10.3389/fcimb.2016.00146
24. Saha B, Harlan DM, Lee KP, June CH, Abe R. Protection against lethal toxic shock by targeted disruption of the CD28 gene. J Exp Med. 1996;183:2675–2680. doi:10.1084/jem.183.6.2675
25. Thomas D, Chou S, Dauwalder O, Lina G. Diversity in Staphylococcus aureus enterotoxins. Chem Immunol Allergy. 2007;93:24–41. doi:10.1159/000100856
26. Song KH, Kim ES, Park KH, et al. Clinical and molecular characterization of panton-valentine leukocidin-positive invasive Staphylococcus aureus infections in Korea. Microb Drug Resist. 2019;25:450–456. doi:10.1089/mdr.2018.0238
27. McClure JA, Conly JM, Lau V, et al. Novel multiplex PCR assay for detection of the staphylococcal virulence marker Panton-Valentine leukocidin genes and simultaneous discrimination of methicillin-susceptible from -resistant staphylococci. J Clin Microbiol. 2006;44:1141–1144. doi:10.1128/JCM.44.3.1141-1144.2006
28. Capparelli R, Parlato M, Borriello G, Salvatore P, Iannelli D. Experimental phage therapy against Staphylococcus aureus in mice. Antimicrob Agents Chemother. 2007;51:2765–2773. doi:10.1128/AAC.01513-06
29. Fenton M, Keary R, McAuliffe O, Ross RP, O’Mahony J, Coffey A. Bacteriophage-derived peptidase CHAP(K) eliminates and prevents Staphylococcal biofilms. Int J Microbiol. 2013;2013:625341. doi:10.1155/2013/625341
30. O’Flaherty S, Ross RP, Meaney W, Fitzgerald GF, Elbreki MF, Coffey A. Potential of the polyvalent anti-Staphylococcus bacteriophage K for control of antibiotic-resistant staphylococci from hospitals. Appl Environ Microbiol. 2005;71:1836–1842. doi:10.1128/AEM.71.4.1836-1842.2005
 © 2019 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.
© 2019 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.