Back to Journals » Cancer Management and Research » Volume 12
Assessment of Nine Driver Gene Mutations in Surgically Resected Samples from Patients with Non-Small-Cell Lung Cancer
Authors Wang S, Qu X, Cao L, Hu X, Hou K, Liu Y , Che X
Received 22 February 2020
Accepted for publication 13 May 2020
Published 28 May 2020 Volume 2020:12 Pages 4029—4038
DOI https://doi.org/10.2147/CMAR.S250822
Checked for plagiarism Yes
Review by Single anonymous peer review
Peer reviewer comments 2
Editor who approved publication: Dr Antonella D'Anneo
Shuo Wang,1– 3 Xiujuan Qu,1– 3 Lili Cao,1– 3 Xuejun Hu,4 Kezuo Hou,1– 3 Yunpeng Liu,1– 3 Xiaofang Che1– 3
1Department of Medical Oncology, The First Hospital of China Medical University, Shenyang 110001, People’s Republic of China; 2Key Laboratory of Anticancer Drugs and Biotherapy of Liaoning Province, The First Hospital of China Medical University, Shenyang 110001, People’s Republic of China; 3Liaoning Province Clinical Research Center for Cancer, Shenyang 110001, People’s Republic of China; 4Department of Respiratory Medicine, The First Hospital of China Medical University, Shenyang 110001, People’s Republic of China
Correspondence: Xiaofang Che; Xiujuan Qu Email [email protected]; [email protected]
Background: The mutational profile of oncogenic driver genes play an important role in non-small-cell lung cancer (NSCLC). The need of a testing panel capable of comprehensively determining patient genotypes in limited amounts of material has increased since the recent association of nine core oncogenic driver genes as tumor predictive biomarkers.
Methods: Surgically resected samples from 214 NSCLC patients (168 patients with adenocarcinomas and 46 with squamous cell cancers) were included. A multiplexed PCR-based assay was developed to simultaneously test 118 hotspot mutations and fusions in nine driver genes.
Results: The sensitivity of the kit was 1% for gene mutation and 450 copies for gene fusion. Genetic alterations were detected in 143 (66.8%) patients by the assay. The three most common alterations identified were EGFR mutations (50.9%), KRAS mutations (8.4%) and ALK fusions (4.7%). Eight (3.7%) patients harbored concurrent mutations, and the most common partners were EGFR mutations which were observed in the eight patients. No associations between survival and EGFR, KRAS, and ALK status were observed. Patients with two or more alterations exhibited shorter DFS compared to those with single mutations (P=0.032), whilst had no significant difference in OS (P=0.245). However, only TNM stage was an independent predictor of OS (HR=2.905, P< 0.001) as well as DFS (HR=2.114, P< 0.001) in our cohort in multivariate analysis. Furthermore, patients with the L858R mutation had longer DFS (P=0.014) compared to other sensitizing mutations and tended to have better OS but the differences were not significant (P=0.06).
Conclusion: These findings suggest this multiplex gene panel testing technique can be efficiently used to detect nine driver genes in a limited number of specimens. This methodology would have the potential to save both specimens and time compared to the combination of all assays by other methods.
Keywords: drive gene, lung cancer, multi-mutational profiling, prognosis
Introduction
Lung cancer is one of the most common malignancies worldwide, and seen as a major cause of cancer death.1 Approximately 85% of all lung cancers are non-small-cell lung cancer (NSCLCs),1 with the most common pathological types being adenocarcinoma (ADC) and squamous cell carcinoma (SCC).2 In the past decade, one of the most important advances in the treatment of NSCLC is the concept of individualized therapy which requires accurate molecular diagnosis to stratify patients and inform treatment. The discovery of mutations in epidermal growth factor receptor (EGFR) tyrosine kinase has also opened the door of molecular-targeted therapy. The selective use of tyrosine kinase inhibitors (TKIs) for patients with EGFR-sensitizing mutations has dramatically improved the survival of these patients.3–5 Anaplastic lymphoma kinase (ALK) inhibitors also have demonstrated clinical activity in ALK gene rearrangements group.6–8 Histologic subtype alone is not sufficient to inform individualized treatments and so molecular genetic testing has become central to the clinical management of NSCLC. This paradigm is further evidenced by the NCCN Guidelines for NSCLC stating that gene mutation testing is required before the prescription of targeted therapy.
There is growing recognition of novel gene alterations that may impact therapy selection and correlate with prognosis of patients with KRAS, BRAF, HER2, and ROS1, etc.9–12 To date, most studies have focused on a single genetic mutation, however, there is a necessary to develop an integrated multi-mutational profile of patients to further identify potentially effective treatments and to avoid the use of treatments that are unlikely to provide clinical benefit. In addition, a comprehensive genetic testing can simultaneously obtain the mutational status of genes of interest with fewer samples and less time. These requirements provide a strong rationale for the development of a multi-target test to determine the optimal precision oncology treatments. In addition, the prevalence of EGFR mutations has been intensively investigated and the frequencies of these mutations are shown to differ between Asian and white populations.13 In contrast, the frequency of ALK gene fusions is 1% to 7% of all NSCLC patients with no obvious racial differences.14 However, epidemiological studies of other mutations remain unclear.
In this context, we have developed a multiplex genotyping panel to determine alterations in nine oncogenic genes. In our current study, we demonstrated the frequencies of these driver genes in 214 Chinese patients with NSCLC and found the correlations between the genotype and clinico-pathologic features. Finally, the prognostic values of driver gene mutations were also determined.
Materials and Methods
Ethical Approval
All procedures performed in studies involving human participants were in accordance with the ethical standards of the institutional research committee and with the 1964 Helsinki Declaration and its later amendments or comparable ethical standards. The study was approved by the Ethics Committee of China Medical University (No. 2017-236), and all patients have signed the informed consent.
Study Population
Clinical data of 214 primary NSCLC patients who received a complete resection between 2010 and 2014 were collected. The pathological TNM stage was defined on the American Joint Committee on Cancer (AJCC) 7th edition. No patient received neoadjuvant or adjuvant therapies, including chemotherapy, radiotherapy and target therapy when the specimens were collected. Follow-up information was obtained from medical records or phone interviews after surgery. Overall survival (OS) is the date from surgery to death or the last follow-up date. Disease free survival (DFS) is the date from surgery to recurrence or metastasis or the last follow-up date.
Tissue Specimens and Molecular Analyses
Three formalin-fixed paraffin-embedded (FFPE) specimens were needed for the test which were sectioned into thickness of 5-μm and contained at least 30% tumor cells in specimens. The molecular analysis was carried out by AmoyDx® Multi-Gene Mutations Detection Kit (Amoy Diagnostics, Xiamen, China), which is a real-time PCR assay for qualitative detection of 118 hotspot mutations/fusions in EGFR, KRAS, BRAF, NRAS, HER2, PIK3CA, ALK, ROS1 and RET genes (Details are described in Table S1). This kit contains RNA gene fusion and DNA gene mutation detection systems. The RNA gene fusion detection system uses specific primers and fluorescent probes which are designed for both side genes of the fusion, combining a one-step RT-PCR procedure to detect ALK, ROS1 and RET gene fusions based on tumor message RNA. The DNA gene mutation detection system uses ADx-ARMS technology, which comprises specific primers and fluorescent probes to detect gene mutations. All experiments were performed according to the manufacturer’s instructions.
Statistics Analysis
Analysis were performed using SPSS (version 16.0). χ2 or Fisher’s exact tests as appropriate were used to assess the Association of clinico-pathological parameters with genotype. Kaplan-Meier analysis and Log rank test were used for survival analysis. P<0.05 was considered statistically significant (two‐sided).
Results
Patient Characteristics
Of the total 214 patients, 168 patients (78.5%) had adenocarcinomas and 46 (21.5%) had squamous cell cancers. The patients had a median age of 59 years (age ranging between 30 and 75 years). 46.3% of the patients were female and 57.9% of the patients were never-smokers. According to 7th AJCC TNM staging, 37.9%, 27.6% and 34.5% of patients were classified as stage I, stage II, and stage III, respectively. Table 1 shows the primary details of the patients, including gender, smoking status, histological type, TNM stage, and tumor differentiation.
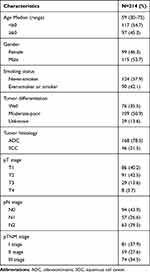 |
Table 1 Clinical Characteristics of the Patient Cohort |
Validation of This Assay Kit
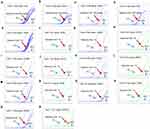
The performance characteristics of this kit were validated by real-time PCR assay and the results were interpreted according to the Ct values of the samples. ALK, ROS1 and RET gene fusions status were analyzed in tubes 1–4. If FAM Ct value of tube 1–4 is <35, the sample was determined as positive, otherwise it was negative. The kit allows detection of 450 copies of gene fusions armored RNA in 0.09–4.5μg FFPE sample RNA (Figure 1A–D). For tubes 5–11, DNA gene mutation status were analyzed. If FAM or VIC Ct value of tube 5–12 is<31, the sample was determined as positive. The sensitivity was 1% for each gene mutation status (Figure 1E–R).
Frequencies of Genetic Alterations
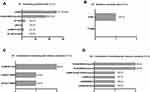
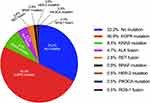
Overall, 143 (66.8%) patients were detected with at least one genetic alteration, whilst 71 (33.2%) patients showed no alterations. The three most commonly mutated genes were EGFR (50.9%, including sensitizing and resistance mutations), KRAS (8.4%) and ALK fusions (4.7%). The frequencies of other genetic alterations were 2.8% for RET fusions, 0.9% for BRAF mutations, 0.5% for HER-2 mutations, 0.5% for PIK3CA mutations, and 0.5% for ROS-1 fusions, respectively. No NRAS mutation was detected (Figure 2).
Eight (3.7%) patients harbored a combination of two or three mutations. EGFR mutations were the most common form for concurrent alterations, which appeared in all 8 patients. EGFR mutations concurrently occurred with other alterations in KRAS (two patients), HER-2 (two patients), PIK3CA (one patient), BRAF (one patient), and ALK (two patients).
The Correlation Between Clinico-Pathologic Correlations and Genotype
As expected, the frequency of EGFR mutation was much higher in female patients (P<0.001), never-smokers (P<0.001), the ADC subtype (P<0.001) and early-stage disease (P<0.001) (Table 2). KRAS mutations were associated with smoking history (P=0.011). ALK fusion patients were significantly younger than wild-type patients, and ALK fusion was well correlated with differentiated tumors.
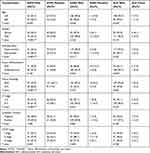 |
Table 2 Frequencies of Genetic Alterations with Clinical Characteristics |
EGFR Mutations
The frequencies and types of EGFR mutations are summarized in Figure 3. Of the 214 patients, 96 (44.9%) had sensitizing mutations alone, one (0.5%) had resistance mutations alone (S768I), four (1.9%) had dual mutations of both sensitizing and resistance, and eight (3.7%) had both of sensitizing mutations and other types of mutations. The most common mutation types detected were exon 21 L858R point mutation and deletion in exon 19. The frequency of L858R (27.6%) was higher compared to deletion in exon 19 (20.6%). Dual L858R/19del mutations were detected in three patients. No patient had the T790M mutation alone, whilst two patients demonstrated the T790M mutation combined with sensitizing mutations.
Prognostic Value of Gene Alternation Status
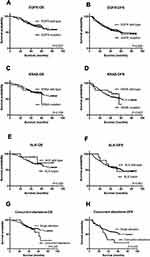
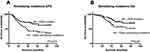
One hundred and one (47.2%) patients relapsed and 70 (32.7%) patients died due to the disease at final follow-up (June 2017). The median follow-up period was 45 months (ranging between 3 and 89 months). OS and DFS were evaluated according to gene mutation status. In Kaplan-Meier (KM) analysis, no associations between survival and EGFR (P=0.977 for OS and P=0.836 for DFS), KRAS (P=0.758 for OS and P=0.607 for DFS), and ALK status (P=0.322 for OS and P=0.861 for DFS) were observed (Figure 4A–F). However, patients with two or more alternations exhibited shorter DFS compared to those with single mutations (P=0.032), whilst had no significant difference in OS (P=0.245) (Figure 4G–H). In further univariate analysis, tumor differentiation and TNM stage showed association with OS and TNM stage, concurrent mutation were correlated with DFS. Finally, multivariate analysis showed that only TNM stage was an independent predictor of OS (HR=2.905, P<0.001) as well as DFS (HR=2.114, P<0.001) in our cohort (Table 3).
 |
Table 3 Univariate and Multivariate Survival Analysis |
In the population of patients with EGFR sensitizing mutation, it was found that patients harboring the L858R mutation had longer DFS (P=0.014) compared to other sensitizing mutation types (Figure 5A). Besides, patients with L858R mutation tended to have better OS although the differences were not significant (P=0.06) (Figure 5B).
Discussion
Tumor mutation analysis of driver genes is essential for realizing personalized medicine in NSCLC involving the optimum use of molecular targeting therapy. To satisfy these requirements, great efforts have been made to identify alterations in critical genes. Previous studies have focused either on the genotype of lung adenocarcinoma,15–17 or in the white population.18–20 In contrast, in the current study, we analyzed data from 214 Chinese lung cancer patients which included 168 patients with adenocarcinomas and 46 with squamous cell cancers.
A multiplexed PCR-based assay was developed to simultaneously test 118 hotspot mutations and fusions in nine driver genes. 66.8% of patients were positive for the detected genes. The most three common mutations identified were detected in EGFR (50.9%), KRAS (8.4%) and ALK fusions (4.7%). Sequist et al20 studied the profile of gene mutations in patients from the USA using a similar detection method. The overall frequency of genetic alterations in our study was 66.8%, which was higher than 51.0% that reported by Sequist et al. One of the most likely reasons is that we demonstrate more EGFR mutations (44.9% in our study versus 13% in Sequist et al) which reflects the ethnic differences of EGFR mutations. A further study16 analyzed data on the mutation status in Japanese patients with lung adenocarcinoma, demonstrating that 48% of patients had mutations or fusions, and the frequency of EGFR mutations was 35%. There were significant differences of smokers between the two studies (42.1% in our study versus 68%). Meanwhile, Li et al15 reported information in lung adenocarcinoma who never smoked, showing 89% of the cohort were identified with at least one genetic alteration. This was higher than our data which showed that the mutation frequency of never-smokers with adenocarcinoma was 80.9%, with no obvious differences between these studies.
Whilst it is widely accepted that testing for EGFR mutations and ALK fusions before therapy is required for patient stratification, it is also important to identify uncommon mutations in driver genes such as BRAF, PIK3CA and HER2, which may also impact therapy selection or be correlated with the prognosis of patients. In our study, amongst the 214 patients, 14 patients were tested with mutations and fusions in PIK3CA, HER2, BRAF, and RET genes. With the rapid development of clinical trial focusing on genotype-specific therapy,10,11 more patients could potentially benefit from these treatments. Furthermore, most oncogenic alternations were found to be mutually exclusive as reported in a previous study.21 However, 7.5% of patients were identified with two or more driver alterations in this study. EGFR mutations were the most common mutation type in combination with other alterations. It has previously been reported that alternations of PIK3CA and MET,22–24 may confer resistance to EGFR-TKIs, it is also crucial to extensively conducting multi-mutational testing in routine clinical practice.
In the present study, we reported that EGFR mutation status had no impact on OS and DFS in resected lung cancer patients. Some previous studies have demonstrated that patients with EGFR sensitizing mutations treated with TKIs had longer survival time than patients without EGFR mutations.25,26 At first sight, our results differ from previous reports. However, our results are based on the patients who received complete resection and did not receive TKIs when the samples were obtained. Similarly, Yotsukura et al27 also did not find any correlations between survival and EGFR mutation status in early-stage lung adenocarcinoma patients who were not treated with TKIs. A meta-analysis based on data from 16 studies indicated that EGFR mutation was not a prognostic factor for NSCLC patients who received a complete resection.28 The above results show that the survival benefit of EGFR mutation may be attributed to the use of TKIs, not the intrinsic survival differences between different EGFR mutations. In addition, patients with two or more alternations had significantly shorter DFS compared to those with a single mutation. Although the size of concurrent alternations in the subset was small, which may have reduced our statistical power, future studies with more samples should be carried out to investigate their impact on patient outcomes.
In summary, our results identified the profile of nine genes alterations in NSCLC patients after complete resection and prognostic values of genotype. Multiplex gene panel testing can efficiently detect nine driver genes with a limited number of specimens. The outcome is expected to be valuable in influencing treatment decisions in patients with lung cancer.
Conclusions
The mutational profile of oncogenic driver genes plays an important role in non-small-cell lung cancer (NSCLC) as several core oncogenic driver genes have been considered to be tumor predictive biomarkers. Our study suggested a multiplex gene panel testing technique can be efficiently used to detect nine driver genes in a limited number of specimens. This methodology would have the potential to save both specimens and time compared to the combination of all assays by other methods.
Acknowledgments
This study was supported by National Science and Technology Major Project of the Ministry of Science and Technology of China (No. 2017ZX09304025); National Natural Science Foundation of China (No. 81972197; No. 81972751; No. 81972331); The Key Research and Development Program of Liaoning Province (No. 2018225060); Science and Technology Plan Project of Shenyang city (No. 1900940).
Author Contributions
All authors made substantial contributions to conception and design, acquisition of data, or analysis and interpretation of data; took part in drafting the article or revising it critically for important intellectual content; gave final approval of the version to be published; and agree to be accountable for all aspects of the work.
Disclosure
The authors report no conflicts of interest in this work.
References
1. Siegel RL, Miller KD, Jemal A. Cancer statistics, 2019. CA Cancer J Clin. 2019;69:7–34. doi:10.3322/caac.21551
2. Herbst RS, Heymach JV, Lippman SM. Lung cancer. N Engl J Med. 2008;359(13):1367–1380. doi:10.1056/NEJMra0802714
3. Mok TS, Wu YL, Thongprasert S, et al. Gefitinib or carboplatin-paclitaxel in pulmonary adenocarcinoma. N Engl J Med. 2009;361:947–957. doi:10.1056/NEJMoa0810699
4. Yang JC, Wu YL, Schuler M, et al. Afatinib versus cisplatin-based chemotherapy for EGFR mutation-positive lung adenocarcinoma (LUX-Lung 3 and LUX-Lung 6): analysis of overall survival data from two randomised, Phase 3 trials. Lancet Oncol. 2015;16:141–151. doi:10.1016/S1470-2045(14)71173-8
5. Osell R, Carcereny E, Gervais R, et al. Erlotinib versus standard chemotherapy as first-line treatment for European patients with advanced EGFR mutation-positive non-small-cell lung cancer (EURTAC): a multicentre, open-label, randomised phase 3 trial. Lancet Oncol. 2012;13(3):239–246. doi:10.1016/S1470-2045(11)70393-X
6. Shaw AT, Kim DW, Nakagawa K, et al. Crizotinib versus chemotherapy in advanced ALK-positive lung cancer. N Engl J Med. 2013;368:2385–2394. doi:10.1056/NEJMoa1214886
7. Peters S, Camidge DR, Shaw AT, et al. Alectinib versus crizotinib in untreated ALK-positive non-small-cell lung cancer. N Engl J Med. 2017;377:829–838. doi:10.1056/NEJMoa1704795
8. Camidge DR, Kim HR, Ahn MJ, et al. Brigatinib versus crizotinib in ALK-positive non-small-cell lung cancer. N Engl J Med. 2018;379:2027–2039. doi:10.1056/NEJMoa1810171
9. Peters S, Gettinger S, Johnson ML, et al. Phase II trial of atezolizumab as first-line or subsequent therapy for patients with programmed death-ligand 1-selected advanced non-small-cell lung cancer (BIRCH). J Clin Oncol. 2017;35:2781–2789. doi:10.1200/JCO.2016.71.9476
10. Planchard D, Smit EF, Groen HJM, et al. Dabrafenib plus trametinib in patients with previously untreated BRAF (V600E)-mutant metastatic non-small-cell lung cancer: an open-label, Phase 2 trial. Lancet Oncol. 2017;18:1307–1316. doi:10.1016/S1470-2045(17)30679-4
11. Peters S, Stahel R, Bubendorf L, et al. Trastuzumab emtansine (T-DM1) in patients with previously treated HER2-overexpressing metastatic non-small cell lung cancer: efficacy, safety, and biomarkers. Clin Cancer Res. 2019;25:64–72. doi:10.1158/1078-0432.CCR-18-1590
12. Chen YF, Hsieh MS, Wu SG, et al. Efficacy of pemetrexed-based chemotherapy in patients with ROS1 fusion-positive lung adenocarcinoma compared with in patients harboring other driver mutations in East Asian populations. J Thorac Oncol. 2016;11:1140–1152. doi:10.1016/j.jtho.2016.03.022
13. Shi Y, Au JS, Thongprasert S, et al. A prospective, molecular epidemiology study of EGFR mutations in Asian patients with advanced non-small-cell lung cancer of adenocarcinoma histology (PIONEER). J Thorac Oncol. 2014;9:154–162. doi:10.1097/JTO.0000000000000033
14. Li T, Kung HJ, Mack PC, et al. Genotyping and genomic profiling of non-small-cell lung cancer: implications for current and future therapies. J Clin Oncol. 2013;31:1039–1049. doi:10.1200/JCO.2012.45.3753
15. Li C, Fang R, Sun Y, et al. Spectrum of oncogenic driver mutations in lung adenocarcinomas from East Asian never smokers. PLoS One. 2011;6(11):e28204. doi:10.1371/journal.pone.0028204
16. Serizawa M, Koh Y, Kenmotsu H, et al. Assessment of mutational profile of Japanese lung adenocarcinoma patients by multitarget assays: a prospective, single-institute study. Cancer. 2014;120(10):1471–1481. doi:10.1002/cncr.28604
17. Sun Y, Ren Y, Fang Z, et al. Lung adenocarcinoma from East Asian never-smokers is a disease largely defined by targetable oncogenic mutant kinases. J Clin Oncol. 2010;28(30):4616–4620. doi:10.1200/JCO.2010.29.6038
18. Barlesi F, Mazieres J, Merlio J-P, et al. Routine molecular profiling of patients with advanced non-small-cell lung cancer: results of a 1-year nationwide programme of the French Cooperative Thoracic Intergroup (IFCT). Lancet. 2016;387(10026):1415–1426. doi:10.1016/S0140-6736(16)00004-0
19. Sholl LM, Aisner DL, Varella-Garcia M, et al. Multi-institutional oncogenic driver mutation analysis in lung adenocarcinoma: the lung cancer mutation consortium experience. J Thorac Oncol. 2015;10(5):768–777. doi:10.1097/JTO.0000000000000516
20. Sequist LV, Heist RS, Shaw AT, et al. Implementing multiplexed genotyping of non-small-cell lung cancers into routine clinical practice. Ann Oncol. 2011;22(12):2616–2624. doi:10.1093/annonc/mdr489
21. Gainor JF, Varghese AM, Ou SHI, et al. ALK rearrangements are mutually exclusive with mutations in EGFR or KRAS: an analysis of 1683 patients with non-small cell lung cancer. Clin Cancer Res. 2013;19(15):4273–4281. doi:10.1158/1078-0432.CCR-13-0318
22. Engelman JA. Targeting PI3K signalling in cancer: opportunities, challenges and limitations. Nat Rev Cancer. 2009;9:550–562. doi:10.1038/nrc2664
23. Sequist LV, Waltman BA, Dias-Santagata D, et al. Genotypic and histological evolution of lung cancers acquiring resistance to EGFR Inhibitors. Sci Transl Med. 2011;3(75):75. doi:10.1126/scitranslmed.3002003
24. Engelman JA, Zejnullahu K, Mitsudomi T, et al. MET amplification leads to gefitinib resistance in lung cancer by activating ERBB3 signaling. Science (Washington D C). 2007;316(5827):1039–1043. doi:10.1126/science.1141478
25. Mitsudomi T. Mutations of the epidermal growth factor receptor gene predict prolonged survival after gefitinib treatment in patients with non-small-cell lung cancer with postoperative recurrence. J Clin Oncol. 2005;23:2513–2520. doi:10.1200/JCO.2005.00.992
26. Chou T-Y. Mutation in the tyrosine kinase domain of epidermal growth factor receptor is a predictive and prognostic factor for gefitinib treatment in patients with non-small cell lung cancer. Clin Cancer Res. 2005;11(10):3750–3757. doi:10.1158/1078-0432.CCR-04-1981
27. Yotsukura M, Yasuda H, Shigenobu T, et al. Clinical and pathological characteristics of, EGFR, mutation in operable early-stage lung adenocarcinoma. Lung Cancer. 2017;109:49–51. doi:10.1016/j.lungcan.2017.04.014
28. Zhang Z, Wang T, Zhang J, et al. Prognostic value of epidermal growth factor receptor mutations in resected non-small cell lung cancer: a systematic review with meta-analysis. PLoS One. 2013;9:e106053. doi:10.1371/journal.pone.0106053
 © 2020 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.
© 2020 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.