Back to Journals » OncoTargets and Therapy » Volume 14
ARID4B Knockdown Suppresses PI3K/AKT Signaling and Induces Apoptosis in Human Glioma Cells
Authors Luo SM , Tsai WC, Tsai CK, Chen Y, Hueng DY
Received 14 October 2020
Accepted for publication 9 December 2020
Published 10 March 2021 Volume 2021:14 Pages 1843—1855
DOI https://doi.org/10.2147/OTT.S286837
Checked for plagiarism Yes
Review by Single anonymous peer review
Peer reviewer comments 2
Editor who approved publication: Dr William C. Cho
Siou-Min Luo,1 Wen-Chiuan Tsai,2 Chia-Kuang Tsai,1,3 Ying Chen,4 Dueng-Yuan Hueng1,5,6
1Graduate Institute of Medical Sciences, National Defense Medical Center, Taipei, Taiwan, Republic of China; 2Department of Pathology, Tri-Service General Hospital, National Defense Medical Center, Taipei, Taiwan, Republic of China; 3Department of Neurology, Tri-Service General Hospital, National Defense Medical Center, Taipei, Taiwan, Republic of China; 4Department of Biology and Anatomy, National Defense Medical Center, Taipei, Taiwan, Republic of China; 5Department of Neurological Surgery, Tri-Service General Hospital, National Defense Medical Center, Taipei, Taiwan, Republic of China; 6Department of Biochemistry, National Defense Medical Center, Taipei, Taiwan, Republic of China
Correspondence: Dueng-Yuan Hueng No. 325, Sec. 2, Chenggong Road, Neihu Dist., Taipei City, 114, Taiwan, Republic of China
Tel +886-2-8792-7177
Email [email protected]
Purpose: Glioblastoma multiforme is a highly malignant primary brain cancer with a poor prognosis. We recently reported that ARID4B could potentially serve as a biomarker associated with poor survival in glioma patients. However, the function of ARID4B in human gliomas remains unclear. The aim of this study is to investigate the molecular cell biology role of ARID4B in human glioma cells.
Materials and Methods: Gene Expression Omnibus (GEO) and Human Protein Atlas (HPA) datasets were analyzed for the expression of ARID4B in WHO pathological grading, overall survival and immunohistochemical staining. Using quantitative RT-PCR and Western blotting, those findings were confirmed in normal brain tissue and glioma cell lines. ARID4B knockdown was conducted via lentivirus-based transfection of small hairpin RNA in human glioma cells to investigate cell proliferation, cell cycle, and apoptosis.
Results: In the present study, our analysis of GEO datasets showed that ARID4B mRNA expression is higher in WHO grade IV tumors (n = 81) than in non-tumor control tissue (n = 23, P < 0.0001). ARID4B knockdown suppressed glioma cell proliferation and induced G1 phase arrest via the PI3K/AKT pathway. It also increased expression of HDAC1, leading to higher acetyl-p53 and acetyl-H3 levels and reduced glioma cell migration and invasion. These effects were mediated via downregulation of AKT pathway components, including p-mTOR, p-PI3K and p-AKT. ARID4B knockdown also led to downregulation of Cyclin D1, which increased apoptosis in human glioma cells.
Conclusion: These findings that ARID4B expression correlates positively with WHO pathologic grading in glioma. ARID4B knockdown suppresses PI3K/AKT signaling and induces apoptosis in human glioma cells. These results suggests that ARID4B acts as an oncogene in human gliomas.
Keywords: invasion, migration, cell proliferation, mTOR, Cyclin D1
Introduction
Glioblastoma multiforme (GBM) is the most common primary brain cancer and is highly aggressive. Despite treatment with chemoradiotherapy and/or surgical resection, GBM patient outcomes are poor.1 Average survival among GBM patients treated with chemoradiotherapy is 14.6 months.2
AT-Rich Interaction Domain 4B (ARID4B) was initially detected in breast cancer tissues and is believed to promote tumor metastasis.3 Identified as a component of the histone deacetylase (HDAC)-dependent SIN3A chromatin remodeling complex,4 ARID4B contains both a DNA-binding domain and a tudor domain, and may function as an adapter protein, but in leukemia, ARID4B acts as a tumor suppressor.5 In addition, ARID4B functions as a transcriptional activator of PIK3CA and PIK3R2, which regulate the PTEN-PI3K pathway in PTEN-deficient prostate cancers. ARID4B is thus a potential therapeutic target in prostate cancers lacking PTEN, as it is necessary for prostate cancer progression and initiation.6 Expression of ARID4B is also elevated in various other cancer types. For example, it promotes mammary tumor metastasis and proliferation,7 and it is reportedly a predictive biomarker in gliomas and meningiomas and is highly expressed in high-grade gliomas.8 But despite these early findings, the role of ARID4B in cell biology of gliomas remains unclear. In the present study, therefore, we knocked down ARID4B in two glioma cell lines to investigate its function in high-grade gliomas.
Materials and Methods
Analysis of GEO Datasets for Human Gliomas
Sample profiles from the Gene Expression Omnibus (GEO) were analyzed. This included 180 files of de-linked data (GDS1962_214934) on ARID4B mRNA expression https://www.ncbi.nlm.nih.gov/geo/tools/profileGraph.cgi?ID=GDS1815:214943_s_at. Information on pathological grading and ARID4B gene expression from the GDS1962_214934 dataset included 130 data files from 23 patients without tumors (controls), 7 with grade II gliomas, 18 with grade III gliomas, and 81 with grade IV gliomas. The survival analysis of the datasets from GDS1815/214943_s_at (https://www.ncbi.nlm.nih.gov/geo/tools/profileGraph.cgi?ID=GDS1815:214943_s_at) contained 100 high-grade gliomas samples (HG-U133A), which excluded unknown survival of 23 datasets; therefore, 77 patient survival data were analyzed and showed.
Analysis of The Cancer Genome Atlas (TCGA) Datasets for Human Gliomas from the Human Protein Atlas (HPA)
The survival analysis of the datasets from TCGA in the Human Protein Atlas (https://www.proteinatlas.org/ENSG00000054267-ARID4B/pathology/glioma) contained 153 high-grade gliomas samples survival data were analyzed and showed.
Datasets of Immunohistochemical Staining from the Human Protein Atlas (HPA)
Immunohistochemical staining for ARID4B in human gliomas was examined using the Human Protein Atlas (HPA) (https://www.proteinatlas.org/ENSG00000054267-ARID4B/pathology/tissue/glioma#). The dataset included pathological glioma sections from 11 patients that were stained using anti-ARID4B antibody (HPA027333). Four of the 11 samples were low-grade gliomas, while the other 7 samples were high-grade. The quantity was defined as 0 (Negative), 1 (<25%), 2 (25–75%), and 3 (>75%). The intensity was scored as 0 (Negative), 1 (Weak), 2 (Moderate), and 3 (Strong). Labeling index was scored based on multiplying quantity by intensity.
Cell Culture and shRNA Transfection
LN229, U87MG and U118 glioma cell lines purchased from American Type Culture Collection (ATCC). GBM8401 glioma cell line, provided by Dr. Wei-Hwa Lee, was obtained from the patient before the establishment of the Institutional Review Board (IRB) of the National Defense Medical Center (NDMC). The characterization of GBM8401 glioma cell line has published in article entitled as Establishment and characterization of a malignant glioma cell line, GBM8401/TSGH, NDMC.9 GBM8401, U87MG and U118 human glioma cells were cultured in high-glucose (4.5 g/L) Dulbecco’s modified Eagle’s medium (DMEM, Corning, USA REF:10–013-CV) containing 10% fetal bovine serum (FBS, Gibco USA REF:10,437–028), penicillin, and streptomycin at 37°C in a 5% CO2 incubator. LN229 human glioma cells were cultured in low-glucose (1 g/L) DMEM (Corning, USA REF: 10–014-CV) containing 10% FBS (Corning, USA REF: 10,437–028), penicillin, and streptomycin (Corning, USA REF: 30–004-CI) at 37°C in a 5% CO2 incubator.
For ARID4B knockdown experiments, GBM8401 or LN229 cells were infected for 24 h with a shLuc control and ARID4B-shRNA-containing lentiviral vector (TRCN0000146977, RNAi core, Academia Sinica, Taiwan, ROC), after which the transfected cells were selected for 14 days in medium containing puromycin (2 g/L InvivoGen CAS:59–58-2). Stable clones were cultured in medium containing puromycin (0.2 mg/L).
cDNA Synthesis and RNA Isolation
Extraction of total mRNA, reverse transcription and quantitative RT-PCR were performed according to the manufacturer’s protocol. For cDNA synthesis, using Oligo dT primer reverse transcribed 1.0 μg RNA into cDNA with Bioline Tetro Reverse Transcriptase (London, UK Cat.No.BIO-65,050). Normal brain cDNA was obtained from Origene Technologies (Rockville, MD, USA cat.CH1001). The reverse transcripts were amplified and quantified using a Onestep RT-qPCR system. GAPDH served as an internal control to normalize relative gene expression quantified using the 2−ΔΔ CT method. The primer pairs used were as follows: for ARID4B, 5′-ATGAGCCTCCCTATTTGACAGT-3′ (forward) and 5′-GGCCCTTTATGTGGTCATCCT-3′ (reverse); for GADPH, 5′-CTTCATTGACCTCAACTAC-3′ (forward) and 5′-GCCATCCACAGTCTTCTG-3′ (reverse); for HDAC1, 5′-CGCCCTCACAAAGCCAATG-3′ (forward) and 5′-CTGCTTGCTGTACTCCGACA-3′ (reverse); for PIK3CA, 5′-CCACGACCATCATCAGGTGAA-3′ (forward) and 5′-CCTCACGGAGGCATTCTAAAGT-3′ (reverse); for CCND1, 5′-GCTGCGAAGTGGAAACCATC −3′ (forward) and 5′-CCTCCTTCTGCACACATTTGAA-3′ (reverse); for PTEN, 5′- TGGATTCGACTTAGACTTGACCT −3′ (forward) and 5′-GGTGGGTTATGGTCTTCAAAAGG-3′ (reverse); for SIN3A, 5ʹ- GGTGGAGGATGCGCTATCTTA-3ʹ (forward) and 5ʹ- GGGTGTCGATGCTCTGAGATTT −3ʹ (reverse).
Cell Lysate Preparation and Western Blotting
GBM8401, LN229 and U87MG cells were lysed in RIPA buffer (100 mM Tris-HCl, 150 mM NaCl, 0.1% SDS, and 1% Triton-X-100) and centrifuged at 13,500 rpm for 10 min, after which the supernatant was collected as the cell lysate. Cell lysates each containing 20 μg of protein were analyzed by 8% and 12% sodium dodecyl sulfate polyacrylamide gel electrophoresis. Following electrophoresis, the separated proteins were transferred to PVDF membranes, which were then blocked for 1 h at room temperature in 5% skim milk in TBST buffer. The antibodies used included anti-ARID4B (BETHYL A302-223A, 1:2000), Cyclin D1 (Cell Signaling Technology CST2978, 1:1000), phosphorylated (p)-PI3K (Cell Signaling Technology CST4228, 1:1000), PI3K (Cell Signaling Technology CST4292, 1:2000), p-AKT (Ser473) (Cell Signaling Technology CST9271 1:2000), AKT (Cell Signaling Technology CST4691 1:3000), p-mTOR (Ser2448) (Cell Signaling Technology CST2971, 1:1000), mTOR (Cell Signaling Technology CST2972, 1:1000), acetyl-p53 (Cell Signaling Technology CST2972, 1:2000), p53 (Santa Cruz Biotechnology E2918, 1:3000), acetyl-H3 (Cell Signaling Technology CST9677, 1:1000), GAPDH (Santa Cruz Biotechnology SC-47724, 1:5000) and ACTN (Santa Cruz Biotechnology SC-17829, 1:5000). Bands were developed using enhanced chemiluminescence and X-ray film (GE Healthcare, RPN2106, Piscataway, NJ, USA).
Soft Agar Colony Formation Assays
To assay colony formation in soft agar, we seeded LN229 and GBM8401 cells with knockdown ARID4B, and control stable clones at a density of 2 × 103 cells/well into 6-well plates with medium containing 0.35% SeaPlaque Agarose (Lonza Rockland, Inc. ME, USA cat.50002) layered on top of a base medium containing 0.5% agarose. After 4 weeks, the colonies were stained with 0.1% crystal violet (Merck KGaA DE, Cat.115940), and those greater than 0.05 mm in size were counted using ImageJ software (NIH, Bethesda, MD).
Cell Migration and Invasion Assay
Cells were seeded into 12-well plates and grown at 37°C in a 5% CO2 incubator. When the cells reached 90% confluence, we removed the medium, made a wound in the monolayer with a 200 μL pipette tip, and washed the plate three times to remove non-adherent cells. The wound area was photographed immediately after wounding (0 hr) and again 16 h later. The migration rates were computed based on the change in wound area measured using ImageJ software (NIH, Bethesda, MD).
Invasion assays were carried out in Transwell chambers (Corning BioCoat cat.354480). A Matrigel matrix layer was coated onto the upper side of the filter separating the upper and lower chambers. Glioma cells in serum-free DMEM were then added to the upper chambers, and medium containing 10% FBS was added to the lower chambers. After incubation for 16 h at 37°C in a CO2 incubator, the non-migrated cells on the upper side of the filter were removed. Cells that invaded the Matrigel and migrated to the lower side of the filter chamber were stained with 0.1% crystal violet for 10 mins. The glioma cells in upper side of the filter were removed by swab. The invaded glioma cells in lower side of the filter was photographed. Finally, the invading cells were dissolved in 1% SDS, and the optical density at 595 nm was measured.
Cell Proliferation and Flow Cytometric Cell Cycle Analysis
For cell cycle analysis, glioma cells were fixed in 70% ethanol at 4°C and kept at –20°C overnight. The cells were then washed twice with cold phosphate-buffered saline (PBS) and stained with propidium iodide (PI, Sigma-Aldrich, MO, USA Cas.25535–16-4) (50 μg/mL PI in PBS, 1% Tween 20 and 10 μg/mL RNase A) for 30 min in the dark. Cell cycle analysis was performed by measuring DNA content using fluorescence activated cell sorting (BD Biosciences, San Jose, CA, USA). Two independent experiments were performed. For the proliferation assays, glioma cells were processed using FITC-BrdU Flow Kits according to the manufacturer’s instructions (BD Biosciences, San Jose, CA, USA cat.51–2354AK). The cells were then trypsinized with trypsin EDTA and analyzed immediately using a FACS Calibur flow cytometer (BD Biosciences, San Jose, CA, USA). The data collected were analyzed using FACSDiva software (BD Biosciences, San Jose, CA, USA).
Protein–Protein Interaction Network Analysis and Genomic Data Commons Data Portal
The distribution of ARID4B in datasets from the Genomic Data Commons Data Portal were determined and analyzed for known and predicted protein–protein interactions using the Search Tool for the Retrieval of Interacting Genes/Proteins (STRING) database (https://string-db.org/).
Apoptosis Assay
To evaluate the incidence of apoptosis, we used the PE Annexin V Apoptosis Detection Kit I according to the manufacturer’s instructions (BD Biosciences San Jose, CA, USA, cat.559763). Briefly, glioma cells were seeded into 6-well dishes at a density of 3 × 105 cells per well. Stained cells were detected using Annexin V-FITC/7-AAD. The cells were categorized into 3 groups: viable (annexin V− and FITC−), early apoptotic (annexin V+ and FITC−), and later apoptotic (annexin V+ and FITC+).
Statistical Analysis
Student’s t-test or one-way analysis of variance was used to make comparisons among groups. The results are presented as the means ± s.d. or as specified. Values of *P < 0.05 were considered significant.
Results
ARID4B Expression Correlates Positively with WHO Pathological Grading of Gliomas and with Poor Survival in Patients with High-Grade Gliomas
To correlate WHO pathological grading of gliomas with ARID4B expression, we analyzed the association between ARID4B expression and pathological grading of human gliomas using a GEO mRNA de-linked databases (GDS1962_214934). We found that ARID4B expression was significantly lower in non-tumor controls (n = 23) than in WHO grade III (n = 19; P = 2 × 10−6) and grade IV (n = 81; P =1.36 × 10−9) tumors (Figure 1A).
ARID4B Expression and Survival analyses in High-Grade Gliomas
GEO profiles were used to analyze the relationship between ARID4B mRNA expression and survival. As shown in Figure 1B, the group from GDS1815/214943_s_at HG-U133A showing high ARID4B expression had poorer survival than that showing low ARID4B expression (median survival: 70 vs 115 weeks, *P = 0.0232, Gehan-Breslow-Wilcoxon Test; hazard ratio: 0.4210, 95% CI: 0.2398-0.7393, cut-off value set at 923.01.). Thus, ARID4B expression is higher in high- grade than in low-grade gliomas, which suggests ARID4B overexpression contributes to the poor prognosis of GBM patients. In contrast, the survival curves from TCGA database showed the survival probability in glioma patients with low ARID4B expression (<3.52) (n = 96) was shorter than high ARID4B expression, **P = 0.0015 (Figure 1C).
Immunohistochemical Staining of ARID4B in Human Glioma Patient Samples
Using HPA database of human proteins, we found that the immunohistochemical staining of ARID4B in high-grade glioma was higher than that in low-grade glioma, but it was not statistically significant (Figure1D).
ARID4B Knockdown in Human Glioma Cell Lines
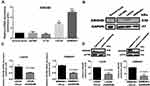
Confirming ARID4B expression levels in human glioma cell lines, RT-qPCR showed that both LN229 and GBM8401 human glioma cells expressed higher levels of ARID4B mRNA than normal brain cells (Figure 2A). Correspondingly, Western blotting showed higher ARID4B protein expression in the two human glioma cell lines (Figure 2B). We therefore chose LN229 and GBM8401 cells for subsequent ARID4B knockdown experiments. To determine the mechanism by which ARID4B contributes to glioma tumorigenesis, we used ARID4B shRNA purchased from the RNAi core at Academia Sinica. We then established GBM8401 and LN229 glioma cells stably expressing shARID4B, which was confirmed using RT-qPCR.The results showed that the ARID4B mRNA expression was significantly suppressed in the ARID4B knockdown cells (*P = 0.0282 and **P = 0.0028, respectively) (Figure 2C). Western blotting revealed similar suppression of ARID4B protein expression in the knockdown cells (***P = 0.0006 and **P = 0.001, respectively) (Figure 2D).
ARID4B Knockdown Suppresses Glioma Cell Migration and Invasion
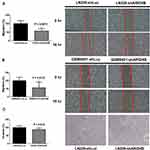
To assess the contribution of ARID4B to glioma cell migration behavior, we performed wound healing assays with LN229 and GBM8401 glioma knockdown cells. The results demonstrate that ARID4B knockdown significantly reduces glioma cell migration (**P = 0.0011, and *P =0.0123, respectively) (Figure 3A and B). In addition, Matrigel-based invasion assays similarly showed that silencing ARID4B expression in LN229 glioma cells significantly suppressed the invasiveness of the cells (*P= 0.019) (Figure 3C).
ARID4B Knockdown Suppresses Cell Proliferation and Increases Apoptosis
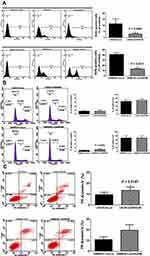
We used BrdU assays to investigate LN229 and GBM8401 cell proliferation. Flow cytometric analysis showed that ARID4B knockdown significantly reduced proliferation of both LN229 (*P = 0.0288) and GBM8401 (**P = 0.0011) glioma cells (Figure 4A). ARID4B knockdown also reduced expression of Cyclin D1, a pro-survival effector protein, and increased the glioma cell fraction in subG1 phase (Figure 4B). In addition, Annexin-V assays showed that ARID4B knockdown significantly increased Annexin-V/7-AAD (*P = 0.0147), indicating that ARID4B knockdown enhanced apoptosis (Figure 4C).
ARID4B Knockdown Suppresses Colony Formation by Human Glioma Cells
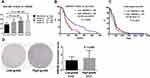
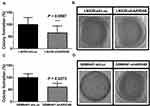
Because ARID4B overexpressed in high-grade gliomas, we investigated the effect of ARID4B knockdown on the tumorigenicity of LN229 and GBM8401 glioma cells by carrying out soft-agar colony formation assays. We found that ARID4B knockdown led to a significant reduction of colony formation by LN229 and GBM8401 glioma cells (***P = 0.0007 and *P = 0.0273, respectively) (Figure 5).
ARID4B Inhibits Glioma Growth via the PTEN-ARID4B-PI3K Pathway
The ARID4B protein–protein interaction (PPI) network obtained with the Search Tool for the Retrieval of Interacting Genes/Proteins (STRING) database showed in Figure 6. We found that ARID4B interacts with HDAC1, HDAC2, chromodomain-helicase-DNA-binding protein 3 (CHD3), chromodomain-helicase-DNA-binding protein 4 (CHD4), TP53, Cyclin D1 (CCND1), phosphatase and tensin homolog (PTEN), SIN3 Transcription Regulator Family Member A (SIN3A), phosphoinositide 3-kinase (PI3K), AKT, and mammalian target of rapamycin (mTOR). AKT, also known as protein kinase B (PKB), is a serine/threonine protein kinase that functions as an effector protein situated downstream of PI3K. The PI3K/AKT/mTOR signaling pathway is important for mediating cell survival and growth. PTEN suppresses PI3K/AKT signaling by dephosphorylating PIP3. HDAC1 is a key regulator of TP53 activity that acts through deacetylation. ARID4B is involved in the SIN3/HDAC1 complex and recognized to be a transcriptional activator of class I PI3K subunit p110α (encoded by PIK3CA), which is a regulator of the PTEN-PI3K pathway.
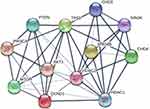 |
Figure 6 ARID4B protein-protein interaction (PPI) network. The PPI network was generated using the STRING database. |
ARID4B was recently identified as a component of the HDAC1/SIN3A chromatin remodeling complex and as a regulator or PTEN-PI3K pathway that acts by targeting a PIK3CA transcriptional activator. Using real-time qPCR, we found here that ARID4B knockdown reduced PI3KCA and Cyclin D1 gene expression and increased gene expression of SIN3A, HDAC1 and PTEN in LN229 (Figure 7A) and GBM8401 (Figure 7B) glioma cells. In addition, Western blotting showed that ARID4B knockdown reduced the protein expression of p-AKT, p-mTOR, and the p110α subunit of PI3K. On the other hand, levels of HDAC1 were increased in ARID4B knockdown glioma cells. The elevated HDAC1 levels led in turn to decreases in acetyl-p53 and acetyl-H3 in LN229 (Figure 8 and Table 1) glioma cells. Taken together, our results demonstrate that ARID4B knockdown increases HDAC1 levels, leading to reductions in acetyl-p53 (Figure 9).
 |
Table 1 The Proteins are Upregulated and Downregulated in ARID4B Knockdown |
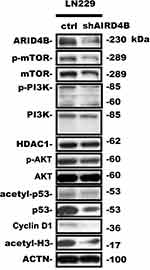 |
Figure 8 Effect of ARID4B knockdown on expression of the indicated proteins. Western blot analysis of expression of the indicated proteins in LN229 cells. Statistical data are presented as the mean ± s.d.; n = 3; in Table 1. |
Discussion
Our previous study showed that ARID4B expression tended to shorten survival rates, but it did not reach statistical significance.8 We further enrolled more data to analyze, as shown in Fig1B and Fig1C, these two cohorts showed that the ARID4B expression and survival outcome of human gliomas have some differences. The possible explanations included the different cohorts and different analytical methods.10 Therefore, further enroll large cohort human gliomas are mandatory in the future. In the present study, we showed that ARID4B knockdown induces apoptosis via a PI3K/AKT signaling pathway in human glioma cells. To our knowledge, this is the first study investigating the molecular mechanism of action of ARID4B in human gliomas. Previous studies showed that ARID4B took part in differentiation and cell growth.11 miR-290–targeted ARID4B inhibited the promoting effects in breast cancer.12 ARID4B reported that gastric cancer can be detected through molecular imaging. Furthermore, ARID4B might serve as a treatment target in gastric cancer.3,13–14 ARID4B is considered to be an oncogene in hepatocellular carcinoma and can be used as a biomarker for the prognoses of patients.15
STRING network analysis showed that ARID4B interacts with HDAC1, Cyclin D1, PTEN, SIN3A, PI3K, AKT, TP53, and mTOR, which form important signaling pathways that play key roles in a variety of crucial cellular processes,16 including cell migration and invasion.17 Reducing levels of p-PI3K and p-AKT previously showed to suppress glioma cell migration and invasion.18 Consistent with those findings, we observed that ARID4B knockdown reduced expression of both p-AKT and p-mTOR and suppressed cell migration and invasion. Recent studies showed that ARID4B is a component of the SIN3A/HDAC1 repressor complex; however, ARID4B does not exhibit acetyltransferase or methyltransferase activity,19 though it does act as a regulator of the PI3K/AKT pathway.6 Similarly, we found that ARID4B knockdown reduces expression of the PI3K p110α (PIK3CA) and p85 subunits, as well as the levels of p-AKT, p-mTOR, and Cyclin D1. Cyclin D1 is an important regulator of cell proliferation that also controls tumorigenesis, cellular senescence, and apoptosis.20 Through its actions, Cyclin D1 exerts cytoprotective effects against induction of apoptosis.16 The PI3K/AKT signal transduction pathway is involved in the regulating survival and apoptosis. Inhibiting AKT phosphorylation may decrease the anti-apoptotic effect of AKT. Thus, reducing p-AKT, Cyclin D1 and p-mTOR levels relieved the suppression of apoptosis in glioma cells. It follows then that the reduced expression of p-AKT, p-mTOR and Cyclin D1 following ARID4B knockdown leads to enhanced apoptosis in glioma cells.18
Our results show that ARID4B knockdown increases expression of HDAC1, which is known to deacetylate p53 and H3. Histone deacetylase inhibitors (HDACIs) induce apoptosis and growth arrest in many cancers.20 In addition, p53 plays an important role in tumor oncogenic stress and cell death, acting as a crucial tumor suppressor mediating cell cycle arrest and apoptosis.21 Inhibiting HDAC1 and SIRT1 leads to enhancement of p53 acetylation, which increases p53-dependent senescence and apoptosis in many cancer cell types.22 This suggests p53 acetylation is an important regulator of apoptosis. The p53 pathway is highly dysregulated in GBM.23 This is due in part to missense mutations in p53 that often lead to reduced tumor suppressor activity.24 In our study, we used LN229 and GBM8401 cells, two p53-mutant and PTEN-wild type glioblastoma cell lines. LN229 cells, for example, harbor a mutated p53 with possible homozygous deletions.11 We suggest that the ability of HDAC1 to suppress LN229 cell apoptosis may be decreased by the p53 mutation.
Conclusion
In summary, our data show that the level of ARID4B expression correlates positively with WHO pathologic grades of gliomas. ARID4B appears to be a potentially useful biomarker of the prognosis in glioma patients, and we suggest this may enable identification of novel therapeutic targets for treatment of human gliomas in the coming future.
Data Sharing Statement
All data are included in this article.
Consent for Publication
All participants agreed to publish the study.
Acknowledgments
We appreciated the help for lentiviruses carrying a ARID4B-specific shRNA (shARID4B) from National RNAi Core Facility, Academia Sinica, Taiwan. We thank Professor Shih-Ming Huang (Graduate Institute of Biochemistry, National Defense Medical Center, Taipei, Taiwan) for providing the experimental help.
Author Contributions
All authors made a significant contribution to the work reported in the conception, study design, execution, and acquisition of data, analysis and interpretation; took part in drafting, revising, and critically reviewing the article. All authors gave final approval of the version to be published; have agreed on the journal to which the article has been resubmitted; and agree to be accountable for all aspects of the work.
Funding
This work was supported, in part, by grants from the Ministry of Science and Technology (MOST 106-2314-B-016-012-MY3, and MOST 108-2314-B-016-026 -MY3 to D.-Y.H.), Tri-Service General Hospital (TSGH-C107-008-S05, ATSGH-C107-008-S05, TSGH-C108-007-008-S05, TSGH-C01-109016, TSGH-2-C107-106-2314-B-016-012-MY3,TSGH-1-C108-106-2314-B-016-012-MY3,TSGH-2-C108-106-2314-B-016-012-MY3, TSGH-2-C108-108-2314-B-016-0-MY3 to D.-Y.H), and Medical Affairs Bureau, Ministry of National Defense (MAB-106-019, MAB-107-009, MAB-108-022, MAB-109-014 to D.-Y.H).
Disclosure
The authors report no conflicts of interest in this work.
References
1. Koul N, Sharma V, Dixit D, Ghosh S, and Sen E. Bicyclic triterpenoid Iripallidal induces apoptosis and inhibits Akt/mTOR pathway in glioma cells. BMC Cancer. 2010;10(328). doi:10.1186/1471-2407-10-328
2. Chu CW, Ko HJ, Chou CH, et al. Thioridazine Enhances P62-Mediated Autophagy and Apoptosis Through Wnt/β-Catenin Signaling Pathway in Glioma Cells. International Journal of Molecular Sciences. 2019;20(473). doi:10.3390/ijms20030473
3. Wang K, Ruan J, Qian Q, et al. BRCAA1 monoclonal antibody conjugated fluorescent magnetic nanoparticles for in vivo targeted magnetofluorescent imaging of gastric cancer. Journal of Nanobiotechnology. 2011;9(23). doi:10.1186/1477-3155-9-23
4. Wu RC, Zeng Y, Pan IW, and Wu MY. Androgen Receptor Coactivator ARID4B Is Required for the Function of Sertoli Cells in Spermatogenesis. Molecular Endocrinology. 2015;29(9):1334–1446. doi:10.1210/me.2015-1089
5. Wu MY, Eldin KW, and Beaudet AL. Identification of Chromatin Remodeling Genes Arid4a and Arid4b as Leukemia Suppressor Genes. Journal of the National Cancer Institute. 2008;100:1247–1259. doi:10.1093/jnci/djn253
6. Wu RC, Young IC, Chen YF, Chuang ST, Toubaji A, and Wu MY. Identification of the PTEN-ARID4B-PI3K pathway reveals the dependency on ARID4B by PTEN deficient prostate cancer. Nature Communication. 2019;10(4332). doi:10.1038/s41467-019-12184-8
7. Winter SF, Lukes L, Walker RC, Welch DR, Hunter and KW. Allelic Variation and Differential Expression of the mSIN3A Histone Deacetylase Complex Gene Arid4b Promote Mammary Tumor Growth and Metastasis. PLoS Genetics. 2012;8(5). doi:10.1371/journal.pgen.1002735
8. Tsai WC, Hueng DY, Nieh S, and Gao HW. ARID4B is a good biomarker to predict tumour behaviour and decide WHO grades in gliomas and meningiomas. Journal of Clinical Pathology. 2017;70:162–167. doi:10.1038/s41467-019-12184-8
9. Lee WH, Yeh MY, Tu YC, Han SH and Wang YC. Establishment and characterization of a malignant glioma cell line, GBM8401/TSGH, NDMC. Journal of Surgical Oncology. 1988;38(3):173–181. doi:10.1002/jso.2930380309
10. Sharma T, Schwalbe EC, Williamson D, et al. Second‑generation molecular subgrouping of medulloblastoma: an international meta‑analysis of Group 3 and Group 4 subtypes. Acta Neuropathologica. 2019;138(2):309–326. doi:10.1007/s00401-019-02020-0.
11. Kato S, Han SY, Liu W, et al. Understanding the function–structure and function–mutation relationships of p53 tumor suppressor protein by high-resolution missense mutation analysis. PANS. 2003;100(14):8424–8429. doi:10.1073/pnas.1431692100
12. Goldberger N, Walker RC, Kim CH, Winter S, and Hunter KW. Inherited variation in miR-290 expression suppresses breast cancer progression by targeting the metastasis susceptibility gene Arid4b. Cancer Research. 2013;73(8):11. doi:10.1158/0008-5472.CAN-12-3513
13. Chao L, Yang J, Can W, et al. BRCAA1 antibody- and Her2 antibody-conjugated amphiphilic polymer engineered CdSe/ZnS quantum dots for targeted imaging of gastric cancer. Nanoscale Research Letters. 2014;9(244). doi:10.1186/1556-276X-9-244
14. Daxiang C, Chunlei Zh, Bing L, et al. Regression of Gastric Cancer by Systemic Injection of RNA Nanoparticles Carrying both Ligand and siRNA. Scientific Reports. 2015;5(10726). doi:10.1186/1477-3155-9-23
15. Rongchang W, Zheng Y, Fan Ch, Chunlian L, Qian W and Xiaohui H. Overexpression of ARID4B predicts poor survival in patients with hepatocellular carcinoma. Human Pathology. 2018;73. doi:10.1016/j.humpath.2017.12.012..
16. Abraham AG and O’Neill E. PI3K/Akt-mediated regulation of p53 in cancer. Signalling and Acquired Resistance to Targeted Cancer Therapeutics. 2014;42:798–803. doi:10.1042/BST20140070
17. Xu WT, Yang Z, and Lu NG. A new role for the PI3K/Akt signaling pathway in the epithelial-mesenchymal transition. Cell Adhesion & Migration. 2015;9(4):317–324. doi:10.1080/19336918.2015.1016686
18. Liu YG, Zheng J, Zhang Y, et al. Fucoxanthin Activates Apoptosis via Inhibition of PI3K/Akt/mTOR Pathway and Suppresses Invasion and Migration by Restriction of p38-MMP-2/9 Pathway in Human Glioblastoma Cells. Neurochemical Research. 2016;41:2728–2751. doi:10.1007/s11064-016-1989-7
19. Raul AG, Hugo GR, Emmanuel ML, et al. SurvExpress: An Online Biomarker Validation Tool and Database for Cancer Gene Expression Data Using Survival Analysis. PLOS ONE. 2013;8(9). doi:10.1371/journal.pone.0074250
20. Roué G, Pichereau V, Lincet H, Colomer D, and Sola B. Cycling D1 mediates resistance to apoptosis through upregulation of molecular chaperones and consequent redistribution of cell death regulators. Oncogene. 2008;27:4909–4920. doi:10.1038/onc.2008.126
21. Seo SK, Jin HO, Woo SH, et al. Histone Deacetylase Inhibitors Sensitize Human Non-small Cell Lung Cancer Cells to Ionizing Radiation Through Acetyl p53-Mediated c-myc Down-Regulation. Journal of Thoracic Oncology. 2011;6:1313–1319. doi:10.1097/JTO.0b013e318220caff
22. Fre ´de ´ric LJ and Ueli SS. Combined HDAC1 and HDAC2 Depletion Promotes Retinal Ganglion Cell Survival After Injury Through Reduction of p53 Target Gene Expression. American Society for Neurochemistry. 2015;7(3). doi:10.1177/1759091415593066
23. Reed SM and Quelle DE. p53 Acetylation: Regulation and Consequences. p53 Acetylation: Regulation and Consequences. 2015;7:30–69. doi:10.3390/cancers7010030.
24. Zhang Y, Dube C, Gibert Jr.M, et al. The p53 Pathway in Glioblastoma. Cancers. 2018;10(297). doi:10.1073/pnas.1431692100.
 © 2021 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.
© 2021 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.