Back to Journals » Cancer Management and Research » Volume 12
ARHGAP6 Promotes Apoptosis and Inhibits Glycolysis in Lung Adenocarcinoma Through STAT3 Signaling Pathway
Authors Li P, Lv H, Xu M, Zang B, Ma Y
Received 10 April 2020
Accepted for publication 21 July 2020
Published 6 October 2020 Volume 2020:12 Pages 9665—9678
DOI https://doi.org/10.2147/CMAR.S257759
Checked for plagiarism Yes
Review by Single anonymous peer review
Peer reviewer comments 2
Editor who approved publication: Professor Yong Teng
Pengfei Li,1,* Huina Lv,2,* Min Xu,1 Bin Zang,1 Yegang Ma1
1Department of Thoracic Surgery, Cancer Hospital of China Medical University, Liaoning Cancer Hospital and Institute, Shenyang 110042, People’s Republic of China; 2Department of Ultrasound, Shengjing Hospital of China Medical University, Shenyang, 110004, People’s Republic of China
*These authors contributed equally to this work
Correspondence: Yegang Ma Email [email protected]
Objective: Constitutively activated signal transducer and activator of transcription 3 (STAT3) has been linked to cisplatin (DDP)-resistance in a wide range of cancers. Recent work has indicated that Rho GTPase-activating protein 6 (ARHGAP6) promotes cell cycle arrest and apoptosis in cervical and breast cancers. However, the role of ARHGAP6 in lung adenocarcinoma and DDP-resistance remains unknown.
Materials and Methods: Bioinformatic analysis, quantitative RT-PCR and IHC staining were used to explore ARHGAP6 expression patterns in The Cancer Genome Atlas (TCGA) dataset and patient samples. Statistical analysis was performed to establish the association of ARHGAP6 expression with the resistance to DDP-based chemotherapy in lung adenocarcinoma patients. Functional assays were then conducted to examine the effect of ARHGAP6 on the apoptosis and glycolysis in DDP-resistant/sensitive A549/DPP cells in vitro. Finally, the effects of ARHGAP6 on the chemosensitivity of DDP were explored in vivo.
Results: We show that decreased ARHGAP6 levels are a reliable marker of lung adenocarcinoma across published datasets, cell culture lines, and clinical samples. Low ARHGAP6 expression was linked to decreased apoptosis and increased metabolic activity, which highlights ARHGAP6’s role as a tumor suppressor. Furthermore, activated p-STAT3 levels increased dramatically in the absence of ARHGAP6, which suggests that ARHGAP6 can inhibit the STAT3 pathway. In agreement with previous studies that linked p-STAT3 levels to DDP-resistance, our in vitro and in vivo data indicate that tumors became more resistant to DDP-therapy with reduced ARHGAP6 levels and an associated increase in p-STAT3.
Conclusion: ARHGAP6 presents a novel study target for overcoming p-STAT3-associated DDP-resistance in lung adenocarcinoma and potentially other cancers.
Keywords: lung adenocarcinoma, ARHGAP6, STAT3, cisplatin, RhoA, cisplatin-resistance
Introduction
Despite an overall decline in the incidence and mortality, lung adenocarcinoma accounts for about 40% of all lung cancer and continues to be a leading cause of cancer mortality worldwide.1 Because the majority of lung adenocarcinomas are incurable and patients have a high risk of relapse after surgery, chemotherapy is often used. Despite significant advances in chemotherapy, patient survival remains poor. A major cause of this continued mortality is the development of resistance against established therapies. Cisplatin (DDP), for example, is one of the most common treatments for a wide range of cancers, yet acquired resistance to DPP-therapy continues to pose a significant challenge.2 Inhibitors of signal transducer and activator of transcription-3 (STAT3), a transcription factor that promotes cell survival and proliferation, have been shown to improve the clinical effectiveness of DDP.3,4 This suggests that targeting the active, phosphorylated form of STAT3, p-STAT3, is a potential strategy for overcoming resistance to DDP-therapy.4
In normal cells, STAT3 is transiently activated (p-STAT3) for half an hour up to several hours. However, in solid and hematological tumors, STAT3 is commonly overexpressed or constitutively activated,5 and this phenomenon has been associated with poor overall survival rates in patients with lung cancer.6,7 Aberrant STAT3 activation upregulates the expression of genes implicated in cell survival and proliferation, such as c-Myc and survivin, which promotes cancer development and progression.8,9 c-Myc, responsible for the upregulation of glycolytic proteins such as hexokinase 2 (HK2), is required to sustain the increased metabolic demands of a cancerous cell.10,11
The STAT3 signaling cascade, including its upstream regulators, harbors multiple promising therapeutic targets. Rho GTPase-activating protein 6 (ARHGAP6), a signaling molecule critical in controlling STAT3 activation, acts through the intermediary, RhoA. RhoA directly activates nuclear STAT3 through tyrosine phosphorylation.12–14 ARHGAP6 inhibits RhoA and prevents the activation of STAT3 and subsequent cell survival pathways.13 Accordingly, ARHGAP6 promotes cell cycle arrest and induces apoptosis in some cervical carcinomas.12,13,15 Reduced levels of ARHGAP6 have also been seen in tumor tissue from patients with lung cancer.1 Furthermore, in vitro upregulation of ARHGAP6 inhibited the growth of A549 lung cancer cells and reduced p-STAT3 levels.16
In this study, we found that primary lung adenocarcinoma tissue shows a significant reduction in ARHGAP6 mRNA expression as compared to that in normal adjacent tissue. In these tissue samples, low ARHGAP6 levels served as a robust biomarker for lung adenocarcinoma and correlated with a worse prognosis. Knockdown of ARHGAP6 inhibited apoptosis and resulted in an increase in glycolysis in DDP-sensitive A549 cells through STAT3-signaling, reminiscent of a cancerous phenotype. Conversely, overexpression of ARHGAP6 mRNA in vitro reduced survival and metabolism in DDP-resistant A549 cells. Increased levels of ARHGAP6 enhanced the chemosensitivity of cells to DDP in vitro and in vivo. These results indicate that the upregulation of ARHGAP6 may be useful in the treatment of DDP-resistant lung adenocarcinomas.
Materials and Methods
Bioinformatic Analysis
The mRNA RNAseq data related to ARHGAP6 of patients with lung adenocarcinoma, including 515 cancerous and 59 normal adjacent lung tissues, and other cancers were collected from The Cancer Genome Atlas (TCGA) dataset. Kaplan–Meier analysis of overall survival in patients with lung adenocarcinoma was obtained from the GSE31210 dataset.
Clinical Samples
Tumor samples were collected from 120 treatment-naïve patients, with lung adenocarcinomas and no co-malignancies, undergoing radical surgery at the Liaoning Cancer Hospital and Institute between January 2010 and July 2013. Ninety-seven patients received either chemo- or radiotherapy in the post-operative period. Among those patients receiving adjuvant therapy, 75 underwent three or more rounds of DDP chemotherapy. All patients signed an informed consent form before participation. The study was approved by the Ethics Committee of the Liaoning Cancer Hospital and Institute. Tissue was flash-frozen in liquid nitrogen before long-term storage at −80°C. Progression-free survival refers to the time of diagnosis to disease progression, relapse, or death. Overall survival refers to the time of diagnosis to death. Death may be the result of any cause.
Immunohistochemistry (IHC)
Lung adenocarcinoma (n=120) and the normal adjacent lung (n=35) tissues were embedded in paraffin and used for IHC studies. Samples were incubated in primary antibody anti-ARHGAP6 (PA5-57812; Thermofisher; 1:100 dilution) or p-STAT3 (ab76315; Abcam; 1:100 dilution), followed by incubation in secondary antibody (D-3004; Shanghai Long Island Biotec. Co., Ltd). Two pathologists, without prior knowledge of each sample’s origin, quantified the stained samples.
Cell Culture
A549 and the cisplatin-resistant (A549/DDP) cell lines (JRDUN Biotech., Shanghai, China) were grown in RPMI 1640 medium (Hyclone, Logan, UT, USA) whereas 16HBE cells were cultured in DMEM medium (Invitrogen, Gaithersburg, MD, USA) supplemented with 10% FBS serum (Gibco, Carlsbad, CA, USA) at 37 °C and 5% CO2.
RNA Interference and Gene Overexpression
To create the overexpression line, the coding sequence of ARHGAP6 was inserted into a pLVX-Puro plasmid (Clontech, USA). For the knockdown lines, the following RNA interference sequences targeting ARHGAP6 were designed and inserted into linearized pLKO.1 plasmids (Addgen, USA): ARHGAP6 (shRNA-1, position 1957–1975, 5ʹ-GCGAATGACAGGGCCTATA-3ʹ; shRNA-2, position 2607–2625, 5ʹ-GCTCCTCATATACCTTCTA-3ʹ; shRNA-3, position 2787–2805, 5ʹ-GCAGAAGTCATCAGACAAA-3ʹ). Complete plasmids (1000 ng) were then co-transfected with the packaging plasmids psPAX2 (900 ng) and pMD2-G (100 ng) into HEK-293T cells in serum-free medium using Lipofectamine-2000 (Invitrogen) in accordance with the manufacturer’s protocol. After 4–6 hours transfection, the medium was replaced with a fresh complete medium. After 48 hours incubation, A549 and A549/DDP cells were transduced with the recombinant lentiviral vectors units at an MOI of 20 in the presence of 8 μg/mL polybrene (Sigma-Aldrich, St. Louis, MO, USA). Cells transduced with pLKO.1-scramble shRNA (shNC) or blank pLVX-Puro (vector) were used as negative controls.
CCK-8 Assay
A549 and A549/DDP cell proliferation assay was based on the Cell Counting Kit (CCK-8; Dojindo, Kumamoto, Japan) method. Cells were seeded in a 96-well plate at a density of 3 × 103 cells/well for 12 h. Cells were transduced with indicated vectors in the absence or presence of 10 μM DDP and allowed to grow for 0, 24, 48 and 72 h. At the end of the experiment, CCK-8 reagent was added to each well and cultured for 1 h at 37°C and 5% CO2. The absorbance of each well was determined at 450 nm.
Cell Apoptosis Assay
A549 and A549/DDP cells were seeded at 3 × 105 cells/well in six-well plates and allowed to grow to 50% confluence. Cells were transduced with the relevant vector in the absence or presence of AG490 (10 μM) or DDP (10 μM), and 48 hours later, apoptosis rates were assessed. Briefly, 5 μL Annexin V-FITC was added to cells, and the cells were incubated in the dark for 15 minutes at 4°C, followed by 5 μL of propidium iodide (PI) for 15 minutes at 4°C before proceeding to flow cytometry.
Measurement of Glucose Uptake
A549 and A549/DDP cells were seeded at 5 × 105 cells/well in six-well plates and grown for one day at 37°C and 5% CO2. Cells were transduced with the relevant vector in the absence or presence of AG490 (10 μM), and 48 hours later, media was replaced with low-glucose DMEM for 3 hours. Cells were then washed with Krebs-Ringer Bicarbonate Buffer with 2% BSA and incubated in glucose-free DMEM containing 100 μM of 2-NBDG (Cayman Chemical, Ann Arbor, Michigan, USA) for 45 min. Flow cytometry was used to measure the uptake of glucose by cells.
Metabolic Assays
A549 and A549/DDP cells were seeded at 5 × 105 cells/well in six-well plates and grown for one day at 37°C and 5% CO2. Cells were transduced with the relevant vector in the absence or presence of AG490 (10 μM), and 48 hours later, a lactic acid assay kit (Nanjing Jiancheng Bioengineering Institute, China) was used to determine lactate production. The manufacturer’s instructions were followed.
Quantitative RT-PCR
16HBE, A549 and A549/DDP cells were seeded at 5 × 105 cells/well in six-well plates and grown for one day at 37°C and 5% CO2. Cells were transduced with or without the relevant vector, and 48 hours later, total RNA from these cell lines was extracted using TRIzol reagent (Life Technologies, Inc., Waltham, MA, USA) according to the manufacturer’s instruction. Similarly, TRIzol reagent was also used to extract total RNA from the lung adenocarcinomas tissues as mentioned above. Following the manufacturer’s protocol, the PrimeScript kit (Takara Biotechnology, Dalian, China) was used to synthesize cDNA. qPCR was performed in an ABI 9700 real-time PCR system (Applied Biosystem) using SYBR green PCR master mix (Applied Biosystems, Foster, CA, USA). The primers used are as follows: ARHGAP6-F: 5ʹ-GAATTTGACCGTGGGATTG-3ʹ, ARHGAP6-R: 5ʹ-CAGGGAGGTAGAAGGTATATG-3ʹ; GAPDH-F: 5ʹ-AATCCCATCACCATCTTC-3ʹ, GAPDH-R: 5ʹ-AGGCTGTTGTCATACTTC-3ʹ. The fold changes of mRNA were determined by the 2−ΔΔCT method.
Western Blot Analysis
16HBE, A549 and A549/DDP cells were seeded at 5 × 105 cells/well in six-well plates and grown for one day at 37°C and 5% CO2. Cells were transduced with or without the relevant vector in the absence or presence of AG490 (10 μM), and 6 hour (measurement for STAT3 and p-STAT3 level) and 48 hours (measurement for other protein levels) later, RIPA lysis buffer, freshly supplemented with protease inhibitor (Sigma, St. Louis, MO, USA), was used to lyse these cells. Similarly, lung adenocarcinomas tissues were also lysed as mentioned above. Proteins were then separated by SDS-PAGE gel and transferred onto a nitrocellulose membrane (Millipore, Bedford, USA). Membranes were blocked with 5% skim milk and incubated with antibodies against ARHGAP6 (PA5-57812; Invitrogen), c-Myc (ab32072; Abcam), PFKL (ab181064; Abcam), HK2 (227198; Abcam), Survivin (#2802; CST, Danvers, MA, USA), STAT3 (ab19352; Abcam), p-STAT3 (phospho Y705) (ab76315; Abcam), GAPDH (5174; CST), and HRP-conjugated secondary antibodies (A0208; Beyotime, Shanghai, China). An enhanced chemiluminescence system (Bio-Rad, Richmond, CA, USA) was used to visualize the resulting signal.
Establishment of Stable Cell Lines and Xenograft Study
To establish the patient-derived xenograft (PDX) model, tumor tissue samples were collected at the time of interventional surgery at Liaoning Cancer Hospital and Institute from lung adenocarcinoma patients who had not received chemotherapy. The patient samples were divided into two groups. One group was fixed for detection of ARHGAP6 expression in Quantitative RT-PCR, and the remaining fresh tumor tissues were cut into pieces and subcutaneously transplanted into 6-8-week-old nude mice (5×5×3 mm3 per mouse) within 1 h of surgical excision from patients. DDP (5 mg/kg; every week) chemotherapy was initiated at day 12 after injection (n=5 per group). At day 33 after injection, mice were sacrificed and tumor size was measured. The xenografts were subjected to TUNEL (Terminal deoxynucleotidyl transferase dUTP Nick-End Labeling) analyses (Roche, Indianapolis, IN, USA). Animal ethics guidelines and approved protocols of Liaoning Cancer Hospital and Institute were followed for all experiments on animals (2015PS152K).
Statistical Analysis
All experiments were performed in triplicate, and the results are expressed as mean ± SD. GraphPad Prism 7.0 (GraphPad Software, San Diego, CA, USA) was used for all statistical analyses. ANOVA followed by Tukey’s post-multiple test was used to compare different experimental groups. Kaplan–Meier method and Cox’s proportional hazards regression model were used to calculate overall survival and progression-free survival. Differences were analyzed by a Log rank test. A P value of less than 0.05 was considered to statistically significant.
Results
Reduced ARHGAP6 mRNA Expression is a Biomarker for Adenocarcinoma in Lung Tissue and Correlates with a Worse Prognosis
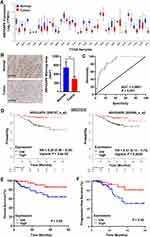
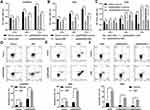
Screening of The Cancer Genome Atlas (TCGA) dataset, including 20,000 malignant and normal samples across 33 cancer types, revealed a reduction in ARHGAP6 mRNA expression across a range of cancers (Figure 1A). Among those, 515 lung adenocarcinoma tissue samples showed significantly lower ARHGAP6 mRNA expression levels compared to that in 59 normal adjacent tissue samples (P<0.001). Immunohistochemical staining also shows the lower expression of ARHGAP6 protein in tumor tissue (P<0.01) (Figure 1B).
The diagnostic value of ARHGAP6 mRNA expression was determined by receiver operating characteristics (ROC) curve analysis (Figure 1C). With an area under the curve of 0.8681 (P<0.001), ARHGAP6 mRNA is a useful biomarker for identifying patients with lung adenocarcinoma. Based on the ROC curve, ARHGAP6 expression level at the value of 0.58 was selected as the cut-off value to define ARHGAP6low and ARHGAP6high expression. Using the defined high and low ranges of ARHGAP6 mRNA expression, a variety of clinicopathological features were analyzed (Table 1). In those patients with reduced levels of ARHGAP6 mRNA, there was a worsening in tumor size, metastasis to lymph nodes and disease stage. However, age, gender, smoker status, and tumor grade did not correlate with the level of ARHGAP6 mRNA present.
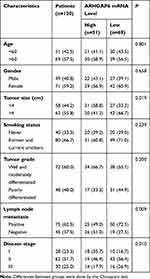 |
Table 1 Association of ARHGAP6 Expression with Clinical Characteristics of 120 Patients with Lung Adenocarcinoma |
Survival rates of each group were then measured for patients in the “gene expression data for pathological stage I–II lung adenocarcinomas” (GSE31210) dataset using two probe sets, 206,167 and 208,085 (Figure 1D, right and left), respectively. ARHGAP6low expression levels, measured with two separate probe sets, correlated with reduced survival in patients with lung adenocarcinoma. Similar results were seen in clinical samples: low ARHGAP6 mRNA levels correlated with worse overall and progression-free survival (Figure 1E-F). We next performed a univariate analysis of prognostic factors for overall survival with the Cox regression model (Table 2). ARHGAP6 expression (p=0.016, hazard ratio [HR]=1.20, 95% confidence interval [CI]=1.04–1.41) and lymph node metastasis (p=0.012, HR=0.82, 95% CI=0.71–0.95) were independent prognostic indicators of lung adenocarcinomas patients. We then performed a multivariate analysis with the Cox regression model (Table 2). ARHGAP6 expression (p=0.004, HR=0.79, 95% CI=0.68–0.92) was an independent prognostic factor in multivariate analysis. All of these data demonstrate that ARHGAP6 is a prognostic factor and may act as a tumor suppressor to prevent the progression of lung adenocarcinoma.
 |
Table 2 Univariate and Multivariate Analysis of Overall Survival in Patients with Lung Adenocarcinoma |
ARHGAP6 mRNA Expression Levels Correlate with the Responsiveness of Lung Adenocarcinoma Tissue to DDP-Therapy
Among patients receiving adjuvant therapy, 75 patients with lung adenocarcinoma underwent three or more rounds of DDP chemotherapy (Table 3) were separated into a DDP-sensitive (n=48) and a DDP-resistant group (n=27).17 Twenty-two of the DDP-resistant tumors (81.5%) showed ARHGAP6low mRNA expression levels (P<0.01). However, only five of the 48 DDP-sensitive tumors (10.4%) showed ARHGAP6low mRNA levels (P<0.01). These results suggest that the role of ARHGAP6 in lung adenocarcinoma and responsiveness to DDP-therapy is of great clinical and scientific interest.
 |
Table 3 The Therapeutic Status for Patients with Lung Adenocarcinoma After Surgery |
Overexpression of ARHGAP6 Enhances Apoptosis and Reduces Glycolysis in DDP-Resistant A549/DDP Cells
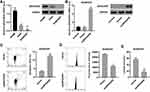
We began investigating ARHGAP6’s role in lung adenocarcinoma by measuring the relative amount of ARHGAP6 across several lung cell lines. ARHGAP6 mRNA and protein expression levels were reduced in the lung adenocarcinoma cell line, A549, compared to those in the healthy human bronchial epithelial cell line 16HBE (P<0.001). A549/DPP cells, the A549 cells that were resistant to DDP-therapy, showed a further reduction in ARHGAP6 mRNA and protein expression (P<0.001) (Figure 2A). We developed an overexpression system using these lowest expressing, treatment-resistant cells for further experiments.
ARHGAP6 levels were increased in the A549/DDP cells that were transduced with an ARHGAP6 overexpressing lentivirus (Figure 2B). Flow cytometric analysis of the transduced cells showed a significant increase in apoptotic activity (Figure 2C) and a reduction in glucose uptake by the cells (Figure 2D). The reduction in metabolic activity was also confirmed by decreased lactate production by ARHGAP6-overexpressing cells (Figure 2E). Together, these results indicate that ARHGAP6 serves as a tumor suppressor to counter the pro-survival signaling in lung adenocarcinoma cells.
The Knockdown of ARHGAP6 Inhibits Apoptosis and Promotes Glycolysis in DDP-Sensitive A549 Cells
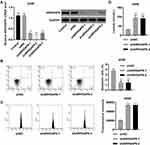
Next, we replicated the ARHGAP6low environment by establishing an in vitro knockdown model. DDP-therapy sensitive A549 cells were transduced with one of three targeted short hairpin RNAs (shARHGAP6-1, −2, and −3) or scrambled control (shNC). ARHGAP6 mRNA and protein expression were both significantly reduced in A549 cells transduced with shARHGAP6-1, −2, and −3, especially in A549 cells transduced with shARHGAP6-1 and −2; the control showed no effect (P<0.001) (Figure 3A).
Flow cytometric analysis of cells transduced with shARHGAP6-1 or −2 revealed that decreased expression of ARHGAP6 reduced cell death (Figure 3B) and increased glucose uptake by cells (Figure 3C) compared to those transduced with shNC (P<0.001). Lactate production was also increased following a reduction in ARHGAP6 expression, which indicates an increased metabolic demand (Figure 3D). In the absence of normal ARHGAP6 levels, cells showed greater survival rates and metabolic demands. These results support ARHGAP6’s role in suppressing uncontrolled cell growth and survival.
ARHGAP6 Regulates Apoptosis and Glycolysis Through the STAT3 Signaling Pathway
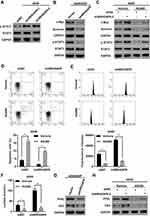
We then investigated how ARHGAP6 affects cell survival and metabolism. Western blot analysis of the ARHGAP6-knockdown model revealed an increase in p-STAT3 protein levels (Figure 4A). Conversely, the overexpression model showed decreased p-STAT3 protein expression and those of downstream effectors, c-Myc and survivin (Figure 4B). The change in ARHGAP6 did not appear to influence the level of STAT3; it only affected the level of activation.
To determine whether STAT3 activation accounts for the effect of ARHGAP6 on cell apoptosis and glycolysis, we added AG490, a specific Jak2/STAT3 inhibitor, to DDP-sensitive A549 cells, with and without ARHGAP6 knockdown. As expected, knockdown of ARHGAP6 markedly elevated p-STAT3 protein levels. Strikingly, the increase in p-STAT3 protein expression induced by ARHGAP6 knockdown was inhibited by the addition of AG490 (Figure 4C), indicating a functional relationship among ARHGAP6 and p-STAT3 in DPP-sensitive A549 cells. Moreover, the treatment with AG490 also reversed the effect of ARHGAP6 on cell apoptosis and glycolysis. As shown in Figure 4D–F, knockdown of ARHGAP6 significantly decreased cell apoptosis but increased glucose uptake and lactose production, but these effects were abolished upon the treatment with AG490. Furthermore, glycolytic proteins, such as liver-type phosphofructokinase (PFKL) and hexokinase-2 (HK2), exhibited a modest decrease in the ARHGAP6 overexpression model (Figure 4G), but they showed a modest increase upon ARHGAP6 knockdown (Figure 4H). However, the increase in PFKL and HK2 protein expression induced by ARHGAP6 knockdown was abolished by the addition of AG490 (Figure 4H). These results collectively support the hypothesis that ARHGAP6 regulates the apoptosis and glycolysis via the STAT3 signaling pathways.
ARHGAP6 Influences DDP-Induced Apoptosis in vitro
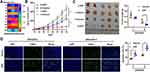
Since we found that STAT3 inhibitors could improve the effectiveness of DDP-therapy, we administered DDP to the overexpression and knockdown model cell lines. When ARHGAP6 was overexpressed in DDP-resistant and DDP-sensitive cells, the response to DDP was improved, evidenced by decreased cell proliferation (Figure 5A and B) and increased cell apoptosis (Figure 5C and D). In contrast, DDP-sensitive cells that had ARHGAP6 knocked down had a reduced response to DDP (Figure 5E and F), which indicates that ARHGAP6 played a role in the initial sensitivity.
Higher ARHGAP6 Levels Enhanced the Chemosensitivity of DDP in vivo
Finally, we determined if ARHGAP6’s impact on DDP-sensitivity was reproducible in a mouse model. Tissue samples were collected from patients with lung adenocarcinoma, fixed for detection of ARHGAP6 expression in Quantitative RT-PCR and characterized as ARHGAP6 high or ARHGAP6 low groups (median value used as cut-off) (Figure 6A). Tumors with low levels of ARHGAP6 showed larger tumor volume, as well as a reduced response to DDP-therapy (Figure 6B). ARHGAP6high expression tumors had a smaller volume, and tumor growth was markedly more sensitive to DDP-therapy.
On day 33, mice were sacrificed, and final tumor size and weight were measured. Again, the ARHGAP6high tumors were smaller after excision and showed a significantly reduced weight following DDP-therapy (Figure 6C). The tumors were then analyzed by the TUNEL assay to detect DNA breaks indicative of the final stages of apoptosis. ARHGAP6high tumors displayed a higher percentage of apoptotic cells and a more dramatic increase in apoptotic cells following DDP-therapy (Figure 6D). Taken together, these results suggest that higher expression of ARHGAP6 enhances the sensitivity of lung adenocarcinoma cells to DDP chemotherapy.
Correlation Analyses in Lung Adenocarcinoma Tissues
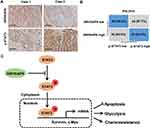
Immunohistochemical staining showed the expression of ARHGAP6 and p-STAT3 expression in tumor tissue (Figure 7A). Interestingly, the samples with ARHGAP6 higher expression displayed low levels for p-STAT3. In addition, samples with low expression of ARHGAP6 appeared strongly staining of p-STAT3. Statistically, ARHGAP6 expression was negatively correlated with p-STAT3 expression in lung adenocarcinoma tissues (Figure 7B).
Discussion
In this study, we have shown that ARHGAP6 acts as a tumor suppressor and can serve as a biomarker for lung adenocarcinoma. Screening of TCGA revealed that ARHGAP6 expression in lung adenocarcinoma tissue was markedly lower than that in normal adjacent tissue. These results are in line with recent studies that showed a reduction in ARHGAP6 levels in breast, cervical, and lung cancer tissue.15,16,18 The relationship between reduced expression and tumor phenotype held true for both cell culture lines and novel patient samples. Using ROC curve analysis, we were able to define ARHGAP6high and ARHGAP6low mRNA expression levels as a reliable biomarker of lung adenocarcinoma. ARHGAP6low levels correlated with a number of poor clinicopathological outcomes, including resistance to DDP-therapy and poor overall survival. Consistent with our results, breast cancer patients with ARHGAP6low mRNA expression also showed similar low overall survival rates.18
ARHGAP6 regulates the survival, metabolism, and proliferation of lung cancer cells. Recent work has shown that ARHGAP6 regulates the proliferation of lung cancer cells, in part, by reducing the levels of p-STAT3.16 Activated STAT3 serves as a transcription factor that is responsible for the activation of numerous pro-proliferative and survival genes. One such gene is c-Myc, a well-studied oncogene.19 Here, we show that reduction of ARHGAP6 led to increased levels of p-STAT3, as well as downstream targets like c-Myc and survivin. To our knowledge, ARHGAP6 has not previously been linked to these more distant oncogenes. Reduced levels of ARHGAP6 also resulted in unregulated metabolism. Consistent with STAT3-overactivation, ARHGAP6 suppression led to a dramatic increase in glycolytic enzymes PFKL and HK2, which was measured by lactate production, and ultimately contribute to anaerobic respiration. Further exploration into the link between ARHGAP6, the downstream targets of STAT3, and metabolic demands would be of great clinical importance in cancer research.
ARHGAP6’s action as a STAT3-inhibitor has great critical importance because STAT3 is credited for causing the bulk of acquired drug-resistance clinically. A great deal of research has focused on identifying functional inhibitors of STAT3 for clinical use.4 In this study, we focused on resistance to DDP, a common treatment for lung adenocarcinoma. As expected, there was a direct correlation between ARHGAP6 expression and DDP-sensitivity. Interestingly, freshly collected patient tissue showed that increased levels of ARHGAP6 led to an improved response to DDP-treatment. This exciting result has major implications for the treatment of lung adenocarcinoma.
The mechanism by which ARHGAP6 inhibits the STAT3 pathway remains unclear. However, ARHGAP6 is known to inactivate the tyrosine kinase RhoA, a molecule responsible for the phosphorylation and activation of nuclear STAT3.7,12 This prevents the transcription factor, p-STAT3, from targeting several genes implicated in tumorigenesis. We hypothesize that ARHGAP6 is indirectly inhibiting the activation of STAT3 by inhibiting RhoA. c-Myc is a cell cycle regulator that also works in concert with STAT3 to regulate the transcription of proto-oncogene Cdc25A.20,21 As a Ras Regulator, Cdc25A could be involved in a potential feedback mechanism and warrants further research, alongside ARHGAP6.22 The full mechanism by which ARHGAP6 regulates STAT3 is of critical importance for future research. Taken together, the results of this study suggest ARHGAP6 negatively regulates the STAT3 pathway, a known cause of acquired DDP-resistance in lung adenocarcinoma.
Conclusion
Our study highlights the importance of ARHGAP6 in maintaining DDP-sensitivity in lung adenocarcinoma cells and identifies it as a potential biomarker for lung adenocarcinoma. We show that that activation of the STAT3 pathway is a critical component in the interaction between ARHGAP6 and DDP-resistance (Figure 7C). ARHGAP6 presents a novel research target for overcoming p-STAT3-associated DDP-resistance in lung adenocarcinoma and potentially other cancers.
Disclosure
The authors report no competing financial interests in relation to this work.
References
1. Ferlay J, Colombet M, Soerjomataram I, Mathers C, Parkin DM. Estimating the global cancer incidence and mortality in 2018: GLOBOCAN sources and methods. Int J Cancer. 2019;144(8):1941–1953. doi:10.1002/ijc.31937
2. Achkar IW, Abdulrahman N, Al-Sulaiti H, Joseph JM, Uddin S, Mraiche F. Cisplatin based therapy: the role of the mitogen activated protein kinase signaling pathway. J Transl Med. 2018;16(1):96. doi:10.1186/s12967-018-1471-1
3. Abroun S, Saki N, Ahmadvand M, Asghari F, Salari F, Rahim F. STATs: an old story, yet mesmerizing. Cell J. 2015;17(3):395–411. doi:10.22074/cellj.2015.1
4. Sun CY, Nie J, Huang JP, Zheng GJ, Feng B. Targeting STAT3 inhibition to reverse cisplatin resistance. Biomed Pharmacother. 2019;117:109135. doi:10.1016/j.biopha.2019.109135
5. Morita Y, Naka T, Kawazoe Y, et al. Signals transducers and activators of transcription (STAT)-induced STAT inhibitor-1 (SSI-1)/suppressor of cytokine signaling-1 (SOCS-1) suppresses tumor necrosis factor alpha-induced cell death in fibroblasts. Proc Natl Acad Sci U S A. 2000;97(10):5405–5410. doi:10.1073/pnas.090084797
6. Tong M, Wang J, Jiang N, Pan H, Li D. Correlation between p-STAT3 overexpression and prognosis in lung cancer: a systematic review and meta-analysis. PLoS One. 2017;12(8):e0182282. doi:10.1371/journal.pone.0182282
7. Vega FM, Ridley AJ. Rho GTPases in cancer cell biology. FEBS Lett. 2008;582(14):2093–2101. doi:10.1016/j.febslet.2008.04.039
8. Gritsko T, Williams A, Turkson J, et al. Persistent activation of stat3 signaling induces survivin gene expression and confers resistance to apoptosis in human breast cancer cells. Clin Cancer Res. 2006;12(1):11–19. doi:10.1158/1078-0432.CCR-04-1752
9. Kiuchi N, Nakajima K, Ichiba M, et al. STAT3 is required for the gp130-mediated full activation of the c-myc gene. J Exp Med. 1999;189(1):63–73.
10. Nakajima K, Kawashima I, Koshiisi M, et al. Glycolytic enzyme hexokinase II is a putative therapeutic target in B-cell malignant lymphoma. Exp Hematol. 2019;78:46–55.e43. doi:10.1016/j.exphem.2019.09.023
11. Zhang S, Li J, Xie P, et al. STAT3/c-Myc axis-mediated metabolism alternations of inflammation-related glycolysis involve with colorectal carcinogenesis. Rejuvenation Res. 2019;22(2):138–145. doi:10.1089/rej.2018.2089
12. Aznar S, Valeron PF, Del Rincon SV, Perez LF, Perona R, Lacal JC. Simultaneous tyrosine and serine phosphorylation of STAT3 transcription factor is involved in Rho A GTPase oncogenic transformation. Mol Biol Cell. 2001;12(10):3282–3294. doi:10.1091/mbc.12.10.3282
13. Prakash SK, Paylor R, Jenna S, et al. Functional analysis of ARHGAP6, a novel GTPase-activating protein for RhoA. Hum Mol Genet. 2000;9(4):477–488. doi:10.1093/hmg/9.4.477
14. Schaefer L, Prakash S, Zoghbi HY. Cloning and characterization of a novel rho-type GTPase-activating protein gene (ARHGAP6) from the critical region for microphthalmia with linear skin defects. Genomics. 1997;46(2):268–277. doi:10.1006/geno.1997.5040
15. Li J, Liu Y, Yin Y. Inhibitory effects of Arhgap6 on cervical carcinoma cells. Tumour Biology. 2016;37(2):1411–1425. doi:10.1007/s13277-015-4502-z
16. Wu Y, Xu M, He R, Xu K, Ma Y. ARHGAP6 regulates the proliferation, migration and invasion of lung cancer cells. Oncol Rep. 2019;41(4):2281–2888. doi:10.3892/or.2019.7031
17. Pan X, Chen Y, Shen Y, Tantai J. Knockdown of TRIM65 inhibits autophagy and cisplatin resistance in A549/DDP cells by regulating miR-138-5p/ATG7. Cell Death & Dis. 2019;10(6):429. doi:10.1038/s41419-019-1660-8
18. Chen WX, Lou M, Cheng L, et al. Bioinformatics analysis of potential therapeutic targets among ARHGAP genes in breast cancer. Oncol Lett. 2019;18(6):6017–6025. doi:10.3892/ol.2019.10949
19. Gao Q, Zhou R, Meng Y, et al. Long noncoding RNA CMPK2 promotes colorectal cancer progression by activating the FUBP3-c-Myc axis. Oncogene. 2020;39(19):3926–3938. doi:10.1038/s41388-020-1266-8
20. Chen H, Liu H, Qing G. Targeting oncogenic Myc as a strategy for cancer treatment. Signal Transduct Target Ther. 2018;3:5. doi:10.1038/s41392-018-0008-7
21. Galaktionov K, Chen X, Beach D. Cdc25 cell-cycle phosphatase as a target of c-myc. Nature. 1996;382(6591):511–517. doi:10.1038/382511a0
22. Ray D, Kiyokawa H. CDC25A phosphatase: a rate-limiting oncogene that determines genomic stability. Cancer Res. 2008;68(5):1251–1253. doi:10.1158/0008-5472.CAN-07-5983
 © 2020 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.
© 2020 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.