Back to Journals » OncoTargets and Therapy » Volume 11
Antibody-sandwich ELISA analysis of a novel blood biomarker of CST4 in gastrointestinal cancers
Authors Dou YL, Lv YL, Zhou XJ, He LF, Liu LH, Li PF, Sun YL, Wang MH, Gao MJ, Wang C
Received 15 August 2017
Accepted for publication 17 December 2017
Published 28 March 2018 Volume 2018:11 Pages 1743—1756
DOI https://doi.org/10.2147/OTT.S149204
Checked for plagiarism Yes
Review by Single anonymous peer review
Peer reviewer comments 2
Editor who approved publication: Dr Yao Dai
Yaling Dou,1,* Yali Lv,2,* Xiaojin Zhou,3,* Linfu He,4,* Lihong Liu,2 Pengfei Li,2 Yulong Sun,3 Minghui Wang,3 Meijuan Gao,3 Chong Wang3
1Department of Clinical Laboratory, Peking Union Medical College Hospital, Chinese Academy of Medical Sciences, Peking Union Medical College, Beijing, People’s Republic of China; 2Department of Pharmacy, Beijing Chao-Yang Hospital, Capital Medical University, Beijing, People’s Republic of China; 3Shanghai Liangrun Biomedicine Technology Co. Ltd., Shanghai, People’s Republic of China; 4Institute of Bioengineering, Jiangsu University, Zhenjiang, People’s Republic of China
*These authors contributed equally to this work
Background: Members of the cystatin family have increasingly been proven to be involved in several tumors, including gastric cancer (GC) and colorectal cancer (CRC). Cystatin S (CST4) was found to be upregulated at the gene expression level in GC cells, making it a potential novel biomarker for the early diagnosis of gastrointestinal cancer.
Materials and methods: Quantitative real-time polymerase chain reaction and Western blotting analysis were used to explore CST4 expression in gastrointestinal cancer tissues and cell lines. We purified CST4 recombinant protein and generated anti-CST4 monoclonal antibodies to develop an antibody-sandwich enzyme-linked immunosorbent assay (ELISA) analysis system for blood CST4 detection. The performance and clinical efficacy of the detection method were evaluated using a training set and validation set, respectively.
Results: According to the quantitative real-time polymerase chain reaction and Western blotting results, CST4-mRNA expression and protein expression were upregulated in gastrointestinal cancer tissues and cell lines. The ELISA detection system for CST4 showed significantly better sensitivities of 69.0% and 69.0% and specificities of 85.6% and 83.6% for GC and CRC, respectively, than other common clinical biomarkers, carcinoembryonic antigen, CA19-9, CA125, and CA72-4. Clinical verification experiments using GC and CRC validation sets also found distinguishable CST4 median concentrations (177.7 pg·mL-1 and 174.2 pg·mL-1 respectively) and high positive detection rates (72.3% and 88.4% respectively), further confirming the specificity and sensitivity of this method.
Conclusion: We validated the overexpression of CST4 in gastrointestinal cancer tissues and cell lines and developed an antibody-sandwich ELISA analysis system for blood CST4 detection, which exhibited high specificity and sensitivity. Novel blood biomarkers of CST4 have enormous potential in terms of clinical diagnostic value in GC and CRC.
Keywords: cystatin S, ELISA, gastrointestinal cancer, biomarker
Introduction
Gastric cancer (GC) and colorectal cancer (CRC) have the second and third highest mortality rates among all cancers, and are an enormous threat to public health.1,2 Owing to the high fatality rate and metastasis risk, early detection and therapy of these tumors are crucial for curative surgery and survival rate of patients.3 Clinically, fecal occult blood tests, fecal DNA tests,4 and colonoscopy are typically used in mass screening for CRC, while endoscopy, imageological examination, and oncogene testing are mainly used to detect GC. However, these methods are invasive, inconvenient in practice, inaccurate, or expensive.5–7 It seems that detection of circulating blood biomarkers will be a more appropriate method for early screening or auxiliary diagnosis of GC and CRC.
The current mainstream clinical auxiliary diagnostic markers for GC and CRC are carcinoembryonic antigen (CEA), CA19-9, CA125, and CA72-4; however, there is still an urgent need for novel biomarkers with higher sensitivity and specificity. Recent research concerning cystatins has supported their potential role as blood tumor biomarkers. For example, Yoneda et al found that cystatin SN (CST1) was notably upregulated in terms of gene expression in colorectal tissues compared with normal samples, and proposed a more sensitive enzyme-linked immunosorbent assay (ELISA) uniting CST1, CEA, and CA19-9.8 The cystatin M (CST6) gene was found to have lower expression in breast cancer tissues, and it was speculated that it may participate in tumor cell metastasis.9 In addition, cystatin C (CST3) has been implicated in tumor invasion and metastasis and is highly correlated with mortality rate in patients with CRC.10 The cystatin S (CST4), CST6, and CST3 genes are significantly upregulated in GC cells and may be important in gastric tumor progression.11
As a member of the cystatin superfamily, CST4 (also known as salivary acidic protein 1 or cystatin-SA-III) has a molecular weight of about 15 kD and two special disulfide bonds at the C terminus.12–14 According to Akiba et al, CST4 regulates cysteine protease activity by specifically combining with cysteine protease and it prevents hydrolysis of the extracellular matrix.15 Located in the cytoplasm, CST4 has the required characteristics of a blood biomarker (low molecular weight, secreted in blood, etc.). We propose that CST4 might be a biomarker, along with other cystatins, especially in gastrointestinal cancer.
In this research, we first explored the expression of CST4 in gastrointestinal cancer tissues and cells. Then, we developed an antibody-sandwich ELISA analysis system for blood CST4 detection and tentatively verified its clinical utility in gastrointestinal cancer diagnosis.
Materials and methods
Ethics statement
This study was approved by the ethics committees of the Peking Union Medical College Hospital and Beijing Chao-Yang Hospital. All human blood samples and gastrointestinal (cancer) tissues were obtained with written informed consent.
Materials
Materials and instruments
Fetal bovine serum (FBS) was purchased from Thermo Fisher Scientific (Waltham, MA, USA). Horseradish peroxidase (HRP) and 3,3′,5,5′-tetramethylbenzidine (TMB) were purchased from Beyotime Biotechnology (Jiangsu, People’s Republic of China). Endo Free Plasmid Kit was purchased from QIAGEN. GC cell lines MKN-45 and HGC-27, gastric mucosal cell lines GES-1 and RGM-1, CRC cell lines HCT-116 and SW480, and intestinal epithelial cell lines HIEC-6 and NCM-460 were purchased from BeNa Culture Collection (Shanghai, People’s Republic of China). Cells were cultured in RPMI1640 medium (Gibco-BRL, Grand Island, NY, USA) supplied with 10% FBS, penicillin (100 U·mL−1), and streptomycin (100 μg·mL−1) at 37°C in a cell incubator with 5% CO2. Radioimmunoprecipitation assay buffer, extraction buffer, and protein A/G beads were purchased from Beyotime Biotechnology. All other chemicals and reagents (which were of analytic grade) were purchased from Sino Pharm Chemical Reagent Co. Ltd. and used as received.
The chemiluminescence signal of TMB was detected with an iMARKT Microplate Reader (Bio-Rad, Hercules, CA, USA). The ultraviolet-visible light measurements were performed on a NanoDrop 2000 spectrometer (Thermo Fisher Scientific). The Bio-Rad 1575 Plate Washer was purchased from Bio-Rad.
Tissues and serum samples
Hundred tumor samples and 100 samples from the corresponding adjacent tissue for GC and CRC, respectively, were collected following surgery from Peking Union Medical College Hospital.
Overall, two sets of blood samples were collected to perform CST4 detection experiments, defined as the training set and validation set, respectively. For the training set, a total of 620 serum samples were collected from Peking Union Medical College Hospital, from patients with GC, CRC, benign gastric disease, benign colorectal disease, and other cancers, and from healthy people (detailed information is provided in Table S1). For the validation set, another 588 serum samples from patients diagnosed with GC, CRC, gastric diseases, colorectal diseases, gastrointestinal diseases, and other cancers were collected, as well as samples containing interfering substance (bilirubin, heme, and so on, detailed in Table S2), and controls from healthy people. These serum samples were obtained from Beijing Chao-Yang Hospital (detailed information is provided in Table 1).
All blood samples from patients were collected into vacuum blood tubes without any additive. Serum was separated after blood coagulation and distributed into fresh tubes. All the serum samples were stored at −80°C before use.
Components of detection reagent
Preparation and purification of CST4 recombinant protein
The cDNA was reversed transcribed with random primers, using total RNA extracted from gastric carcinoma tissues as the template.
F: CCCAAGCTTGCCACCATGGCCCAGTATCTGAGTACC (containing the HindIII restriction enzyme cutting site and Kozak sequence) was designed as the forward primer and R: GGATTCTTGACACCTGGATTTCAC (containing the BamHI restriction enzyme cutting site) was designed as the reverse primer to clone the CST4 sequence, with cDNA as the amplification template.
CST4 fragments and the pcDNA3.1 plasmid were double digested with HindIII and BamHI, and ligated to produce the recombinant plasmid CST4-pcDNA3.1, which contained a 6× His tag sequence. The ligation product was then transformed to DH5α Escherichia coli to yield abundant recombinant plasmids. After verification of the double digestion, agarose gel electrophoresis, and gene sequencing, 1 μg·μL−1 CST4-pcDNA3.1 was transformed into COS-7 cells by lipofectamine 2000, cultivated in DMEM containing 10% FBS at 37°C with 5% CO2 for 72 h. Cell culture medium was then collected and filtered through a 0.22 μm filter membrane for CST4 purification.
In order to collect a highly purified protein solution of CST4, both Ni-nitrilotriacetic acid affinity chromatography and anion exchange chromatography were applied using 500 mL of cell culture filtrate (previously collected and filtered). The equilibration buffer (pH 7.6) for the Ni-nitrilotriacetic acid affinity chromatography contained 50 mM PBS, 10 mM imidazole, and 150 mM NaCl. The elution buffer (pH 7.6) contained 50 mM PBS, 250 mM imidazole, and 150 mM NaCl. Ultrafiltration devices of molecular weight 3 kD were used to concentrate the obtained protein solution using an exchange buffer (pH 7.4) containing 20 mM PBS, 1 mM EDTA, and 10 mM NaCl. The collected recombinant protein eluant was purified by anion exchange chromatography. The equilibration buffer (pH 7.4) contained 20 mM PBS, 1 mM EDTA, and 10 mM NaCl; the elution buffer (pH 7.4) contained 20 mM PBS, 1 mM EDTA, and 250 mM NaCl. Purified protein samples were stored in a buffer (pH 7.4) containing 50 mM PBS, 150 mM NaCl, and 0.02% P300 at −80°C. The purity of purified protein was authenticated by sodium dodecyl sulfate polyacrylamide gel electrophoresis (SDS-PAGE) analysis.
Preparation and purification of antibodies
The experiments were performed with the approval of the Institutional Animal Care and Use Committee, Peking Union Medical College Hospital, and according to the rules of the Animal Welfare and “3Rs” (Replacement, Reduction, and Refinement) guidelines.
The purified recombinant CST4 protein (50 μg) was used to immunize BALB/c mice. Hybridoma cell strains secreting specific monoclonal antibodies were collected, and the subtypes of monoclonal antibodies were authenticated with a mouse antibody subtype identification kit (Baiaotong, Luoyang, People’s Republic of China). The selected hybridoma cells were injected into the abdomen of BALB/c mice, and the ascites were extracted after 7–10 d. The antibodies were purified from the ascites and analyzed by SDS-PAGE using a 10% gel.
Selection of antibodies
Monoclonal antibody titer was analyzed by ELISA. Each antibody was prepared at dilutions of 1:5,000, 1:10,000, 1:20,000, 1:40,000, and 1:80,000 for reaction with CST4 recombinant protein. One hundred microliters of goat anti-rabbit (GAR)–HRP conjugate was added into each well to initiate the reaction.
For monoclonal antibody epitope analysis, an ELISA-mediated antibody overlap experiment16 was performed. Briefly, any two monoclonal antibodies were added, either separately (saturation) or simultaneously (half of saturation), to react with CST4, and the amount of bound antibody was quantitatively measured by ELISA detection with the initial addition of GAR–HRP. The A450 values were measured as A1, A2, and A1 + A2, respectively, and the overlap index I was calculated according the formula: I = 2A1 + 2/(A1 + A2). Two monoclonal antibodies are considered to identify the different antigen epitopes when I is >0.5.
The matching experiments were used to choose appropriate capture and detection antibodies. The two antibodies with the highest positive-to-negative ratio (P/N) were chosen as the coating (capture) antibody and the labeling (detection) antibody.
The specificity tests of the capture and detection antibodies against CST family proteins (CST1, CST2, CST3, CST4, CST5, and CST6) were conducted using an ELISA assay with GAR–HRP.
Western blotting and immunoprecipitation
Cells were centrifuged and washed with PBS (pH 7.5) before collection. All cell samples were lysed with radioimmunoprecipitation assay lysis and extraction buffer and 1 mM phenylmethylsulfonyl fluoride (PMSF) on ice for 20 min. Lysates were centrifuged at 12,000 g and 4°C for 15 min, and supernatants were transferred for bicinchoninic acid protein assay (Beyotime). Fifty micrograms of total protein was loaded for 15% SDS-PAGE and then transferred onto a nitrocellulose filter membrane (EMD Millipore, Billerica, MA, USA). The membranes were blocked with 3% skimmed milk for 1 h and incubated with primary antibodies for 2 h at room temperature. After washing three times with tris-buffered saline and polysorbate 20, the membranes were incubated with HRP-conjugated secondary antibodies for 1 h at room temperature. An enhanced chemiluminescence kit (Beyotime) was used for detection.
Twenty microliters of protein A/G beads were added into a 2 mL centrifuge tube and washed with PBS (pH 7.5), after which 5 μg of capture antibody 5D2F2 was added and the mixture incubated on a turnover mixer at room temperature for 30 min. Five micrograms of CST4 recombinant protein was then added to the bead/antibody mixture, which was incubated on a turnover mixer at room temperature for 2 h. Immunoprecipitated products were centrifuged at 500 g for 2 min and resuspended in 100 μL PBS before Western blotting analysis with detection antibody 5E4G5.
RNA extraction and quantitative real-time polymerase chain reaction (qRT-PCR)
Total RNA was extracted using TRIzol reagent (Invitrogen, Carlsbad, CA, USA) according to the manufacturer’s protocols. RNA product quality was assessed by A260/A280 ratio, and RNA integrity was validated using an Agilent 2100 Bioanalyzer. Briefly, for each sample, 1 μg of total RNA was reverse transcribed into cDNA using Moloney Murine Leukemia Virus Reverse Transcriptase (NEB, Omaha, NE, USA) with random primers according to the manufacturer’s standard protocols. The quantitative polymerase chain reaction experiment was performed using a Light Cycler 480 device (Hoffman-La Roche Ltd., Basel, Switzerland) with a specific TaqMan probe (5′-FAM-ctccagctttgtgctctgcctctg-TAMRA-3′) and amplification primers. For each reaction, experiments were performed with the following 20 μL reaction volume: 2 μL 10X polymerase chain reaction (PCR) mix, 14.7 μL ddH2O, 0.5 μL forward and reverse primers (10 μM), 1 μL probe (10 μM), 1 μL cDNA template, and 0.3 μL Fast-Taq enzyme. The recombinant CST4-pcDNA3.1 plasmids were series diluted and used as reference templates to build standard curves. The CST4-mRNA copies in gastrointestinal tissues and corresponding adjacent tissues were calculated based on the cycle threshold value according to standard curves. Three biologic duplicates were prepared for each reaction, and U6 gene was chosen as the internal control.
Preparation of microplates
CST4 monoclonal antibody 5D2F2 was diluted to 2.5 μg·mL−1 with coating buffer and added into microplates at 100 μL per well. After coating at 4°C for 16 h, 200 μL enzyme-labeled plate stabilizing solution was added into wells and kept at 37°C for 1 h. Blocking solution was added and incubated for 1 h before vacuum drying at 30°C for 15 min. Dried microplates with desiccant were put into aluminum foil bags and vacuumed. All microplates were stored at 4°C until use.
Detection performance evaluation
Linear range
Linear range represents the effective concentration range of CST4 that the ELISA assay can detect. Twelve series-diluted concentrations of CST4 protein (2,800, 2,400, 2,000, 1,600, 1,200, 400, 200, 100, 50, 25, 12.5, and 6.25 pg·mL−1) were detected by double-hole detection. The linear range was confirmed when R≥0.99 and the minimum of the linear range was higher than the limit of detection (LOD).
LOD and precision
The diluent was detected as 0 pg·mL−1 20 times in order to determine the lower LOD. We calculated the mean (M) and SD of 20 detection results and used the formula M + 2SD to calculate the LOD.
The precision was evaluated using three different batches of ELISA kits when detecting two different concentration samples for three consecutive days. Each detection was repeated 10 times. According to M and SD, in-batch/batch-to-batch and in-day/day-to-day coefficients of variation (CV) were calculated with the equation: CV = SD/M ×100%. The detection concentrations of CST4 of two samples for CV analysis were 1,200 and 200 pg·mL−1.
Hook effect
The Hook effect appears when excessive antigen in a sample binds to limited available binding sites of antibodies, leading to failure to form the antibody–antigen–antibody sandwich and causing a measured OD value lower than the actual OD value. Concentrations of CST4 up to 12,800 pg·mL−1 were detected to investigate the Hook effect.
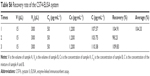
Recovery rate
The recovery rate was used to assess the accuracy of the ELISA assay. Briefly, a high-concentration CST4 sample (A) was added to a low-concentration CST4 sample (B) with a volume ratio (A:B) ≤1:9; that is, 15 μL of 1,200 pg·mL−1 reference A was added to 300 μL of 50 pg·mL−1 reference B. The recovery rate R was calculated with the equation, R = C × (V0 + V) − (C0 × V0)/(V × CS) ×100%, where V is the volume of sample A, V0 is the volume of sample B, C is the concentration of the mixture of A and B, C0 is the concentration of sample B, and CS is the concentration of sample A.
Experimental procedure for ELISA
The detection reagents, inspected samples, and other detection materials were equilibrated at room temperature for 30 min. The detection procedure was as follows.
Standard sample solutions were prepared as a gradient of six dilutions, and 100 μL of each dilution was added into wells to provide standard curves. Quality control protein solution (200 pg·mL−1) and serum samples were added to the wells simultaneously. The plate was sealed with sealing film and kept at 37°C for 60 min.
The reaction liquid was then removed, and the wells were washed three times. One hundred microliters of HRP anti-CST4 solution was added to each well and kept at 37°C for 60 min, after which the reaction liquid was removed and the wells washed three times again. One hundred microliters of TMB substrate solution was added to each well and kept at 37°C for 15 min.
Finally, 100 μL terminating solution was added and the OD value at 450 nm was read within 30 min.
Statistical methods
All experiments were repeated three times. All data are presented as M ± SD. Comparisons between groups were conducted using the Mann–Whitney, McNemar t-test, or the chi-square test. All statistical analyses were performed with SPSS 19.0 software. P<0.05 was considered to be statistically significant.
The cut-off value was chosen by a percentile method and the receiver-operating curve (ROC), as well as reference to the literature. The detection effect was evaluated according to the area under the ROC curve.
Results
Overexpression of CST4
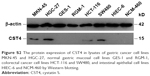
To verify the overexpression of CST4, qRT-PCR analysis of gastrointestinal tissues and Western blotting assay of gastrointestinal cell lysates were conducted. The qRT-PCR results (Figure S1) showed a significantly higher expression of CST4-mRNA in gastrointestinal cancer tissues compared with corresponding adjacent tissues. Positive bands (shown in Figure S2) of CST4 protein were also detected in lysates of gastrointestinal cancer cell lines (MKN-45, HGC-27, HCT-116, and SW480) in contrast to normal gastrointestinal cell lines (GES-1, RGM-1, HIEC-6, and NCM-460). The overexpression of CST4 in gastrointestinal cancer tissues and cells indicates its potential as a novel gastrointestinal biomarker.
Composition of the reaction system
Preparation of CST4 and antibodies
The agarose gel electrophoresis of the PCR amplification product and HindIII and BamHI double enzyme digestion product showed obvious positive bands at around 450 bp (Figure S3), proving that the CST4-pcDNA3.1 plasmids were successfully constructed. In addition, the gene sequencing result was consistent with the putative gene sequence of CST4 (Table S3).
The recombinant purified protein (Figure 1) was validated as being the P01036 protein, based on the alignment results with predicted sequence from the UniProt data bank, and its purity was over 95%.
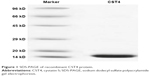  | Figure 1 SDS-PAGE of recombinant CST4 protein. |
Five monoclonal antibodies, 5D2F2, 5E4G5, 2G7A9, 3F4G8, and 4F1B6, were obtained. Their concentrations were determined by bicinchoninic acid as 5.4, 4.6, 3.7, 4.9, and 5.2 mg·mL−1, respectively.
The titer of antibodies was determined according to the maximum dilution rate of P/N >2.1, P–N >0.2. The titers of 5D2F2, 5E4G5, 2G7A9, 3F4G8, and 4F1B6 were 1:80,000, 1:40,000, 1:20,000, 1:20,000, and 1:20,000, respectively, which met the demand of further analysis.
Selection of paired antibodies
The types of epitopes (linear or conformation) were analyzed by Western blotting analysis (Figure S4). The results showed that monoclonal antibodies 5D2F2, 5E4G5, and 2G7A9 were capable of identification of the linear epitope of CST4 protein.
An antibody overlap test was performed to analyze the epitopes of antibodies. Each epitope respectively recognized by the 5D2F2, 5E4G5, 2G7A9, 3F4G8, and 4F1B6 monoclonal antibodies was confirmed. The results (Figure S4) showed that the overlap index of 5D2F2 and 5E4G5 was 0.86, which meant that 5D2F2 and 5E4G5 could identify different surface sites of CST4. The other overlap indices were all lower than 0.5, indicating probable overlap between the epitopes recognized by the antibodies. The Western blotting results showed that 5D2F2, 5E4G5, and 2G7A9 could identify linear surface sites, having titer type of both ELISA and Western blotting. Based on these results, 5D2F2 and 5E4G5 were selected for further analysis.
Matching experiments were used to choose appropriate capture and detection antibodies. As shown in Table S4, in terms of coated/detected antibodies, the negative ODs of 5D2F2/5E4G5 and 5E4G5/5D2F2 were both <0.1 and the P/N values were 22.26 and 17.74, respectively. Therefore, according to the standard, 5D2F2 was selected as the capture antibody and 5E4G5 as the detection antibody.
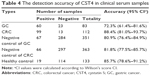
Specificity testing of the capture and detection antibodies (Table 2) showed that a basic weak signal was detected against CST1, CST2, CST3, CST5, and CST6 even at an extremely high protein concentration of 5 μg·mL−1. Conversely, a low concentration of CST4 (200 pg·mL−1) gave a positive signal, with an A450 value of 0.29. Notably, the specificities of 5D2F2 and 5E4G5 as antibodies against CST4 were verified, excluding cross-reactivity with other CST family members.
To verify that the antibodies (5D2F2 and 5E4G5) recognize the native forms, as detected by ELISA, CST4 recombinant protein was first immunoprecipitated by 5D2F2 and then band-detected by 5E4G5 for Western blotting assay. The Western blotting results (Figure S5) show positive bands for CST4 and for the heavy and light chains of the antibody. This implies that the capture antibody of 5D2F2 is capable of recognizing CST4 in its natural conformation as it exists in extracorporeal blood. To maximize the sensitivity of ELISA detection, citrate-buffered saline (0.05 M, pH 9.6) was selected as the coating buffer. We also found that P/N reached its maximum reading of 24.83 at a coating concentration of 100 ng·mL−1 of enzyme-labeled antibody (N<0.1); at the same time, the sensitivity reached a plateau when the coating antibody concentration approached 2.5 μg·mL−1. Hence, 100 ng·mL−1 of enzyme-labeled antibody and 2.5 μg·mL−1 of coating antibody were selected for our detection system.
Performance analysis of detection system
The overall detection parameters of the CST4-ELISA system are shown in Table 3. The OD values for detection of different concentrations of CST4 (Figure S6A) showed that the linear range was below 2,000 pg·mL−1. There were negligibly small changes at concentrations of 6.25–25 pg·mL−1, which could not be separated from the background (less than M + 2SD; data shown in Table S5). The correlation coefficient was R>0.990, with an efficient concentration range of 50–1,600 pg·mL−1. The standard curve is shown in Figure 2. Thus, the linear range was determined to be 50–1,600 pg·mL−1.
  | Figure 2 CST4 linear regressions of OD and concentration in the linear range. |
We calculated the M + 2SD of blank samples as the LOD. Consistent with the industry standard, the LOD was confirmed as 40 pg·mL−1.
The Hook effect was also investigated; the curve in Figure S6B shows the results below the upper limit of the microplate reader, that is, up to 6,400 pg·mL−1, in which the Hook effect was not observed.
The precision was evaluated using the detection results with CST concentrations of 1,200 and 200 pg·mL−1, for which the in-batch/batch-to-batch and in-day/day-to-day CV were 3.69% and 4.43% (data not shown), respectively, which reach the industry standard.
The accuracy was determined by recovery experiments; 15 μL of 1,200 pg·mL−1 reference A was added to 300 μL of 50 pg·mL−1 reference B. As shown in Table S6, the recovery rates for three experiments using three different batch samples were 98.64%, 103.90%, and 104.33%, respectively. Consistent with the industry standard, the recovery rate was determined in the range of 80%–120%. For interference experiments, CST4 samples with three different concentrations were detected after adding different kinds of interfering substance (detailed types and concentrations of substances are given in Table S2). All the results showed <10% deviation, which meant that the interfering substance did not affect detection.
Determining the normal reference range
We gathered 100 GC and 100 CRC samples, plus 60 gastric and 60 colorectal benign disease samples, 50 samples of other cancers, and 250 healthy samples as control. According to the ROC (Figure 3), the areas under the ROC of GC and CRC were 0.841 (0.797–0.588, 95%) and 0.836 (0.791–0.880, 95%), respectively. At a CST4 concentration of 101 pg·mL−1, the correct diagnosis index, defined as sensitivity – (1 – specificity), reached its maximum. Thus, for GC and CRC, the cut-off values were chosen as 101.07 and 101.59 pg·mL−1, the sensitivities were 69.0% and 69.0%, and the specificities were 85.6% and 83.6%, respectively.
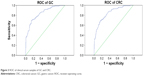  | Figure 3 ROC of clinical serum samples of GC and CRC. |
Clinical verification of antibody-sandwich ELISA analysis system of CST4
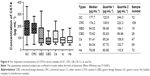
In order to evaluate the clinical detection efficacy of the established antibody-sandwich ELISA analysis system for CST4, another 588 serum samples (Table 1) were used. The concentration ranges of CST4 among all group samples are presented in Figure 4; the median, quartile 1, and quartile 2 values for gastric and colorectal tumors were significantly higher than those of all other groups (P<0.001). This indicates that CST4 is specifically secreted in serum samples of GC and CRC, which are thus highly distinguishable from samples of gastric or colorectal diseases, other tumors, samples containing common interfering blood substance, and healthy tissue, exhibiting specificity to an industry standard.
The accuracy of CST4 detection was analyzed with statistical methods; the results are shown in Table 4. The sensitivity of CST4 detection was 72.3% in GC and 88.4% in CRC. The corresponding specificities for GC and CRC were 80.9% and 81.8%, respectively. The negative control samples included benign disease, other cancers, interference samples, and healthy samples, which represent most kinds of negative samples. The negative coincidence rate (the ratio of detected negative sample numbers divided by total negative sample numbers) of healthy control samples reached 85.7%.
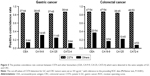
For the sake of comparison with ordinary clinical biomarkers, we collected results for CEA, CA19-9, CA125, and CA72-4, which are commonly detected in hospital tests. The statistical analysis results (Figure 5) demonstrated that the positive detection rate (the ratio of detected positive sample numbers divided by total positive sample numbers) of CST4 (84.09% for GC and 87.03% for CRC) was notably higher than that of the other four common biomarkers (31.1% for GC and 37.07% for CRC at most), showing great superiority in sensitivity (P<0.001).
Discussion
In this work, we have discovered a promising novel human blood-secreted protein, CST4, which is specifically highly expressed in gastrointestinal cancer tissues and cell lines (Figures S1 and S2). Therefore, the idea of establishing an ELISA detection method of blood CST4 from gastrointestinal cancer patients for cancer diagnosis emerged. We generated specific monoclonal antibodies (5D2F2 and 5E4G5) for CST4 and developed an antibody-sandwich ELISA analysis system for CST4. After use on a preliminary training set of 320 total samples and interference samples, this CST4 detection system showed a series of outstanding performance parameters: linear range of 50–1,600 pg·mL−1 and LOD of 40 pg·mL−1, as well as sensitivities of 69.0% and 69.0% and specificities of 85.6% and 83.6%, respectively, for GC and CRC, which meet the industry standards.
To check the specificity and sensitivity of this detection system, we used another 588 of the serum samples (the validation set) to detect CST4 in blood samples. The positive detection rate of CST4 for GC and CRC (84.09% and 87.03%, respectively) indicated significant advantages over other four traditional biomarkers, CEA, CA19-9, CA125, and CA72-4 (31.1% and 37.07% at most), showing great superiority of this CST4 detection system in sensitivity.
Current gastrointestinal clinical markers, including CEA, CA19-9, CA125, and CA72-4, suffer from some deficiencies. CA19-9 has been applied as a typical gastrointestinal marker, yet it shows elevated concentrations in both cancerous and benign lesion samples, indicating that it lacks tissue specificity.17–19 CA72-4 is an epithelioma-sensitive glycoprotein biomarker that can indicate canceration in all digestive system tumors and ovarian cancer with good specificity and sensitivity, but it failed to effectively predict prognosis in advanced gastric carcinoma.20–22 Both CA72-4 and CA19-9 showed poor positive detection rates and sensitivity of only 40%–50% in gastrointestinal carcinomas.23,24 CEA also lacks specificity in GC and CRC, with a positive single detection rate of 48%–60.2%.25,26 CSTs have long been known to balance the enzyme activity of cathepsin to prevent cell matrix hydrolysis; however, in recent years, they have been found to be highly expressed in cancer tissues and to have important roles in cancer growth, angiogenesis, infiltration, metastasis, and so on. Abnormal expression of CSTs, including CST1, CST3, and CST6, has already been shown to be related to cancers.8–11 Keppler proposed that some CSTs share certain characteristics, such as being single-domain, cytoplasmic, and cell-secreted proteins, which may enable them to play an important part in the promotion or suppression of tumor growth, invasion, and metastasis.27 We have provided preliminary verification of the higher sensitivity of CST4 in gastrointestinal carcinoma tissues and cell lines; however, the exact molecular mechanism of CST4 in tumorigenesis remains unrevealed. We will next explore the underlying reasons for the role of CST4 as an excellent blood biomarker.
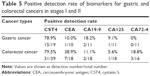
In addition to the above results, the detection differences between CST4 and other biomarkers in the early stages (stage I and II) of GC and CRC are depicted in Table 5. Similar huge advantages in early diagnosis (positive detection rate) of GC and CRC were also observed (78.9% and 79.5% respectively versus 18.2% and 38.9% respectively at most). Moreover, in the training set, high positive detection rates of 60.5% (23/38) and 67.7% (42/62) of CST4 for stage I and II GC and CRC, respectively, were observed (data not shown). This may indicate great potential for clinical applications in early mass screening, which is crucial for effective treatment and survival of patients with GC and CRC. To date, insufficient serum samples (44–54) have been used to determine whether there are significant differences in clinical detection between traditional blood biomarkers (CEA, CA19-9, CA125, and CA72-4) and CST4. Clinical tests for CST4 should be conducted on a larger cohort of clinical samples (along with comparative studies of CEA, CA19-9, and so on) to verify its diagnostic significance in gastrointestinal cancers in the near future.
Conclusion
We have identified the overexpression of CST4 in gastrointestinal cancer tissues and cell lines as a novel tumor biomarker candidate, and developed an antibody-sandwich ELISA analysis system for detection of CST4 in clinical blood samples. This CST4 detection system showed sensitivities of 69.0% and 69.0% and specificities of 85.6% and 83.6% for GC and CRC, respectively, which were significantly higher than those of other common clinical biomarkers. Clinical experiments with a validation set also showed a distinguishable CST4 median concentration and high positive detection rate, further confirming the specificity and sensitivity of this method. The novel blood biomarker CST4 has enormous potential clinical diagnostic value in GC and CRC. In future work, further serum samples from many more sources will be collected to verify the clinical utility of this biomarker.
Acknowledgment
This research was carried out with support from the National “863 program” (number SQ2014SF02D00274) of the People’s Republic of China.
Disclosure
The authors report no conflicts of interest in this work.
References
Ferlay J, Soerjomataram I, Dikshit R, et al. Cancer incidence and mortality worldwide: Sources, methods and major patterns in GLOBOCAN, 2012. Int J Cancer. 2015;136(5):E359–E386. | ||
Sung JJY, Ng EK, Lin JT, et al; Asia Pacific GI Oncology Summit Group. Digestive cancer management in Asia: position statements: a report on GI oncology summit in 2011. J Gastroen Hepatol. 2012;27(9):1417–1422. | ||
Ouyang DL, Chen JJ, Getzenberg RH, Schoen RE. Noninvasive testing for colorectal cancer: a review. Am J Gastroenterol. 2005;100(6):1393–1403. | ||
Imperiale TF, Ransohoff DF, Itzkowitz SH, Turnbull BA, Ross ME; Colorectal Cancer Study Group. Fecal DNA versus fecal occult blood for colorectal cancer screening in an average-risk population. N Engl J Med. 2004;351(26):2704–2714. | ||
Parekh M, Fendrick AM, Ladabaum U. As tests evolve and costs of cancer care rise: reappraising stool-based screening for colorectal neoplasia. Aliment Pharmacol Ther. 2008;27(8):697–712. | ||
Woolf SH. The best screening test for colorectal cancer-a personal choice. N Engl J Med. 2000;343(22):1641–1643. | ||
Winawer S, Fletcher R, Rex D, et al; Gastrointestinal Consortium Panel. Colorectal cancer screening and surveillance: clinical guidelines and rationale – Update based on new evidence. Gastroenterology. 2003;124(2):544–560. | ||
Yoneda K, Iida H, Endo H, et al. Identification of cystatin SN as a novel tumor marker for colorectal cancer. Int J Oncol. 2009;35(1):33–40. | ||
Sotiropoulou G, Anisowicz A, Sager R. Identification, cloning, and characterization of cystatin M, a novel cysteine proteinase inhibitor, down-regulated in breast cancer. J Biol Chem. 1997;272(2):903–910. | ||
Kos J, Krasovec M, Cimerman N, Nielsen HJ, Christensen IJ, Brünner N. Cysteine proteinase inhibitors stefin A, stefin B, and cystatin C in sera from patients with colorectal cancer: relation to prognosis. Clin Cancer Res. 2000;6(2):505–511. | ||
Choi EH, Kim JT, Kim JH, et al. Upregulation of the cysteine protease inhibitor, cystatin SN, contributes to cell proliferation and cathepsin inhibition in gastric cancer. Clin Chim Acta. 2009;406(1–2):45–51. | ||
Bobek LA, Aguirre A, Levine MJ. Human salivary cystatin S. Cloning, sequence analysis, hybridization in situ and immune cytochemistry. Biochem J. 1991;278(3):627–635. | ||
Isemura S, Saitoh E, Ito S, Isemura M, Sanada K. Cystatin S: a cysteine proteinase inhibitor of human saliva. J Biochem. 1984;96(4):1311–1314. | ||
Abe K, Emori Y, Kondo H, Suzuki K, Arai S. Molecular cloning of a cysteine proteinase inhibitor of rice (oryzacystatin). Homology with animal cystatins and transient expression in the ripening process of rice seeds. J Biol Chem. 1987;262(35):16793–16797. | ||
Akiba S, Hayashi Y, Hakamada Y, et al. Extracellular production of human cystatin S and cystatin SA by Bacillus subtilis. Protein Expres Purif. 2006;49(2):203–210. | ||
Friguet B, Djavadi-Ohaniance L, Pages J, Bussard A, Goldberg M. A convenient enzyme-linked immunosorbent assay for testing whether monoclonal antibodies recognize the same antigenic site. Application to hybridomas specific for the beta 2-subunit of Escherichia coli tryptophan synthase. J Immunol Methods. 1983;60(3):351–358. | ||
Ferrone CR, Finkelstein DM, Thayer SP, Muzikansky A, Fernandez-delCastillo C, Warshaw AL. Perioperative CA19-9 levels can predict stage and survival in patients with resectable pancreatic adenocarcinoma. J Clin Oncol. 2006;24(18):2897–2902. | ||
Edwards R, Ho S, Lau WY, et al. Elevated tumor marker CA19-9: clinical interpretation and influence of obstructive jaundice. Eur J Surg Oncol. 2000;26(5):474–479. | ||
Chen CC, Yang SH, Lin JK, et al. Is it reasonable to add preoperative serum level of CEA and CA19-9 to staging for colorectal cancer? J Surg Res. 2005;124(2):169–174. | ||
Tocchi A, Costa G, Lepre L, et al. The role of serum and gastric juice levels of carcinoembryonic antigen, CA19.9 and CA72.4 in patients with gastric cancer. J Cancer Res Clin Oncol. 1998;124(8):450–455. | ||
Emoto S, Ishigami H, Yamashita H, Yamaguchi H, Kaisaki S, Kitayama J. Clinical significance of CA125 and CA72-4 in gastric cancer with peritoneal dissemination. Gastric Cancer. 2012;15(2):154–161. | ||
Holubec L Jr, Topolcan O, Pikner R, et al. The significance of CEA, CA19-9 and CA72-4 in the detection of colorectal carcinoma recurrence. Anticancer Res. 2000;20(6D):5237–5244. | ||
Lai IR, Lee WJ, Huang MT, Lin HH. Comparison of serum CA72-4, CEA, TPA, CA19-9 and CA125 levels in gastric cancer patients and correlation with recurrence. Hepatogastroenterology. 2002;49(46):1157–1160. | ||
Ychou M, Duffour J, Kramar A, Gourgou S, Grenier J. Clinical significance and prognostic value of CA72-4 compared with CEA and CA19-9 in patients with gastric cancer. Dis Markers. 2000;16(3–4):105–110. | ||
Youssef EMI. Comparison between CEA, CA 19-9 and CA 72-4 in patients with colon cancer. Crop Sci. 1977;17(1):897–901. | ||
Huang P, Donghai. Clinical significance of determination of changes of serum CEA, CA242, CA72-4 and CA19-9 levels after operation in patients with gastric cancer. J Radioimmunol. 2009;22(6):598–599. | ||
Keppler D. Towards novel anti-cancer strategies based on cystatin function. Cancer Lett. 2006;235(2):159–176. |
Supplementary materials
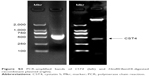  | Figure S3 PCR-amplified bands of CST4 (left) and HindIII/BamHI-digested recombinant plasmid (right). |
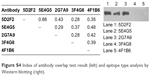  | Figure S4 Index of antibody overlap test result (left) and epitope type analysis by Western blotting (right). |
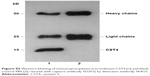  | Figure S5 Western blotting of immunoprecipitates (recombinant CST4 [1] and blank control PBS [2] reacted with capture antibody 5D2F2) by detection antibody 5E4G5. |
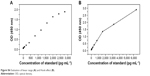  | Figure S6 Evaluation of linear range (A) and Hook effect (B). |
  | Table S2 Types and critical values of interference samples |
  | Table S3 Sequencing result of recombinant plasmid of CST4-pcDNA3.1 |
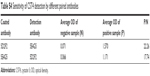  | Table S4 Sensitivity of CST4 detection by different paired antibodies |
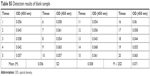  | Table S5 Detection results of blank sample |
 © 2018 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.
© 2018 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.