Back to Journals » International Journal of Nanomedicine » Volume 16
An Ultrasensitive Label-Free Fluorescent Aptasensor Platform for Detection of Sulfamethazine
Authors Wang Y, Yan X, Kou Q, Sun Q, Wang Y, Wu P, Yang L, Tang J, Le T
Received 17 February 2021
Accepted for publication 17 March 2021
Published 9 April 2021 Volume 2021:16 Pages 2751—2759
DOI https://doi.org/10.2147/IJN.S307080
Checked for plagiarism Yes
Review by Single anonymous peer review
Peer reviewer comments 2
Editor who approved publication: Dr Yan Shen
Yarong Wang, Xueling Yan, Qiming Kou, Qi Sun, Yuexin Wang, Ping Wu, Lulan Yang, Jiaming Tang, Tao Le
College of Life Science, Chongqing Normal University, Chongqing, 401331, People’s Republic of China
Correspondence: Tao Le Tel +86 23 65910119
Fax +86 23 65910119
Email [email protected]
Purpose: Sulfamethazine (SMZ) exposed in the environment can enter the human body through the food chain and pose a serious threat to human health. Therefore, it is important to develop a rapid and sensitive method for detecting SMZ in environmental samples. In order to fastly and quantitatively detect SMZ in environmental samples, we developed a label-free fluorescent aptasensor based on specific aptamer (SMZ1S) and fluorescence resonance energy transfer (FRET) between gold nanoparticles (AuNPs) and rhodamine B (RhoB).
Methods: In the absence of SMZ, SMZ1S was adsorbed on the surface of AuNPs, which led to dispersion of the AuNPs in high concentration saline solution, thus effectively quenching the fluorescence of RhoB. With the increase of the SMZ concentration, the specific binding of SMZ1S and SMZ led to the aggregation of AuNPs in the presence of NaCl, which reduced the quenching of RhoB fluorescence and increased the fluorescence intensity. The sensitivity and linearity curve of the label-free fluorescent aptasensor were determined with different concentrations of sulfamethazine standard solutions. The specificity of this fluorescent aptasensor was determined by replacing sulfamethazine with different antibiotics. In addition, the actual water and soil samples were spiked and recovered.
Results: Under optimized conditions, the proposed fluorescent aptasensor demonstrated a good linear detection of SMZ in binding buffer from 1.25 ng mL− 1 to 40 ng mL− 1 and the limit of detection was 0.82 ng mL− 1. The spiked recoveries for SMZ were 94.4% to 108.8% with a relative standard deviation of 1.8– 10.3% in water and soil samples, respectively.
Conclusion: The label-free fluorescent aptasensor investigated in the current study is a promising tool to detect and quantify SMZ in water and soil samples.
Keywords: label-free, fluorescence aptasensor, sulfamethazine, gold nanoparticles, rhodamine B
Introduction
Sulfamethazine (SMZ) is a sulfonamide antibiotic that is frequently used against bacterial and parasitic infections.1 However, excessive and long-term use of the SMZ has led to its accumulation in farmed animals and animal-derived foods.2 Furthermore, SMZ and its metabolites are known to enter water and soil samples through animal urine and feces, thus and posing a significant threat to the environment.3,4 For instance, SMZ was detected in water samples collected from 60 agricultural regions in northern Spain.5 Since environmental SMZ can enter the food chain through aquatic organisms and crops, and it is known that the SMZ can cause severe allergic reactions and thyroid tumors in humans,6 it is essential to develop a rapid and sensitive method to detect SMZ in environmental samples.
To data, SMZ is detected by gas chromatography-mass spectrometry,7 high-performance liquid chromatography (HPLC),8,9 capillary electrophoresis,10 and high-performance liquid chromatography-tandem mass spectrometry (HPLC-MS/MS).11,12 Although these methods are standardized, accurate and reliable, their wide applications are limited by high costs and complex sample preparations.13 In addition, antibody-dependent methods including enzyme-linked immunosorbent assay (ELISA),14 chemiluminescence,15 time-resolved immunoassay,16,17 lateral flow immunoassay18 and electrochemical immunosensor19 also face different challenges such as poor stabilities, high costs, and difficulties in modifying antibody structures.20
Aptamers are single-stranded DNA or RNA that bind specifically to target molecules, and they are selected by an in vitro process called systematic evolution of ligands by exponential enrichment (SELEX). In comparison with antibodies, aptamers are characterized by high binding selectivities, low cost to prepare and facile chemical modification.21 The first SMZ-specific aptamer was selected in 2019 by Yang et al22 and was used to build chemiluminescent and graphene oxide (GO) quantum dots-based fluorescent aptasensors to detect SMZ in milk samples. Recently, we used a non-fixed ssDNA library and improved GO-SELEX technology to obtain a highly specific SMZ-binding aptamer (SMZ1S) after seven rounds of screening,23 and developed a GO-based fluorescent aptasensor to detect SMZ. However, these methods required labeling aptamer, which is time-consuming and technically complex, and may even reduce the efficiency of target binding. Therefore, a rapid and simple label-free method is needed for detecting SMZ.
Due to its simple preparation, high extinction coefficient, high oxidation resistance, and large specific surface area, gold nanoparticles (AuNPs) are an ideal nanomaterial used in optical sensors to improve the sensitivity of the system.24 Many methods for the detection of small organic molecules, metal ions and proteins based on AuNPs have been reported, such as fluorescence25 and colorimetry.26 In addition, since the obvious overlap between the absorption spectrum of AuNPs and the emission spectra of dyes such as rhodamine B (RhoB) and rhodamine 6G, AuNPs have a significant effect on the photoluminescence of these dyes and are used in some fluorescence aptasensors as “super quencher”.27 RhoB is a synthetic dye with strong fluorescence, light stability and water solubility.28 Study has shown that when RhoB is absorbed on the surface of AuNPs through electrostatic interactions, dispersed AuNPs can greatly quench the fluorescence of RhoB, while aggregated AuNPs exhibit weak quenching ability.29
As a series of oligonucleotides, aptamers can be attached to the surface of AuNPs through the van der Waals forces and the coordination of the N atoms in the DNA component and the gold ions of the AuNPs, and prevent the salt-induced aggregation of the AuNPs.30 Li et al have demonstrated that when the aptamer is adsorbed on the surface of AuNPs, the specific binding of the aptamer to its target can cause the aggregation of gold nanoparticles AuNPs.31 Therefore, when using AuNPs combined with aptamers for target detection, the amount of target added can be used to control the aggregation of gold nanoparticles.
In this research, we developed an aptamer-based fluorescent sensing platform based on the specific binding of SMZ1S to SMZ, and the fluorescence resonance energy transfer (FRET) between RhoB and AuNPs to detect SMZ in water and soil samples. Neither SMZ1S nor the AuNPs require any labeling and modification in this aptasensor system, which can potentially increase the detection efficiency of SMZ.
Materials and Methods
Chemicals and Reagents
Sulfamethazine-specific aptamer, SMZ1S (5′-CGTTAGACG-3′), was screened in our laboratory.23 HAuCl4·4H2O, SMZ, sulfadimethoxine, sulfameter and nitrofurantoin were obtained from Sigma-Aldrich (St. Louis, MO, USA). Sulfapyridine, ofloxacin, chlortetracycline, doxycycline and chloramphenicol were purchased from Aladdin Biotechnology Inc. (Shanghai, China). Rhodamine B was obtained from Sangon Biotechnology Inc (Shanghai, China). Unless otherwise mentioned, all other reagents were of analytical grade. Ultrapure water was used for all experiments and obtained using a Millipore Milli-Q system (Millipore, Bedford, MA, USA).
Synthesis and Characterization of AuNPs
Citrate-coated AuNPs were prepared using a previously described protocol with minor modifications.32 Briefly, 0.5 mL 1% HAuCl4·4H2O was added to 50 mL dd∙H2O and heated to boil with vigorous stirring, followed by the addition of 1 mL 1% trisodium citrate. The solution was stirred for 5 minutes till it attained a wine-red color, after which it was cooled to room temperature, and subsequently stored in a dark glass bottle at 4°C. The resulting AuNPs were characterized by measuring the absorption at 520 nm using Varioskan™ LUX Multimode Microplate Reader (Thermo Fisher Scientific Inc., Waltham, MA, USA). Dynamic light scattering (DLS) (NanoBrook Omni, Holtsville, NY, USA) was used to measure the average diameter of the AuNPs.
Interactions Between Aptasensor Components
The AuNPs (180 µL) were incubated with 80 nM SMZ1S, 30 mM NaCl, 2.5 µM RhoB and 1µg mL−1 SMZ in various combinations (total reaction volume 200 µL) at 25°C. The absorbance of AuNPs, AuNPs-NaCl, SMZ1S-AuNPs-NaCl, SMZ-AuNPs and SMZ-SMZ1S- AuNPs-NaCl was measured at 695 nm (A695) and 520 nm (A520) using Varioskan™ LUX Multimode Microplate Reader (Thermo Fisher Scientific Inc., Waltham, MA, USA), respectively, to determine the relative extent of aggregated and dispersed AuNPs. The degree of aggregation was calculated as the A695/A520 ratio. Fluorescence intensity of RhoB measured under the excitation wavelength of 510 nm using Varioskan™ LUX Multimode Microplate Reader (Thermo Fisher Scientific Inc., Waltham, MA, USA). The concentration of SMZ was estimated on the basis of ΔF, which was calculated as F–F0 wherein F and F0 are, respectively, the fluorescence intensities in the presence and in the absence of SMZ, respectively. The fluorescence spectra of RhoB, AuNPs-RhoB, SMZ1S-AuNPs-NaCl-RhoB and SMZ-SMZ1S-AuNPs-NaCl-RhoB were compared. In addition, fluorescence spectra of RhoB, SMZ1S-RhoB, SMZ-RhoB, NaCl-RhoB, SMZ1S-NaCl-RhoB, SMZ1S-SMZ-RhoB and SMZ1S-SMZ-NaCl-RhoB were measured in the absence of AuNPs to assess the effect of the AuNPs on RhoB. Similarly, the effect of NaCl on the interaction between AuNPs and RhoB was determined by comparing AuNPs-RhoB, SMZ-AuNPs-RhoB, SMZ1S-AuNPs-RhoB and SMZ-SMZ1S-AuNPs-RhoB.
Optimization of Detection Conditions
Different concentrations of NaCl (0, 10, 20, 30, 40 and 50 mM) were incubated with the AuNPs for 10 min, the absorbance of each of the samples at 695 nm and 520 nm was measured, and the ratios of the absorbance A695/A520 for each sample were calculated. In addition, the AuNPs were incubated with the optimal concentration of NaCl that resulted in maximum A695/A520 for 0, 2, 4, 6, 8, 10 and 12 min, and A695/A520 was calculated. Different concentrations of SMZ1S (0, 20, 40, 60, 80, 100 and 120 nM) were incubated with the AuNPs for 30 min, followed by the optimized NaCl concentration for 10 min. The A695/A520 was calculated, and the SMZ1S concentration corresponding to minimum A695/A520 was incubated with the AuNPs for 0, 10, 20, 30 and 40 min. After incubating with the optimized concentration of NaCl for 10 min, and the A695 and A520 were measured. The incubation time resulting in the lowest A695/A520 value was considered the optimal incubation time.
The optimized doses of SMZ1S and SMZ were incubated for 0, 5, 10, 15, 20, 25 and 30 min, followed by the AuNPs for 30 min and then NaCl for 10 min. The A695 and A520 of the sample solution were measured, and the incubation time of with the highest A695/A520 was considered the optimal binding duration of SMZ and SMZ1S. To optimize the concentration of RhoB, SMZ1S was incubated with/out SMZ for 20 min, and then sequentially with the AuNPs and NaCl as described above. Different concentrations of RhoB (0, 0.5, 1, 1.5, 2, 2.5 and 3 µM) were then added, and the fluorescence intensities of the SMZ (F) and blank (F0) samples were measured. The ΔF was calculated, and the optimal concentration of RhoB was determined based on the highest value of ΔF.
Analytical Performance of the Fluorescent Aptasensor
To study the sensitivity of the fluorescent aptasensor, SMZ of different concentrations (0, 1.25, 5, 10, 20, 40, 80, 160, 320, 500, 750, 1000 ng mL–1) was incubated with SMZ1S for 20 min, after which the AuNPs was added and incubated for 30 min. Subsequently, NaCl was added and incubated for 10 min. Finally, RhoB was added into the mixture to incubate for another 10 min. The fluorescence intensity (F, F0) was measured and the value of ΔF (ΔF=F-F0) was calculated. To study the specificity, different types of interfering substances were selected to replace SMZ, including sulfonamides: sulfapyridine, sulfadimethoxine, sulfameter; and other types of antibiotics: ofloxacin, chlortetracycline, doxycycline, chloramphenicol, and nitrofurantoin. The final concentration of all interfering substances was 1 µg mL−1. The fluorescence intensity F and F0 were measured, and the value ΔF (ΔF=F-F0) was calculated.
Application to Real Samples
To evaluate the practical feasibility and accuracy of the aptasensor, tap water, lake water and soil samples collected from farms in Chongqing, China were spiked with SMZ to the final concentrations of 5, 10, 15 and 20 ng mL−1. The samples were validated to be SMZ-free by HPLC. The water samples were directly spiked, whereas the soil samples (1g) were vortexed for 1 min with 1 mL binding buffer, and the extracts obtained after 10 min incubation at room temperature were spiked. All samples were centrifuged at 10,000 rpm for 10 minutes and the supernatant was filtered through a 0.22 μm syringe filter (Thermo Fisher Scientific Inc., Wilmington Waltham, MA, USA) before aptasensor analysis. The analysis for each concentration level was repeated in five copies over a period of two months, then the recoveries and coefficients of variation (CVs) of the samples were calculated.
Results and Discussions
Principle of the Fluorescent Aptasensor for SMZ Detection
As shown in Figure 1, the fluorescent aptasensor consists of SMZ, SMZ1S, AuNPs, NaCl and RhoB. In the absence of SMZ, SMZ1S can attach to the surfaces of AuNPs via the interactions between bases and gold ions. The resulting SMZ1S-AuNPs complex allows the AuNPs to remain in a dispersed state even at high NaCl concentrations. As a result, the dispersed AuNPs can effectively quench the initial high fluorescence of RhoB.33 However, in the presence of SMZ, SMZ1S specifically binds with SMZ to form the SMZ1S-SMZ complex, which is more stable compared to SMZ1S-AuNPs. Therefore, the negative charges on the surface of AuNPs interact with the positive charge of NaCl and form aggregates that cannot quench RhoB fluorescence.34,35 The effect of AuNPs aggregation and dispersion on the fluorescence intensity of RhoB can theoretically allow SMZ quantification in water and soil samples.
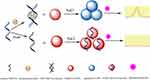 |
Figure 1 Schematic illustration of the fluorescent aptasensor for detecting SMZ based on label-free aptamer and FRET between RhoB and AuNPs. |
Interactions Between Aptasensor Components
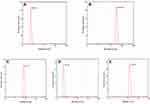
The effect of different sample solutions on SMZ1S-mediated SMZ detection was evaluated in terms of the changes in the absorption spectra and diameter of AuNPs. As shown in Figure 2, the citrate-coated AuNPs revealed an absorption peak at 520 nm, and addition of NaCl significantly reduced the intensity at 520 nm and resulted in a new peak at 695 nm. Since SMZ1 can bind to and stabilize the AuNPs in the absence of SMZ,28,36 NaCl had no effect on the absorption spectrum of AuNPs in the presence of SMZ1. In addition, SMZ did not alter the absorption spectra of AuNPs in the absence of SMZ1, whereas specific binding between SMZ and SMZ1S resulted in agglomeration of the AuNPs upon addition of NaCl, which significantly reduced absorption at 520 nm. The diameter of citrate-coated AuNPs was approximately 0.54 nm (Figure 3A) and increased significantly following NaCl-mediated aggregation (Figure 3B). Adsorption of SMZ1S on the surface of AuNPs also increased their diameter to 6.21 nm (Figure 3C). SMZ did not affect the diameter of AuNPs (Figure 3D) in the absence of SMZ1S, while the presence of both led to the aggregation of AuNPs in saline conditions and increased their diameter (Figure 3E).
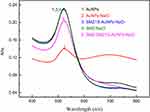 |
Figure 2 Absorption spectra of AuNPs solutions in different sample solutions. AuNPs, 180 µL; NaCl, 30 mM; SMZ1S, 80 nM; SMZ, 1µg mL−1. |
As shown in Figure 4, the strong fluorescence intensity of RhoB (Curve 1) was significantly decreased by the AuNPs (Curve 2) even in the presence of SMZ1S (Curve 3). On the other hand, the binding of SMZ1S to SMZ led to AuNPs aggregation in saline conditions and restored RhoB fluorescence (Curve 4). In the absence of AuNPs, the fluorescence intensity of RhoB was not affected by either NaCl, SMZ or SMZ1S, or by the specific binging of SMZ and SMZ1S (Figure S1A). Likewise, SMZ and SMZ1S interaction did not reverse AuNPs-mediated quenching of RhoB fluorescence in the absence of NaCl (Figure S1B).
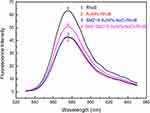 |
Figure 4 Fluorescence spectra of RhoB in different sample solutions. AuNPs, 180 µL; NaCl, 30 mM; SMZ1S, 80 nM; RhoB, 2.5 µM; SMZ, 1µg mL−1. |
Optimization of Aptasensor Performance
The concentrations of NaCl, SMZ1S and RhoB, as well as the incubation time, were optimized to improve the sensitivity of the SMZ1S aptamer to SMZ. As shown in Figures S2 and S3, the A695/A520 was maximum with 30 mM NaCl and incubation time ≥ 10 min, whereas incubation with 80 nM SMZ1S for 30 min or more resulted in the lowest A695/A520 (Figures S4 and S5). In addition, the A695/A520 ratio peaked following incubation of SMZ and SMZ1S for 20 min (Figure S6). Finally, the ΔF value was maximum when the concentration of RhoB was 2.5 µM and decreased thereafter due to the background fluorescence (Figure S7).
Sensitivity of SMZ Detection
As shown in Figure 5, ΔF increased linearly in the SMZ concentration range of 1.25–40 ng mL−1, and the linear regression equation was ΔF=0.1103 CSMZ+0.2511 (R2=0.99). The limit of detection (LOD) was calculated as 0.82 ng mL−1 using the formula 3*SD/S, where SD was the standard deviation of the instrument and S was the slope of the linear curve. Compared to the conventional detection methods reported previously,22 the aptasensor exhibited good sensitivity for detecting SMZ.
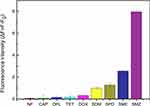
Specificity of SMZ Detection
The selectivity of the aptasensor was determined by testing it against different non-target antibiotics. As shown in Figure 6, the ΔF was significantly lower for nitrofurantoin and other antibiotics compared to that for three sulfonamides with similar structure and functional groups as SMZ. Nevertheless, ΔF was markedly higher for SMZ relative to the analogous sulfonamides, indicating that the SMZ1S aptasensor can detect SMZ residues in real samples with high specificity and selectivity.
Analytical Application in Real Samples
The potential practical application of the SMZ1S aptasensor was further evaluated using water and soil samples spiked with different concentrations of SMZ (5, 10, 15, and 20 ng mL−1). As shown in Table 1, the recoveries ranged from 94.4% to 108.8%, and the CVs ranged from 1.8% to 10.3%. These results indicate that the aptasensor has good reproducibility and can detect SMZ in environmental samples with reliable accuracy.
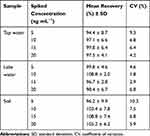 |
Table 1 Mean Recoveries and Coefficients of Variation for the SMZ in Actual Water, Soil Sample Using Optimized Fluorescence Aptasensor (n = 5) |
Conclusion
A label-free fluorescent aptasensor platform for detecting SMZ based on aptamer and FRET between AuNPs and RhoB has been successfully developed. SMZ1S prevented AuNPs aggregation in saline conditions and led to RhoB fluorescence quenching, and the binding of SMZ and SMZ1S led to the aggregation of AuNPs and restored RhoB fluorescence. The LOD of the established aptasensor was 0.82 ng mL−1 with the linear range from 1.25 to 40 ng mL−1. In addition, the recovery was from 94.4% to 108.8%, and the CVs from 1.8% to 10.3%. This novel aptasensor can rapidly detect SMZ in soil and water samples with high sensitivity, reproducibility and accuracy.
Ethical Approval
This article does not contain any studies with human subjects and animal experiments.
Acknowledgments
The authors would like to think the National Natural Science Foundation of China (Grant No. 31671939), the Science and Technology Research Program of Chongqing Municipal Education Commission (KJZD-K201800501), and Chongqing University Innovation Research Group Project (No. CXQT20031) for funding the research.
Disclosure
The authors declare no conflicts of interest.
References
1. Ko E, Song H, Park JH. Direct competitive enzyme-linked immunosorbent assay for sulfamethazine. J Vet Med Sci. 2000;62(10):1121–1123. doi:10.1292/jvms.62.1121
2. Shim WB, Kim JS, Kim MG, et al. Rapid and sensitive immunochromatographic strip for on‐site detection of sulfamethazine in meats and eggs. J Food Sci. 2013;78(10):M1575–M1581. doi:10.1111/1750-3841.12232
3. Grimmett ME. Adsorption of sulfamethazine from environmentally relevant aqueous matrices onto hypercrosslinked adsorbent MN250. J Environ Qual. 2015;44(4):1183–1192. doi:10.2134/jeq2015.02.0109
4. Ghirardini A, Grillini V, Verlicchi P. A review of the occurrence of selected micropollutants and microorganisms in different raw and treated manure–environmental risk due to antibiotics after application to soil. Sci Total Environ. 2020;707:136118. doi:10.1016/j.scitotenv.2019.136118
5. Iglesias A, Nebot C, Vázquez BI, et al. Detection of veterinary drug residues in surface waters collected nearby farming areas in Galicia, North of Spain. Environ Sci Pollut Res. 2014;21(3):2367–2377. doi:10.1007/s11356-013-2142-7
6. Dolliver H, Kumar K, Gupta S. Sulfamethazine uptake by plants from manure‐amended soil. J Environ Qual. 2007;36(4):1224–1230. doi:10.2134/jeq2006.0266
7. Reeves VB. Confirmation of multiple sulfonamide residues in bovine milk by gas chromatography–positive chemical ionization mass spectrometry. J Chromatogr B Biomed Sci Appl. 1999;723(1–2):127–137. doi:10.1016/S0378-4347(98)00548-9
8. Furusawa N. A clean and rapid liquid chromatographic technique for sulfamethazine monitoring in pork tissues without using organic solvents. J Chromatogr Sci. 2003;41(7):377–380. doi:10.1093/chromsci/41.7.377
9. Wen Y, Zhang M, Zhao Q, et al. Monitoring of five sulfonamide antibacterial residues in milk by in-tube solid-phase microextraction coupled to high-performance liquid chromatography. J Agric Food Chem. 2005;53(22):8468–8473. doi:10.1021/jf051319b
10. Tubaon RM, Haddad PR, Quirino JP. High-sensitivity analysis of anionic sulfonamides by capillary electrophoresis using a synergistic stacking approach. J Chromatogr A. 2014;1349:129–134. doi:10.1016/j.chroma.2014.05.007
11. Guofang P, Yanzhong C, Jinjie Z, et al. Simultaneous determination of 16 sulfonamides in poultry meat by high-performance liquid chromatography-tandem mass spectrometry. Chin J Anal Chem. 2005;33(9):1252. doi:10.1038/sj.cr.7290370
12. Wang Y, Liu L, Xiao C, et al. Rapid determination of trace sulfonamides in milk by graphene oxide-based magnetic solid phase extraction coupled with HPLC–MS/MS. Food Anal Method. 2016;9(9):2521–2530. doi:10.1007/s12161-016-0433-6
13. Ha NR, Jung IP, La IJ, et al. Ultra-sensitive detection of kanamycin for food safety using a reduced graphene oxide-based fluorescent aptasensor. Sci Rep. 2017;7(1):1–10. doi:10.1038/srep40305
14. Yang T, Ren X, Li Y, et al. Development of a sensitive monoclonal antibody-based ELISA for the detection of sulfamethazine in cow milk, honey, and swine urine. Hybridoma. 2010;29(5):403–407. doi:10.1089/hyb.2010.0048
15. Ma M, Wen K, Beier RC, et al. Chemiluminescence resonance energy transfer competitive immunoassay employing hapten-functionalized quantum dots for the detection of sulfamethazine. ACS Appl Mater Interfaces. 2016;8(28):17745–17750. doi:10.1021/acsami.6b04171
16. Le T, Yan P, Liu J, et al. Simultaneous detection of sulfamethazine and sulfaquinoxaline using a dual-label time-resolved fluorescence immunoassay. Food Addit Contam A. 2013;30(7):1264–1269. doi:10.1080/19440049.2013.801084
17. Hu S, Li D, Huang Z, et al. Ultra-sensitive method based on time-resolved fluorescence immunoassay for detection of sulfamethazine in raw milk. Food Agric Immunol. 2018;29(1):1137–1149. doi:10.1080/09540105.2018.1520816
18. Wang Z, Hu S, Zhang G, et al. Aggregation-induced emission-based competitive lateral flow immunoassay for rapid detection of sulfamethazine in honey. Food Agric Immunol. 2019;30(1):1303–1317. doi:10.1080/09540105.2019.1689929
19. Yang M, Wu X, Hu X, et al. Electrochemical immunosensor based on Ag+-dependent CTAB-AuNPs for ultrasensitive detection of sulfamethazine. Biosens Bioelectron. 2019;144:111643. doi:10.1016/j.bios.2019.111643
20. Youn H, Lee K, Her J, et al. Aptasensor for multiplex detection of antibiotics based on FRET strategy combined with aptamer/graphene oxide complex. Sci Rep. 2019;9(1):1–9. doi:10.1038/s41598-019-44051-3
21. Song KM, Lee S, Ban C. Aptamers and their biological applications. Sensors. 2012;12(1):612–631. doi:10.3390/s120100612
22. Yang L, Ni H, Li C, et al. Development of a highly specific chemiluminescence aptasensor for sulfamethazine detection in milk based on in vitro selected aptamers. Sens Actuators B Chem. 2019;281:801–811. doi:10.1016/j.snb.2018.10.143
23. Kou Q, Wu P, Sun Q, et al. Selection and truncation of aptamers for ultrasensitive detection of sulfamethazine using a fluorescent biosensor based on graphene oxide. Anal Bioanal Chem. 2021;413(3):901–909. doi:10.1007/s00216-020-03044-2
24. Sang F, Liu J, Zhang X, et al. An aptamer-based colorimetric Pt (II) assay based on the use of gold nanoparticles and a cationic polymer. Microchim Acta. 2018;185(5):1–8. doi:10.1007/s00604-018-2794-6
25. Yun W, Wu H, Chen L, et al. Dual enzyme-free amplification strategy for ultra-sensitive fluorescent detection of bisphenol A in water. Anal Chim Acta. 2018;1020:104–109. doi:10.1016/j.aca.2018.02.064
26. Ramezani M, Danesh NM, Lavaee P, et al. A novel colorimetric triple-helix molecular switch aptasensor for ultrasensitive detection of tetracycline. Biosens Bioelectron. 2015;70:181–187. doi:10.1016/j.bios.2015.03.040
27. Zhan S, Xu H, Zhan X, et al. Determination of silver (I) ion based on the aggregation of gold nanoparticles caused by silver-specific DNA, and its effect on the fluorescence of Rhodamine B. Microchim Acta. 2015;82(7–8):1411–1419. doi:10.1007/s00604-015-1462-3
28. Liu D, Chen W, Wei J, et al. A highly sensitive, dual-readout assay based on gold nanoparticles for organophosphorus and carbamate pesticides. Anal Chem. 2012;84(9):4185–4191. doi:10.1021/ac300545p
29. Zhang H, Wang L, Jiang W. Label free DNA detection based on gold nanoparticles quenching fluorescence of Rhodamine B. Talanta. 2011;85(1):725–729. doi:10.1016/j.talanta.2011.04.057
30. Shahdordizadeh M, Yazdian-Robati R, Ansari N, et al. An aptamer-based colorimetric lead(II) assay based on the use of gold nanoparticles modified with dsDNA and exonuclease I. Microchim Acta. 2018;185(2):1–6. doi:10.1007/s00604-018-2699-4
31. Li H, Rothberg L. Colorimetric detection of DNA sequences based on electrostatic interactions with unmodified gold nanoparticles. Proc Natl Acad Sci U S A. 2004;101(39):14036–14039. doi:10.1073/pnas.0406115101
32. Grabar KC, Freeman RG, Hommer MB, et al. Preparation and characterization of Au colloid monolayers. Anal Chem. 1995;67(4):735–743. doi:10.1021/ac00100a008
33. Zhao W, Brook MA, Li Y. Design of gold nanoparticle‐based colorimetric biosensing assays. ChemBioChem. 2008;9(15):2363–2371. doi:10.1002/cbic.200800282
34. Ni X, Xia B, Wang L, et al. Fluorescent aptasensor for 17β-estradiol determination based on gold nanoparticles quenching the fluorescence of Rhodamine B. Anal Biochem. 2017;523:17–23. doi:10.1016/j.ab.2017.01.021
35. Su L, Wang S, Wang L, et al. Fluorescent aptasensor for carbendazim detection in aqueous samples based on gold nanoparticles quenching Rhodamine B. Spectrochim Acta A. 2020;225:117511. doi:10.1016/j.saa.2019.117511
36. Li H, Rothberg LJ. DNA sequence detection using selective fluorescence quenching of tagged oligonucleotide probes by gold nanoparticles. Anal Chem. 2004;76(18):5414–5417. doi:10.1021/ac049173n
 © 2021 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.
© 2021 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.