Back to Journals » Drug Design, Development and Therapy » Volume 13
Alteration of Amino Acid Profiling Influenced by the Active Ingredients of DanHong Injection After Prescription Optimization
Authors Guo Z , Zhu Y, Xu W, Luo K, Xiao H, Wang Z
Received 21 June 2019
Accepted for publication 12 November 2019
Published 21 November 2019 Volume 2019:13 Pages 3939—3947
DOI https://doi.org/10.2147/DDDT.S220314
Checked for plagiarism Yes
Review by Single anonymous peer review
Peer reviewer comments 2
Editor who approved publication: Dr Qiongyu Guo
Zhili Guo,1 Yan Zhu,2 Wenjuan Xu,3 Kaitao Luo,1 Hongbin Xiao,4 Zhong Wang3
1Chinese Medicine Department, Jiaxing Hospital of Traditional Chinese Medicine, Jiaxing University, Jiaxing, Zhejiang, 314000, People’s Republic of China; 2Chinese Medicine Department, Beijing Electric Power Hospital, Capital Medical University, Beijing 10073, People’s Republic of China; 3Chinese Medicine Pharmacology, Institute of Basic Research in Clinical Medicine. China Academy of Chinese Medical Sciences, Beijing 100700, People’s Republic of China; 4Beijing University of Chinese Medicine, Chinese Medicine Institute, Beijing 100029, People’s Republic of China
Correspondence: Kaitao Luo; Zhong Wang Tel +86057382079269; +861064014411-3308
Email [email protected]; [email protected]
Introduction: The aim of this work was to optimize the formulation composition of DanHong injection and to study the disturbance of microscopic components of cerebral ischemia in amino acid metabolites and metabolic pathways. The subtle relationship among these three substances and the influence of metabolic pathways were also studied.
Methods: In this study, the central composite design (CCD) matrix and response surface methodology (RSM) were used to design the experiments and to evaluate the interactive effects of three substances. Targeted metabolomics was used to detect the amino acid variation in CCD sets.
Results: Response surfaces were generated, and the formulation was optimized by superimposing the contour plots. It was found that the optimum values of the responses could be obtained at an SAB concentration (x1) of 8–9 mg/kg, a TSN concentration (x2) of 14–16 mg/kg, and an HSYA yellow A concentration (x3) of 6 mg/kg. Statistical analysis showed that the three independent variables had significant effects (p < 0.05) on the responses. A total of 22 experimental runs were performed, and the kinetic data were analyzed using a second-order polynomial. Model algorithm calculation indicated that glutamic acid, serine, leucine, glycine, and valine had a very close correlation with the active ingredients. Methionine, aspartic acid, asparagine, glutamic acid, and valine were important for distinguishing different groups, and they were identified as potential biomarkers. Cluster analysis and pathway analysis indicated that the valine, leucine, and isoleucine degradation (VLI degradation) pathway was the major metabolic pathway. Arginine and proline metabolites were most frequently detected, and they were closely associated with other networks according to the network analysis results. VLI degradation pathway and arginine and proline metabolism pathway had a significant influence on cerebral ischemia.
Discussion: The integration of CCD and metabolomics may be an effective strategy for optimizing the formulation composition and identifying the mechanism of action of traditional chinese medicine.
Keywords: amino acid metabolomics, central composite design, metabolic pathways, cerebral ischemia, DanHong injection, HPLC-FLD test, traditional Chinese medicine
Introduction
Ocular surface squamous neoplasia (OSSN) is the most common malignant tumor of the ocular surface with a high incidence in men between 50 and 75 years of age and at latitude of 16 degree south.1,2
Conjunctival squamous cell carcinoma (CSCC) has been causally related to UV-B radiation, human papilloma virus, cigarette smoking, immunosuppressant, and HIV infection.3
CSCC involves the conjunctival interpalpebral zone in a diffuse, nodular, papilliform, gelatinous, leukoplakic or invasive fashion.4
Tumor sizes were divided into small, medium, and large tumors. Large and diffuse tumors were defined as those more than 5 limbal clock hours or more than 8 mm in largest diameter.5,6
The mainstays for the diagnosis of OSSN encompass the spectrum of conjunctival and corneal intraepithelial neoplasia and CSCC remains histopathologic evaluation after a surgical excision.7
Anterior segment optical coherent tomography (AS-OCT) can yield an optical biopsy of tissue, helping in the diagnosis and follow-up of ocular surface lesions. The appearance of OSSN on HR-OCT is a thickened, hyper-reflective epithelium, and an abrupt transition between the normal and affected epithelium.8,9
CSCC is considered a low-grade malignancy. Recurrence is generally higher for severe grades of OSSN and depends also on the adequacy of malignancy free margins at the primary excision. Ascertainment of tumor edges and deep margins are difficult.10–12
The treatment options for conjunctival epithelial malignancies include tumor removal with or without cryotherapy, radiotherapy, electron-based external beam radiation therapy, or topical chemotherapy (i.e., interferon, mitomycin C, or 5-fluorouracil). Primary chemotherapy has also been used as an isolated therapy. Larger, multifocal, or aggressive tumors have required combinations of all previous lines of therapy.13–15
Mitomycin C (MMC) has a significant anti-tumor activity and is effective for treatment of superficial OSSN allowing the treatment of the entire ocular surface including the conjunctival fornices, destroy subclinical disease and prevent new tumors growth elsewhere on the ocular surface.14,16
Amniotic membrane graft (AMG) has been proven to be an effective technique for the management of ocular surface reconstructions following the excision of ocular surface squamous tumors.17,18
Diffuse CSCC has been considered as a therapeutic challenge, despite the available therapeutic modalities. In this cohort, we carried out a prospective analysis to assess the outcomes of diffuse CSCC management. This treatment regimen included surgical resection accompanied with AMG then was followed by postoperative topical MMC 0.04% course. Limited surgical resection of conjunctiva lesion was followed in this cohort to minimize tissue loss.
Subjects And Methods
Patient Selection And Data Collection
Twenty-four cases of diffuse CSCC were included in this nonrandomized prospective study. Diffuse CSCC was defined as a lesion spread more than five clock hours in limbal extent or more than 8 mm in largest diameter.5,6 Fifteen patients underwent amniotic membrane graft after limited resection of diffuse CSSC followed by postoperative topical mitomycin C 0.04% eye drop (MMC group). Nine patients underwent amniotic membrane graft after excision of diffuse CSCC (no MMC group). One surgeon (T.E.) performed all surgeries.
Complete ophthalmic examinations, anterior segment slit-lamp photography, AS-OCT was performed preoperatively and at all subsequent follow-up visits. Data collection included age, sex, laterality, visual acuity, neoplasm characters, AM graft size, histopathological results, recurrence, complications, surface healing, patient comfort (according to the Ocular Surface Disease Index (OSDI) questionnaire) and cosmetic appearance (according to the grades of conjunctival injection as none: 0, mild: 1, moderate: 2, and severe: 3).18
Exclusion criteria were lesions with localized spread (less than five clock hours in limbal extent) and primary chemotherapy treatment.
Imaging
Slit-lamp photography: anterior segment digital images were taken by slit-lamp cam (5.0, integrated and 5 megapixels) attached to the slit-lamp microscope (Zeiss 220 SL, Carl Zeiss Meditec AG, Jena, Germany).
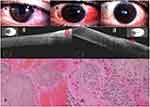
AS-OCT: Anterior Segment 5 Line Raster scanning protocol of Cirrus HD-OCT 5000 system (Carl Zeiss Meditec AG, Jena, Germany) was used in ocular surface lesions imaging. The epithelial thickness of the tumor was measured on its greatest dimension by the distance-measuring tool. Hyper-reflectivity and hypo-reflectivity of conjunctival epithelium and subepithelial component were defined as increased whiteness and darkness compared to normal subjects, respectively.9
The appearance of CSCC on HR-OCT is a thickened, hyper-reflective epithelium and an abrupt transition between the normal and affected epithelium.8,9
The diagnosis was based on clinical and AS-OCT specific criteria of CSSC and was confirmed by histopathological results. Staging of tumors was done according to AJCC classification.
Treatment
Surgery
In MMC group, clinically apparent healthy conjunctiva was marked. The corneal and conjunctival lesions were excised after corneal epithelial scraping. In no MMC group, the corneal and conjunctival lesions excision involves 2-mm margin of normal-looking epithelium was done, then double freeze-thaw cryotherapy was applied to all surgical margins and allowed to thaw. A “no-touch” excision technique was used in both groups.19,20
Excised specimens were sent for histopathologic diagnosis. The amniotic membrane graft (Amnio Tek [3*3 cm]; ISP Surgical) was cut to fit the size of conjunctival defect then was secure with the epithelial side facing up and attached to the sclera and conjunctiva using fibrin glue (Tisseel®, Baxter Inc., Deerfield, IL, USA). Eye pressure patch was applied for 24 hrs to provide a tight graft adherence.
Postoperatively,
- Topical mitomycin C 0.04% eye drop is prescribed 4 times daily per treatment cycle (1 cycle=1 week on & 1 week off) in MMC group. The treatment cycles were repeated if any tumor persistence or recurrence detected clinically and on AS-OCT images.
- Tobramycin–dexamethasone eye drops four times daily on a tapered regimen and a lubricating eye drop were prescribed for a month.
Follow-up visits were performed weekly during the 1st month following the treatment, monthly for 1st year, then twice yearly.
The Primary Outcome Measure
The recurrence of tumor which was detected clinically and/or on AS-OCT at 2-year follow-up while the secondary outcomes included cosmetic appearance and surgical or MMC-related complications.
Study Approval
The study was approved by the local ethics committee of advanced center for daycare surgery, Abu Dhabi, UAE. This study has followed the Tenets of the Declaration of Helsinki and its later amendments or comparable ethical standards. Informed consent was obtained from all individual participants included in the study.
ClinicalTrials.gov ID is NCT03515954
Statistical Analysis
Variables were revealed as a mean±standard deviation or percentage. Paired Student’s t-test was used for variables analysis. A P-value less than 0.05 was considered statistically significant. Statistical analysis was performed using Windows SPSS software version 16.0 (SPSS, Inc., Chicago, IL) (Figures 1 and 2).
Results
This is a prospective study included 24 eyes of 24 patients. Mean postoperative follow-up was 27.1±4.1 (range, 19–31) months. The study population was similar between the two groups; the mean age of patients was 57.4±12.6 years with 11 males (73.3%) and 4 females (26.7%) in MMC group and 60.4±3.8 years with 6 males (66.7%) and 3 females (33.3%) in no MMC group. One patient showed an atypical presentation of CSCC; the patient was young (35 years old) and HIV negative.
The mean extent of the limbal involvement was 7.0±0.7 and 7.25±0.6 clock hours in MMC group and no MMC group, respectively. According to AJCC classification, there were 7 (46.7%) T2 tumors and 8 (53.3%) T3 tumors in MMC group and 3 (22.2%) T2 tumors and 6 (77.8%) T3 tumors in no MMC group. The mean size of conjunctival defect after tumor excision in MMC group (60.8±9.2 mm2) was significantly lower than in no MMC group (92.4±17.3mm2) (P value <0.0001). Mean operative time was 38.6±2.7and 39.8±3.2 mins in MMC group and no MMC group, respectively.
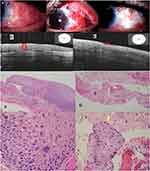
Histopathological diagnosis revealed invasive CSCC in all specimens. Severe dysplasia was associated with invasive SCC in 3 patients in MMC group. Involvement of the excision margin was reported in 6 (40%) and 1 (11%) specimens in MMC and no MMC groups, respectively.
BCVA changed insignificantly from 0.3±0.25 and 0.32±0.18 log MAR preoperatively to 0.25±0.17 and 0.27±0.17 log MAR at 2-year follow-up in MMC group and no MMC group, respectively.
The mean OSDI score changed insignificantly from 10.8±0.25 and 10.4±2.9 preoperatively to 10.0±3.1 and 0.27±3.5 at 2-year follow-up in MMC group and no MMC group, respectively.
According to conjunctival injection grades, there were 12 (80%) grade 0 and 3 (20%) grade 1 in MMC group and 6 (75%) grade 0 and 2 (25%) grade 1 in no MMC group.
The mean preoperative conjunctival epithelial thickness decreased significantly in both groups at 2-year follow-up. It decreased from 267±24.2 & 256±19.1 μm preoperatively to 56.7±11.6 & 60.4±9.6 μm at 2-year follow-up in MMC group and no MMC group, respectively (P value <0.0001). AS-OCT images showed characteristic appearance of CSCC in the form of a thickened, hyper-reflective epithelium and an abrupt transition between the normal and affected epithelium which correlate with histopathology results of primary, postoperative subclinical disease, and recurrent cases. The mean number of MMC treatment cycles was 2.1±0.85 cycles. Recurrence was recorded in zero (0%) and 1 (11%) patients at 2year follow-up in MMC and no MMC groups, respectively (Table 1).
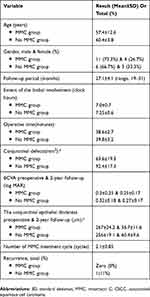 |
Table 1 Summary Of Demographic And Clinical Results |
In MMC group, 3 patients suffered from epiphora, which was relived after nasolacrimal syringing and conjunctival hyperemia and punctuate keratopathy but disappeared on cessation topical MMC. No major operative or MMC-related complications were recorded.
Discussion
We conducted a prospective study on 24 eyes with diffuse CSCC. Fifteen eyes underwent combined surgical and topical chemotherapy treatment lines starting with lesion resection followed by postoperative topical mitomycin C 0.04% eye drop (MMC group) while nine eyes underwent complete excision and cryotherapy (no MMC group). The conjunctival defect was covered with AMT secured by fibrin glue. The diagnosis and treatment of CSCC was monitored by AS-OCT.
The study results were similar between the two groups except the size of conjunctival defect after tumor excision was significantly lower in MMC group than in no MMC group and we think that minimized the burden of surgery on ocular surface and improved postoperative patient comfortless. The recurrence rate was 0% and 11% in MMC and no MMC groups, respectively.
Early diagnosis and proper management of OSSN can prevent significant ocular morbidity.13
The treatment of larger CSCC is a challenging task for all forms of therapy.21
A perfect treatment of OSSN should achieve complete tumor eradication, proper reconstruction of the lost tissue and achieving good functional and cosmetic appearance.22
The primary surgical resection with lesion-free margins is the mainstay of treatment for OSSN providing advantages of surgical debulking, confirming a definitive histopathological diagnosis and exclude life-threatening invasive carcinoma.20,23
The use of topical or intralesional chemotherapeutic agents such as mitomycin C or IFNa2b has become widespread. However, in the review of the literature, it appears that most centers selectively treat large (>8 mm in diameter) tumors with surgery and adjuvant cryotherapy or chemotherapy.24–26
The lack of prior surgical debulking resulted in the higher persistence and recurrence rates noted in cases with diffuse disease. On the other hand, surgical excision of diffuse conjunctival lesions is potentially hazardous due to the risk of limbal stem cell failure and marked tissue loss in the attempt for achieving clear surgical margins. Moreover, the recurrence rate is still high, ranging from 15% to 52%, with the sole primary excision even with lesion-free margins.10,27,28
AS-OCT is a useful adjunctive tool for the diagnosis and management of ocular surface lesions. AS-OCT is a noninvasive technique, able to detect subclinical disease, monitoring for OSSN resolution during medical treatment and prevent the premature termination of topical therapy. Disadvantage of AS-OCT concerning OSSN is the inability to exclude invasive disease and assess intracellular characteristics. Besides, the fornix and caruncle lesions are difficult to be scanned.29–31
In our cohort, AS-OCT showed a characteristic appearance of CSCC in all cases helping not only the diagnosis of primary and postoperative subclinical, persistent or recurrent cases, but also monitoring postoperative topical therapy. AS-OCT is a very helpful and adjunctive tool in diagnosis and management of CSCC but the mainstay for the diagnosis of CSCC remains histopathologic evaluation.
Topical chemotherapy such as MMC, 5FU, and IFN a2b has been used as treatment for OSSN. Some case series of large CSCC showed complete tumor resolution with the use of sole topical chemotherapy.5,24,32
On the other hand, some publications reported that not only the efficacy of isolated chemotherapy in large or invasive OSSN is low but also it is contraindicated for invasive tumors explaining that, topical agents unlikely penetrate to the required level to reach the invasive cells.33,34
Topical chemotherapy needs long treatment duration and patient compliance. IFNa2b shows favorable outcomes in the treatment of primary and recurrent OSSN but it requires a longer duration of treatment than is required with MMC, which may be a disadvantage.23,35
Although zero recurrence rate with sole chemotherapy in Shields et al report, they recommended that extensive tumors may be managed by topical mitomycin C following appropriate biopsy. In Albert et al's study, which carried out excisional biopsy and adjunctive cryotherapy for OSSN treatment, nonetheless, the majority of incompletely excised OSSN (25/28) showed no recurrence. Our study and Bello et al case report of giant OSSN carried out the same therapeutic combination by performing an early tumor excision and followed by topical treatment with MMC and IFNa2b, respectively (Table 2).5,6,14,20,22–24,28,35,37–39
 |
Table 2 Previous Reports On Treatment For Diffuse CSCC |
Reports of successful treatment of giant OSSN with topical treatment may give a false sense of clinical effectiveness of topical therapies for all degrees of invasive disease. Until the depth at which topical therapies lose their anti-neoplastic effectiveness is known, treating CSCC with topical agents alone must be cautious.40
The conjunctival defect created by excision of CSCC was repaired with primary closure techniques, but large one often required tissue replacement. The use of AMG has improved the outcomes for CSCC surgery, as a biological bandage covers underlying ocular structures, facilitates healing. Many publications have reported the successful use of AMG in the reconstruction of conjunctival defects following different types of conjunctival tumors and mentioned little postoperative complication as symblepharon, pyogenic granuloma, and fibrosis. Using fibrin glue shortens the operating time and promotes postoperative patient comfort.22,39,41–43
Immunoreduction with interferons or topical MMC chemoreduction allow complete excision of giant or diffuse conjunctival SCC not involving orbit.24,44
Based on previous discussion, we believe that CSSC management notably diffuse type is an enigma and challenge for ocular surgeons. All lines of therapy either isolated or combined have their advantages and disadvantages.
In our cohort, we started with limited surgical resection of the lesion followed by topical treatment with MMC. An early surgical excision gave the advantage of confirmation of diagnosis and lesion debulking to decrease recurrence and persistence of tumor with the minimal tissue loss. Limited excision had advantage of minimal tissue loss but carried the risk of incomplete excision and tumor persistence.
We think that postoperative topical chemotherapy; as MMC can overcome the possibility of incomplete lesion excision and give advantage of assuring tumor resolution, besides, manage any subtle residual epithelial malignancy. AMG application helps in ocular surface reconstruction attains a good cosmetic appearance.
Prospective study compares current approach (limited excision followed by adjuvant MMC chemotherapy) and chemoreduction followed by surgical excision would answer the question of superiority of either technique in the treatment of diffuse and giant CSSC. Limitations of this study include a relatively small cohort size.
Conclusion
The treatment of diffuse CSCC with AS-OCT guided combined therapy includes limited surgical resection, AMG and postoperative topical MMC achieves a complete tumor resolution and good functional and cosmetic outcomes. This combination minimizes the burden of surgery on ocular surface and improves postoperative patient comfortless in our study cohort.
Abbreviations
CSCC, conjunctival squamous cell carcinoma; OSSN, ocular surface squamous neoplasia; MMC, mitomycin C; AS-OCT, anterior segment optical coherent tomography; AMG, amniotic membrane graft; 5FU, 5 Fluorouracil; INF a2b, interferon alpha 2b; H&E, hematoxylin-eosin; HIV, human immunodeficiency virus; AJCC, American Joint Committee on Cancer; OSDI, Ocular Surface Disease Index.
Data Availability Statement
The datasets generated during and/or analyzed during the current study are available from the corresponding author on reasonable request without end date.
Acknowledgment
Part of this study was presented by Dr Tarek Roshdy Elhamaky as free paper abstract in 38th ESOPRS annual meeting, Hamburg, Germany (11–14 September 2019).
Disclosure
No financial support was received for this submission. No substantial contribution was provided for this submission. The authors declare no conflicts of interest in this work.
References
1. Lee GA, Hirst LW. Ocular surface squamous neoplasia. Surv Ophthalmol. 1995;39(6):429–450. doi:10.1016/S0039-6257(05)80054-2
2. Gichuhi S, Ohnuma SI, Sagoo MS, et al. Pathophysiology of ocular surface squamous neoplasia. Exp Eye Res. 2014;1(129):172–182.
3. Yousef YA, Finger PT. Squamous carcinoma and dysplasia of the conjunctiva and cornea: an analysis of 101 cases. Ophthalmology. 2012;119(2):233–240. doi:10.1016/j.ophtha.2011.08.005
4. Ip MH, George CR, Naing Z, et al. Topical cidofovir for treatment-refractory ocular surface squamous neoplasia. Ophthalmology. 2018;125(4):617–619. doi:10.1016/j.ophtha.2017.11.021
5. Gupta A, Muecke J. Treatment of ocular surface squamous neoplasia with Mitomycin C. Br J Ophthalmol. 2010;94:555–558. doi:10.1136/bjo.2009.168294
6. Shields CL, Naseripour M, Shields JA. Topical mitomycin C for extensive, recurrent conjunctival-corneal squamous cell carcinoma1. Am J Ophthalmol. 2002;133(5):601–606. doi:10.1016/S0002-9394(02)01400-9
7. Kao AA, Galor A, Karp CL, et al. Clinicopathologic correlation of ocular surface squamous neoplasms at Bascom palmer eye institute: 2001 to 2010. Ophthalmology. 2012;119:1773–1776. doi:10.1016/j.ophtha.2012.02.049
8. Shousha MA, Karp CL, Canto AP, et al. Diagnosis of ocular surface lesions using ultra-high-resolution optical coherence tomography. Ophthalmology. 2013;120:883–891. doi:10.1016/j.ophtha.2012.10.025
9. Nanji AA, Sayyad FE, Galor A, et al. High-resolution optical coherence tomography as an adjunctive tool in the diagnosis of corneal and conjunctival pathology. Ocul Surf. 2015;13(3):226–235. doi:10.1016/j.jtos.2015.02.001
10. Midena E, DegliAngeli C, Valenti M, et al. Treatment of conjunctival squamous cell carcinoma with topical 5-fluorouracil. Br J Ophthalmol. 2000;84(3):268–272. doi:10.1136/bjo.84.3.268
11. Semenova EA, Milman T, Finger PT, et al. The diagnostic value of exfoliative cytology vs histopathology for ocular surface squamous neoplasia. Am J Ophthalmol. 2009;148:772–778. doi:10.1016/j.ajo.2009.06.015
12. Mittal V, Narang P, Menon V, et al. Primary simple limbal epithelial transplantation along with excisional biopsy in the management of extensive ocular surface squamous neoplasia. Cornea. 2016;35(12):1650–1652. doi:10.1097/ICO.0000000000000953
13. Tunc M, Char DH, Crawford B, et al. Intraepithelial and invasive squamous cell carcinoma of the conjunctiva: analysis of 60 cases. Br J Ophthalmol. 1999;83(1):98–103. doi:10.1136/bjo.83.1.98
14. Shields CL, Demirci H, Marr BP, et al. Chemoreduction with topical mitomycin C prior to resection of extensive squamous cell carcinoma of the conjunctiva. Arch Ophthalmol. 2005;123:109–113. doi:10.1001/archopht.123.1.109
15. Birkholz ES, Goins KM, Sutphin JE, et al. Treatment of ocular surface squamous cell intraepithelial neoplasia with and without mitomycin C. Cornea. 2011;30:37–41. doi:10.1097/ICO.0b013e3181dee560
16. Prabhasawat P, Tarinvorakup P, Tesavibul N, et al. Topical 0.002% mitomycin C for the treatment of conjunctival-corneal intraepithelial neoplasia and squamous cell carcinoma. Cornea. 2005;24(4):443–448. doi:10.1097/01.ico.0000148314.86557.6a
17. Asoklis RS, Damijonaityte A, Butkiene L, et al. Ocular surface reconstruction using amniotic membrane following excision of conjunctival and limbal tumors. Eur J Ophthalmol. 2011;21(5):552–558.
18. Cheng AM, Zhao D, Chen R, et al. Accelerated restoration of ocular surface health in dry eye disease by self-retained cryopreserved amniotic membrane. Ocul Surf. 2016;14(1):56–63. doi:10.1016/j.jtos.2015.07.003
19. Rudkin AK, Muecke JS. Adjuvant 5-fluorouracil in the treatment of localized ocular surface squamous neoplasia. Br J Ophthalmol. 2011;95:947–950. doi:10.1136/bjo.2010.186171
20. Li AS, Shih CY, Rosen L, et al. Recurrence of ocular surface squamous neoplasia treated with excisional biopsy and cryotherapy. Am J Ophthalmol. 2015;160(2):213–219. doi:10.1016/j.ajo.2015.04.027
21. Barbazetto IA, Lee TC, Abramson DH. Treatment of conjunctival squamous cell carcinoma with photodynamic therapy. Am J Ophthalmol. 2004;138(2):183–189. doi:10.1016/j.ajo.2004.04.007
22. Hanada K, Nishikawa N, Miyokawa N, et al. Long-term outcome of amniotic membrane transplantation combined with mitomycin C for conjunctival reconstruction after ocular surface squamous neoplasia excision. Int Ophthalmol. 2017;37(1):71–78. doi:10.1007/s10792-016-0231-6
23. Bello CM, Pérez AL, Piera JN. Giant conjunctival squamous cell carcinoma. Treatment with surgery following topical interferon alpha-2b. Arch Soc Esp Oftalmol. 2016;91(4):188–190. doi:10.1016/j.oftale.2015.12.018
24. Kim HJ, Shields CL, Shah SU, et al. Giant ocular surface squamous neoplasia managed with interferon alpha2b as immunotherapy or immunoreduction. Ophthalmology. 2012;119:938–944. doi:10.1016/j.ophtha.2011.11.035
25. Nanji AA, Sayyad FE, Karp CL. Topical chemotherapy for ocular surface squamous neoplasia. Curr Opin Ophthalmol. 2013;24(4):336–342. doi:10.1097/ICU.0b013e3283622a13
26. Rudkin AK, Dempster L, Muecke JS. Management of diffuse ocular surface squamous neoplasia: efficacy and complications of topical chemotherapy. Clin Experiment Ophthalmol. 2015;43(1):20–25. doi:10.1111/ceo.2015.43.issue-1
27. Al-Barrag A, Al-Shaer M, Al-Matary N, et al. 5-Fluorouracil for the treatment of intraepithelial neoplasia and squamous cell carcinoma of the conjunctiva, and cornea. Clin Ophthalmol. 2010;4:801. doi:10.2147/OPTH.S9709
28. Joag MG, Sise A, Murillo JC, et al. Topical 5-fluorouracil 1% as primary treatment for ocular surface squamous neoplasia. Ophthalmology. 2016;123(7):1442–1448. doi:10.1016/j.ophtha.2016.02.034
29. Rodriguez M, Yesilirmak N, Chhadva P, et al. High-resolution optical coherence tomography in the differentiation of inflammatory versus non-inflammatory peripheral corneal thinning. Cornea. 2017;36(1):48. doi:10.1097/ICO.0000000000001023
30. Janssens K, Mertens M, Lauwers N, et al. To study and determine the role of anterior segment optical coherence tomography and ultrasound biomicroscopy in corneal and conjunctival tumors. J Ophthalmol. 2016;2016.
31. Chin EK, Cortés DE, Lam A, et al. Anterior segment OCT and confocal microscopy findings in atypical corneal intraepithelial neoplasia. Cornea. 2013;32(6):875–879. doi:10.1097/ICO.0b013e318285cab9
32. Ramasubramanian A, Shields CL, Sinha N, et al. Ocular surface squamous neoplasia after corneal graft. Am J Ophthalmol. 2010;149:
33. Holcombe DJ, Lee GA. Topical interferon alfa-2b for the treatment of recalcitrant ocular surface squamous neoplasia. Am J Ophthalmol. 2006;142:568–571. doi:10.1016/j.ajo.2006.05.058
34. Acharya N, Ramappa R, Thomas RK. Clinical profile of Ocular Surface Squamous Neoplasia: a retrospective case series. J Evol Med Dent Sci. 2016;5(21):1100–1103. doi:10.14260/jemds
35. Chaugule SS, Park J, Finger PT. Topical chemotherapy for giant ocular surface squamous neoplasia of the conjunctiva and cornea: is surgery necessary? Indian J Ophthalmol. 2018;66(1):55. doi:10.4103/ijo.IJO_590_17
36. Bahrami B, Greenwell T, Muecke JS. Long‐term outcomes after adjunctive topical 5‐fluorouracil or mitomycin C for the treatment of surgically excised, localized ocular surface squamous neoplasia. Clin Experiment Ophthalmol. 2014;42(4):317–322. doi:10.1111/ceo.2014.42.issue-4
37. Frucht-Pery J, Rozenman Y, Pe’er J. Topical mitomycin-C for partially excised conjunctival squamous cell carcinoma. Ophthalmology. 2002;109(3):548–552. doi:10.1016/S0161-6420(01)00967-8
38. Khokhar S, Soni A, Singh Sethi H, et al. Combined surgery, cryotherapy and mitomycin-C for recurrent ocular surface squamous neoplasia. Cornea. 2002;21:189–191. doi:10.1097/00003226-200203000-00012
39. Palamar M, Kaya E, Egrilmez S, et al. Amniotic membrane transplantation in surgical management of ocular surface squamous neoplasias: long-term results. Eye. 2014;28(9):1131. doi:10.1038/eye.2014.148
40. Jacinto FA, Margo CE. Clinically occult squamous cell carcinoma of conjunctiva after topical immunotherapy for ocular surface squamous neoplasia. Can J Ophthalmol. 2017;52(4):e152–3. doi:10.1016/j.jcjo.2017.01.009
41. Shields CL, Shields JA, Armstrong T. Management of conjunctival and corneal melanoma with surgical excision, amniotic membrane allograft, and topical chemotherapy. Am J Ophthalmol. 2001;132(4):576–578. doi:10.1016/S0002-9394(01)01085-6
42. Ozcan A, Esen E, Ciloglu E. Sutureless amniotic membrane transplantation following excision of ocular surface neoplasia. Int J Ophthalmol. 2015;8(3):637.
43. Agraval U, Rundle P, Rennie IG, et al. Fresh frozen amniotic membrane for conjunctival reconstruction after excision of neoplastic and presumed neoplastic conjunctival lesions. Eye. 2017;31(6):884. doi:10.1038/eye.2016.322
44. Meel R, Dhiman R, Sen S, et al. Immunoreduction of locally advanced orbito-conjunctival squamous cell carcinoma with intraorbital interferon alpha-2b injection: a globe saving approach. Clin Experiment Ophthalmol. 2018;46(1):87–88. doi:10.1111/ceo.2018.46.issue-1
45. Parrozzani R, Frizziero L, Trainiti S, et al. Topical 1% 5-fluorouracil as a sole treatment of corneoconjunctival ocular surface squamous neoplasia: long-term study. Br J Ophthalmol. 2017;101(8):1094–1099. doi:10.1136/bjophthalmol-2016-309219
 © 2019 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.
© 2019 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.