Back to Journals » International Journal of General Medicine » Volume 15
A Retrospective Cohort Analysis of the Genetic Assay Results of Foetuses with Isolated and Nonisolated Umbilical Cord Cyst
Authors Liu Q, Wei R, Lu J, Ding H, Yi H, Guo L, Wu J
Received 17 February 2022
Accepted for publication 16 June 2022
Published 23 June 2022 Volume 2022:15 Pages 5775—5784
DOI https://doi.org/10.2147/IJGM.S358864
Checked for plagiarism Yes
Review by Single anonymous peer review
Peer reviewer comments 3
Editor who approved publication: Dr Scott Fraser
Qian Liu, Ran Wei, Jian Lu, Hongke Ding, Hui Yi, Li Guo, Jing Wu
Department of Medical Genetics Center, Guangdong Women and Children Hospital, Guangzhou City, Guangdong Province, People’s Republic of China
Correspondence: Jing Wu, Department of Medical Genetics Center, Guangdong Women and Children Hospital, NO. 521 Xingnan Road, Panyu District, Guangzhou city, Guangdong Province, People’s Republic of China, Tel +86 20-39151548, Email [email protected]
Objective: To analyse the risk of clinical chromosomal abnormalities in foetuses with umbilical cord cysts.
Methods: Data from all genetic assays that were performed as part of invasive prenatal diagnoses of umbilical cord cysts between October 2014 and June 2021 were retrospectively collected from Guangdong Women and Children Hospital. We compared the differences in genetic assay findings in isolated and nonisolated umbilical cord cyst cohorts.
Results: A total of 49 singleton pregnancies and 2 foetuses that were one of the cotwins in monochorionic twin pregnancies were enrolled in the cohort; 20 isolated and 31 nonisolated umbilical cord cysts were identified in the cohort. One foetus (5%, 1/20) in the isolated umbilical cord cyst group showed chromosomal abnormalities and 17p12 microduplication. Twelve cases (38.7%, 12/31) of chromosomal abnormalities, including seven cases of trisomy 18, two cases of trisomy 13 and three cases of microdeletion, were identified in the nonisolated umbilical cord cyst group. The incidences of chromosomal abnormalities between the two groups were significantly different (1/20, 5% vs 13/31, 38.7%, p=0.003). There was no relative pathological medical exome sequencing finding in the three foetuses suffering from nonisolated umbilical cord cysts whose parents chose to undergo chromosomal microarray analysis (CMA) and medical exome sequencing.
Conclusion: This retrospective cohort study evaluated the value of CMA in foetuses with umbilical cord cysts and suggested that copy number variants (CNVs) may be the basic genetic aetiological factors that should be considered for diagnostic evaluation. We recommended CMA as a basic genetic evaluation in cases of umbilical cord cysts, especially in nonisolated cases.
Keywords: chromosomal microarray analyses, CMA, umbilical cord cyst, G banding karyotype
Graphical Abstract:
Introduction
Umbilical cord cysts have been described as anechoic masses of the umbilical cord1 that can be detected by prenatal ultrasound from 8 to 9 weeks gestational age when the umbilical cord is well developed.2 Previous studies have reported that the prevalence of umbilical cord cysts in the first trimester ranged from 0.4% to 3.4%.3 Although rare, umbilical cord cysts represent the second most common anomaly of the umbilical cord.4 Umbilical cord cysts can be isolated or nonisolated ultrasound findings during the pregnancy period, and the prognosis varies with the status of multiple or unique presentation and with persistent or progressive size.5 If no other anomaly is found, the prognosis is mostly excellent.
It was documented that the umbilical cord might be considered a prominent prenatal feature of foetal aneuploidy.1,5 With the development of prenatal diagnostic techniques, common foetal aneuploid chromosome abnormalities can be identified by noninvasive prenatal tests (NIPTs).6 However, advice should be given to the parents when the foetus has ultrasound findings of umbilical cord cyst by performing NIPT or invasive prenatal diagnosis with chromosomal microarray analyses (CMA) or other genetic tests, especially in isolated cases. Based on this, we conducted this retrospective cohort study to analyse the foetal chromosome and copy number variation results of isolated and nonisolated umbilical cord cysts.
Materials and Methods
Study Population
This retrospective cohort study was conducted between October 2014 and June 2021 at Guangdong Women and Children Hospital. Genetic assay data from 49 singleton pregnancies and 2 monochorionic diamniotic (MCDA) twin pregnancies diagnosed with umbilical cord cysts were collected. When umbilical cord cysts were suspected or pregnant women were referred from other hospitals for suspicion, ultrasound (Voluson E8 OR Voluson E10, GE Medical Systems, Chicago, IL, USA) was conducted by experienced obstetrical sonographers to confirm the diagnosis and screen additional anomalies thoroughly. Foetal magnetic resonance imaging (MRI) was performed in certain cases in the second or third trimester if necessary.
Confirmed cases were classified into two groups: the isolated umbilical cord cyst group (n = 20) and the nonisolated umbilical cord cyst group (including structural anomalies, n = 24, and sonographic soft markers, n = 7). Foetal samples were obtained through chorionic villi sampling (n = 5), amniocentesis (n = 26) or cord blood sampling (n = 19) at the appropriate gestational age. Fetuses with isolated or non-isolated umbilical cord cyst whose parents refused to undergo interventional prenatal diagnosis were excluded from this cohort and were fully informed of the potential risks.
All parents of foetuses received comprehensive prenatal genetic counselling before the invasive procedures in which they received explanations about the invasive procedures, including the potential risk of the procedure, alternative detection techniques, detection ranges, limitations and the possible significance of the findings. All enrolled foetuses underwent professional genetic testing according to their parents’ choice. G-banding karyotyping and CMA were performed in 41 samples, 3 samples underwent CMA and medical exome sequencing (MES), 2 samples underwent G-banding karyotyping only, and 5 samples underwent CMA only. Informed consent for the treatment was obtained from each pregnant woman accepting invasive prenatal diagnosis.
Ethics/Review Board Approval
The study was performed according to the principles of the Declaration of Helsinki. Ethical approval to conduct this study was obtained from the Research Ethics Committee of Guangdong Women and Children Hospital, Guangzhou, China. All patient details were deidentified. Informed consent from all parents of foetuses enrolled into this study for this research was obtained.
Genetic Assay Analysis
All collected samples were subjected to genetic assays based on the choice of the parents. G-banding karyotype analysis at a level of 320 to 500 bands was performed according to standard procedures. Genomic DNA (gDNA) of foetuses was extracted from uncultured samples using QI Aamp DNA Blood Mini kits (Qiagen, Valencia, CA, USA) and then subjected to CMA or medical exome sequencing (MES).
CMA was conducted using CytoScan 750K Array (Thermo Fisher Scientific, Santa Clara, CA, USA) with 250 ng of gDNA according to the manufacturer’s instructions. Data analysis was conducted using Chromosome Analysis Suite v4.1 (Thermo Fisher Scientific), and the detected copy number variants (CNVs) were assessed for clinical significance based on American College of Medical Genetics and Genomics (ACMG) guidelines.7 The GRCh37 (hg19) genome was used for annotation.
MES was conducted using an Illumina Nextera Rapid Capture Exome Kit (Illumina CA, USA) for library preparation according to the manufacturer’s protocol. Trio-exome gene sequencing was performed using a Nova seq 6000 sequencer (Illumina, CA, USA) according to the manufacturer’s protocols. Data analysis was performed by bioinformation analysis, including 1000 Genomes, dbSNP, GnomAD, ClinVar, HGMD and OMIM. Detected genetic variants were assessed for clinical significance and classified as associated with the foetal phenotype, which was reported in prenatal laboratory and ultrasound findings, based on ACMG guidelines for Medical Pathology recommendations.8
Follow-Up
Clinical follow-up was conducted one month after the invasive procedure and 6 months after the birth using clinical records and multiple interview methods, such as telephone and internet contact.
Data Analysis
After proper categorization, genetic assay results were classified as normal, aneuploidy, pathogenic CNVs and variants of unknown significance (VOUS). The genetic assay results of isolated and non-isolated umbilical cord cyst was presented in Table 1. About data presentation, continuous variables were showed as mean±standard deviations (SD) and categorical data were showed as numbers and proportions.
 |
Table 1 The Genetic Assay Results Abnormal Rate and Pregnancy Outcome of Fetuses with Umbilical Cord Cyst |
Statistical analysis was performed with SPSS 21.0 software (IBM SPSS Statistics for Windows, IBM, USA). The frequency of clinically significant abnormal chromosome results was compared between isolated and nonisolated umbilical cord cyst groups using Fisher’s exact test. Differences were considered significant at P< 0.05.
Results
A total of 49 singleton pregnancies and 2 MCDA twin pregnancies (51 foetuses) were diagnosed with isolated or nonisolated umbilical cord cysts by ultrasound screening based on the obstetric ultrasound screening protocol and received an invasive prenatal diagnostic test at Guangdong Women and Children Hospital. A prenatal diagnosis flow chart used for foetuses affected by isolated and nonisolated umbilical cord cysts is shown in Figure 1. The image of umbilical cord cyst in one of the foetuses is shown in Figure 2. The average maternal age at the time of the invasive prenatal procedure was 29 years (20–40 years), and at the median gestational age of 25.2 weeks (12–35.5 weeks), foetuses were diagnosed with umbilical cord cysts. Five foetuses (9.8%) were diagnosed by ultrasound in the first trimester, and the other 46 foetuses (90.2%) were diagnosed in the 2nd or 3rd trimester.
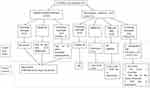 |
Figure 1 The flow chart of the study. |
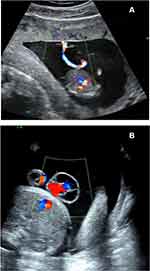 |
Figure 2 (A) Color Doppler ultrasound image of an umbilical cyst near the placenta. (B) Color Doppler ultrasound image of multiple small cysts around the umbilical cord. |
In one MCDA twin pregnancy case, both foetuses underwent amniocentesis and got normal chromosome and CMA result, with the indication of one twin suffering from umbilical cord cyst complicated by omphalocele and choroid plexus cyst and the cotwin suffering from choroid plexus cyst. The foetus suffering from umbilical cord cyst accompanied by omphalocele died intrauterine in the third trimester, and the co-twin was born alive at 36 weeks gestational age. The other MCDA twin pregnancy case was diagnosed with twin-to-twin transfusion Phase III (TTTs) and selected intrauterine growth restriction (sIUGR), the recipient suffered from umbilical cord cysts and TTTs with relative imaging changes, and the foetus was small for gestational age, which underwent amniocentesis and found 20p12.3 micro duplication (VUS). However, the pregnancy was terminated due to the parental choice considering the severity and prognosis of TTTs.
The results of the prenatal genetic assay are presented in Table 2. One of isolated umbilical cord cyst foetuses (5%, 1/20) accept the amniocentesis with the indication of ultrasound finding and positive result of expanding NIPT of 17 micro duplication, with the CNVs 17p12 microduplication detected. Twelve cases (38.3%, 12/31) with chromosomal abnormalities, including seven cases of trisomy 18, two cases of trisomy 13 and three cases of microdeletion, were identified in the nonisolated umbilical cord cyst group. The incidences of chromosomal abnormalities between the two groups were significantly different (1/20, 5% vs 12/31, 38.3%, p=0.003), although the sample size may be too small to reach a conclusion on this point.
 |
Table 2 Genetic Assay Results of Fetuses with Umbilical Cord Cyst |
The clinical characteristics of the fetuses with CNVs detected by CMA are shown in Table 3. Thirteen cases of chromosomal abnormality (13/51, 25.5%) were confirmed through karyotyping and/or CMA. Nine of them were aneuploid, including 7 cases of trisomy 18 and 2 cases of trisomy 13, and all of them showed severe abnormalities by ultrasound examination. The results of six cases with pathogenic CNVs and VOUS, including three cases micro-deletion (15q11.2q13.3 deletion, 22q11.21 deletion and 9q34.3 deletion) and three cases micro-duplication (17p12, 20p12.3 and 6p13.11), the prenatal genetic assay result and prenatal outcome detail are shown in Table 3. Three nonisolated cases underwent medical exome sequencing and did not have relative pathogenic mutation findings.
 |
Table 3 Characteristic of the Fetuses with CNVs Detected by CMA |
The pregnancy outcomes of isolated and non-isolated umbilical cord cyst are shown in Table 1. Out of 51 foetuses, 32 were born alive (62.7%). The foetus with deletion of 15q11.2q13.3 was prematurely born several days before the result of CMA was reported, and the diagnosis of Prader-Willi syndrome (PWS) was confirmed by a methylation test postpartum. The foetus showed 16p11.11 duplication, with the ultrasound finding of umbilical cord cyst and mild bilateral lateral ventricle enlargement. The duplication was confirmed to be inherited from the mother by the parental CMA test. The parents of the foetus chose to retain the foetus, and no obvious abnormalities were observed after the foetus was born at term.
Discussion
It was reported that in foetuses, umbilical cord cysts may be related to aneuploidy. However, the frequency of abnormal chromosomes in foetuses with umbilical cord cysts is still unclear. There was no relevant conclusion that umbilical cord cyst, especially isolated ones need to accept which test is more appropriate, non-invasive prenatal genetic testing or interventional prenatal diagnosis, which test plan is more appropriate. So we retrospectively analysed the genetic assay results of foetuses with umbilical cord cysts and found that the frequency of abnormal chromosomes was higher in the nonisolated group than in the isolated group (1/20, 5% vs 13/31, 38.7%, p=0.003). We did not find gene mutations related to umbilical cord cysts in three foetuses undergoing medical exome sequencing.
First, in this study, we found that the chromosome clinical abnormal rate in foetuses with isolated umbilical cord cysts was low (1/20, 5%), which was screened by expanding NIPT. In a cohort reported previously, conducted before 2015 year and CMA was not applied, which found that revealed normal karyotype in 5 out of 13 isolated umbilical cord cyst foetuses, and other 8 cases have not done fetal karyotype.5 Both our study and previous studies3,5 found favorable fetal prognosis with isolated cord cysts. We inferred that expanding NIPT for fetuses with isolated cord cysts may be a good option, but lack support from large sample studies.
Second, we found that in foetuses, umbilical cord cysts may indeed be related to aneuploidy, in accordance with previous studies.5,9,10 In this study, we found that the most common abnormal chromosome in foetuses with umbilical cord cysts was trisomy 18 (13.7%, 7/51). Among them, there were three cases accompanied by foetal intracardiac echogenic foci, one case accompanied by hydramnios and subependymal cysts, one case complicated by omphalocele and choroid plexus cyst, and the other two cases were independently associated with pyelectasia and foetal hydrops. In another cohort of umbilical cord cysts, two foetuses showed trisomy 18 among 27 cases.5 In this cohort, we also found two cases of trisomy 13, one of which was accompanied by subependymal cysts and cerebral ventriculomegaly, and the other case was complicated by omphalocele and increased nuchal translucency and nasal bone dysplasia. Foetuses with umbilical cord cysts showing trisomy 13 were also reported in previous studies.11 This aneuploidy can be diagnosed through karyotyping or CMA.
Third, we found that some foetuses with isolated or nonisolated umbilical cord cysts showed pathogenic CNVs. In the present study, there was one foetus in isolated group with positive result of expanding NIPT of 17 micro duplication, which was confirmed by amniotic fluid CMA analysis and rated as pathogenic CNVs, with the repeat fragment size of 1.3Mb. CMA analysis of parental blood revealed that the pregnant woman (29 years old) carried the same microduplication as the foetus. The mother was phenotypically normal, so the parent considered continuing the pregnancy. The foetus was delivered at term and growth well at the beginning of two years of life. It was reported that 17 p12 duplication may be associated with Charcot–Marie–Tooth (CMT) neuropathy, which is a chronic progressive neuropathy affecting both the motor and sensory nerves.12 This duplication may be detected by expanding NIPT in mothers with identical duplication,13 there may even be no genomic imbalance in the fetus, only NIPT false-positives due to maternal CNVs.14 We have not found evidence about that 17P12 duplication is associated with fetal umbilical cord cysts, and this may be an incidental finding that needs further observation.
The nonisolated group presented a higher incidence of pathogenic CNVs (9.6%, 3/31 vs 5%, 1/20, p<0.05). There were three cases of microdeletion in the nonisolated group, including 22q11.21, 15q11.2-q13.3 and 9q34.3 deletions. Chromosome 22q11.2 deletion syndrome (22q11.2del) is the most common chromosomal microdeletion reported in humans and is implicated in congenital heart defects and other mutiorgan defects.15–17 To the best of our knowledge, we are the first to report that foetuses affected by the 22q11 microdeletion showed umbilical cord cysts complicated with severe hydramnios, moderate lateral ventricle broadening and head circumference larger than 2 SD greater than the mean level with the same gestational age but without congenital heart defect ultrasound findings.
Chromosome 15q11.2 deletion syndrome (15q11.2 del) is the first example of imprinting errors in humans, resulting in disorders including PWS and Angelman syndrome,18 with behavioural and academic outcomes in individuals with PWS. This nutrition-related condition impacts the DNA methylation process and not only alters the development of the foetus, but pregnancy complications may result from large foetal size.18 The largest PWS cohort analysed to date showed that 61% of patients with PWS have the typical 15q11-q13 deletion19 and a severe neurodevelopmental disorder caused by this deletion. In our cohort, we found one foetus affected by the 15q11-q13 deletion, with the prenatal ultrasound finding of umbilical cord cyst accompanied by hydramnios and scalp oedema. The pregnant woman was found to have a foetal umbilical cord cyst in the external hospital and was referred to our hospital at 32 weeks of pregnancy. Unfortunately, the foetus was prematurely born 11 days after the cord blood sampling procedure, at 33 weeks gestational age, before the CMA result was reported, and the diagnosis of PWS was confirmed by a methylation test postpartum. The final diagnosis of this case was in accordance with most of the typical 15q11-q13 deletion cases.
Furthermore, we found two cases of monochorionic twin pregnancies, with one of the twins affected by umbilical cord cysts. One case was complicated by TTTs and sIUGR, with the ultrasound finding that the recipient was affected by umbilical cord cyst and oedema; the pregnancy was terminated due to parental choice, although the foetuses showed VOUS CNVs of 20p12.3 microduplication. Another case was accompanied by choroid plexus cysts in both twins, with the foetuses showing a normal karyotype and CMA. Monochorionic twin pregnancies are more likely to be complicated with foetal structural abnormalities,20 but there are few cases of complicated umbilical cysts. An umbilical cord cyst-affected donor monochorionic twin pregnancy complicated by TTTs was reported previously.21 Chih-Ping Chen reported a case of a thoraco-omphalopagus conjoined twin associated with omphalocele and an umbilical cord cyst that showed a normal karyotype.22 The mechanism by which umbilical cord cysts affect twin pregnancy is not clear and needs more research.
Finally, but not the least important, we found that four foetuses with umbilical cord cysts also had omphalocele, with one of them showing trisomy 13 and other foetuses showing normal karyotype and CNVs. Foetal umbilical cord cysts accompanied by omphalocele have also been observed in previous studies.23–26 In addition, one of the four foetuses had umbilical cord cysts, and omphalocele was found in a monochrionicity twin that showed a normal karyotype and CNV. Chih-Ping Chen recorded a case in which Thoraco-Omphalopagus conjoined twins were associated with omphalocele and an umbilical cord cyst.22 Omphalocele is one of the most common anterior abdominal wall defects, and the coexistence of omphalocele with a simple or multiple umbilical cord cyst or pseudocysts has rarely been reported.11,25 The related mechanism is still unknown.
Our result should be interpreted in consideration of several limitations. First, the sample size in the study was relatively small, which means that the study can only provide a general result but cannot indicate a definite conclusion. Second, we did not assess the long-term outcome of foetus survival, which we have documented recently. A prospective study is needed to determine the long-term outcomes of surviving foetuses with isolated or nonisolated umbilical cord cysts.
Conclusion
Among foetuses, umbilical cord cysts may be related to abnormal chromosomes and CNVs, especially in nonisolated cases. Comprehensive genetic counselling and appropriate testing recommendations are necessary when ultrasound screening reveals an umbilical cord cyst.
Data Sharing Statement
The data analyzed in this study are available at https://osf.io/2wdfp/.
Ethics Approval and Informed Consent
Ethical approval to conduct this study was obtained from the Research Ethics Committee of Guangdong Women and Children Hospital, Guangzhou, China.
Consent for Publication
All authors agree to publish all details and images related to this study and sign the consent to publication.
Acknowledgments
The authors thank all colleagues in the Department of Medical Genetics Center, staff members of Guangdong Women and Children Hospital, and the parents and foetuses who participated.
Author Contributions
Jing Wu and Qian Liu conceived the idea for the article and drafted the manuscript. Jian Lu, Li Guo and Hongke Ding respond for genetics assay detection and result analysis. Ran Wei and Hui Yi respond for collecting patient information. All authors have read and approved the final manuscript. Jing Wu is the guarantor for the article. The decision to publish was jointly agreed by all authors. All authors made a significant contribution to the work reported, whether that is in the conception, study design, execution, acquisition of data, analysis and interpretation, or in all these areas; took part in drafting, revising or critically reviewing the article; gave final approval of the version to be published; have agreed on the journal to which the article has been submitted; and agree to be accountable for all aspects of the work.
Disclosure
The authors declare that they have no competing interests.
References
1. Sepulveda W. Beware of the umbilical cord ‘cyst’. Ultrasound Obstetr Gynecol. 2003;21(3):213–214. doi:10.1002/uog.80
2. Moshiri M, Zaidi SF, Robinson TJ, et al. Comprehensive imaging review of abnormalities of the umbilical cord. Radiographics. 2014;34(1):179–196. doi:10.1148/rg.341125127
3. Hannaford K, Reeves S, Wegner E. Umbilical cord cysts in the first trimester: are they associated with pregnancy complications? J Ultrasound Med. 2013;32(5):801–806. doi:10.7863/ultra.32.5.801
4. Kong CKY, Zi Xean K, Li FX, Chandran S. Umbilical cord anomalies: antenatal ultrasound findings and postnatal correlation. BMJ Case Rep. 2018;2018. doi:10.1136/bcr-2018-226651
5. Ruiz Campo L, Savirón Cornudella R, Gámez Alderete F, et al. Prenatal diagnosis of umbilical cord cyst: clinical significance and prognosis. Taiwan J Obstet Gynecol. 2017;56(5):622–627. doi:10.1016/j.tjog.2017.08.008
6. Alberry MS, Aziz E, Ahmed SR, Abdel-Fattah S. Non invasive prenatal testing (NIPT) for common aneuploidies and beyond. Eur J Obstet Gynecol Reprod Biol. 2021;258:424–429. doi:10.1016/j.ejogrb.2021.01.008
7. South ST, Lee C, Lamb AN, Higgins AW, Kearney HM. ACMG Standards and Guidelines for constitutional cytogenomic microarray analysis, including postnatal and prenatal applications: revision 2013. Genet Med. 2013;15(11):901–909. doi:10.1038/gim.2013.129
8. Richards S, Aziz N, Bale S, et al. Standards and guidelines for the interpretation of sequence variants: a joint consensus recommendation of the American College of Medical Genetics and Genomics and the Association for Molecular Pathology. Genet Med. 2015;17(5):405–424. doi:10.1038/gim.2015.30
9. Kuwata T, Matsubara S, Izumi A, et al. Umbilical cord pseudocyst in a fetus with trisomy 18. Fetal Diagn Ther. 2003;18(1):8–11. doi:10.1159/000066376
10. Bahado-Singh RO, Choi SJ, Oz U, Mendilcioglu I, Rowther M, Persutte W. Early second-trimester individualized estimation of trisomy 18 risk by ultrasound. Obstet Gynecol. 2003;101(3):463–468. doi:10.1016/s0029-7844(02)03078-8
11. Suzuki T, Yamamoto Y, Nakamura H, et al. Fetal umbilical cord cyst may evolve to omphalocele during pregnancy. J Clin Ultrasound. 2019;48(3):181–183. doi:10.1002/jcu.22786
12. Salpietro V, Manole A, Efthymiou S, Houlden H. A review of copy number variants in inherited neuropathies. Curr Genomics. 2018;19(6):412–419. doi:10.2174/1389202919666180330153316
13. Chen CP, Chen SW, Wu PS, Wu FT, Wang W. A false-positive result at non-invasive prenatal testing due to maternal 17p12 microduplication. Taiwan J Obstet Gynecol. 2022;61(3):532–534. doi:10.1016/j.tjog.2022.03.037
14. Kumps C, Niel Bütschi F, Rapin B, et al. Non-invasive prenatal testing leading to a maternal diagnosis of Charcot–Marie–Tooth neuropathy. J Hum Genet. 2020;65(11):1035–1038. doi:10.1038/s10038-020-0789-8
15. Du Q, De la morena MT, van Oers N. The genetics and epigenetics of 22q11.2 deletion syndrome. Front Genet. 2019;10:1365. doi:10.3389/fgene.2019.01365
16. Lv W, Wang S. Detection of chromosomal abnormalities and the 22q11 microdeletion in fetuses with congenital heart defects. Mol Med Rep. 2014;10(5):2465–2470. doi:10.3892/mmr.2014.2564
17. Li S, Han X, Ye M, et al. Prenatal diagnosis of microdeletions or microduplications in the proximal, central, and distal regions of chromosome 22q11.2: ultrasound findings and pregnancy outcome. Front Genet. 2019;10:813. doi:10.3389/fgene.2019.00813
18. Butler MG. Imprinting disorders in humans: a review. Curr Opin Pediatr. 2020;32(6):719–729. doi:10.1097/MOP.0000000000000965
19. Butler MG, Hartin SN, Hossain WA, et al. Molecular genetic classification in Prader-Willi syndrome: a multisite cohort study. J Med Genet. 2019;56(3):149–153. doi:10.1136/jmedgenet-2018-105301
20. Yamamoto R, Nakanishi K, Kawaguchi H, Hayashi S, Ishii K. Prevalence of extraplacental anastomoses in monochorionic twin pregnancies. Fetal Diagn Ther. 2021;48(1):24–27. doi:10.1159/000510636
21. Jacquemyn Y, Markov D, Beckstedde I. Umbilical cord cyst in a monochorionic twin pregnancy: an experiment of nature for the treatment of twin-twin transfusion syndrome. Fetal Diagn Ther. 2002;17(4):233–235. doi:10.1159/000059375
22. Chen CP. Thoraco-omphalopagus conjoined twins associated with omphalocele and an umbilical cord cyst. Taiwan J Obstet Gynecol. 2007;46(2):183–184. doi:10.1016/S1028-4559(07)60017-5
23. Suzuki T, Yamamoto Y, Nakamura H, et al. Fetal umbilical cord cyst may evolve to omphalocele during pregnancy. J Clin Ultrasound. 2020;48(3):181–183. doi:10.1002/jcu.22786
24. Sharma D, Murki S, Pratap T. A newborn with omphalocele and umbilical cord cyst: an interesting entity. Iran J Pediatr. 2014;24(4):449–450.
25. Stella A, Babbo GL. Omphalocele and umbilical cord cyst. Prenatal diagnosis. Minerva Ginecol. 2000;52(5):213–216.
26. Emura T, Kanamori Y, Ito M, et al. Omphalocele associated with a large multilobular umbilical cord pseudocyst. Pediatr Surg Int. 2004;20(8):636–639. doi:10.1007/s00383-004-1247-y
 © 2022 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.
© 2022 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.
