Back to Journals » Clinical and Experimental Gastroenterology » Volume 12
A pilot study of fecal bile acid and microbiota profiles in inflammatory bowel disease and primary sclerosing cholangitis
Authors Vaughn BP , Kaiser T , Staley C , Hamilton MJ, Reich J, Graiziger C, Singroy S, Kabage AJ , Sadowsky MJ, Khoruts A
Received 1 September 2018
Accepted for publication 10 December 2018
Published 10 January 2019 Volume 2019:12 Pages 9—19
DOI https://doi.org/10.2147/CEG.S186097
Checked for plagiarism Yes
Review by Single anonymous peer review
Peer reviewer comments 2
Editor who approved publication: Professor Wing-Kin Syn
Byron P Vaughn,1 Thomas Kaiser,2,3 Christopher Staley,2,3 Matthew J Hamilton,2 Jon Reich,1 Carolyn Graiziger,1 Stephanie Singroy,2 Amanda J Kabage,1 Michael J Sadowsky,2,4,5 Alexander Khoruts1,2
1Inflammatory Bowel Program, Division of Gastroenterology, Hepatology and Nutrition, University of Minnesota, Minneapolis, MN, USA; 2BioTechnology Institute, University of Minnesota, St Paul, MN, USA; 3Department of Surgery, University of Minnesota, Minneapolis, MN, USA; 4Department of Soil, Water, and Climate, University of Minnesota, St Paul, MN, USA; 5Department of Plant and Microbial Biology, University of Minnesota, St Paul, MN, USA
Introduction: Inflammatory bowel disease (IBD) is thought to arise from an abnormal immune response to the gut microbiota. IBD is associated with altered intestinal microbial community structure and functionality, which may contribute to inflammation and complications such as colon cancer and liver disease. Primary sclerosing cholangitis (PSC) is associated with IBD and markedly increases the risk of colon cancer. We hypothesized that secondary bile acids, which are products of microbial metabolism, are increased in PSC patients.
Aim: Here, we profiled the fecal bile acid composition and gut microbiota of participants with IBD and PSC, as well as healthy participants. Additionally, we tested the effects of vancomycin, a proposed treatment for PSC, on gut microbiota and fecal bile acid composition in participants with IBD and PSC.
Methods: Fecal samples were collected from patients with IBD, IBD/PSC and healthy controls and fecal bile acids and DNA for microbiota analysis were extracted. Fecal bile acids were averaged over a seven-day period. For subjects with IBD/PSC, oral vancomycin 500mg twice a day was administered and fecal samples were collected for up to eleven weeks.
Results: Participants with IBD and PSC had less fecal microbial diversity at baseline relative to controls. While there was some evidence of altered conversion of cholic acid to deoxycholic acid, no substantial differences were found in the fecal bile acid profiles of patients with IBD and PSC (n=7) compared to IBD alone (n=8) or healthy controls (n=8). Oral vancomycin was a potent inhibitor of secondary bile acid production in participants with IBD and PSC, particularly deoxycholic acid, although no changes in liver biochemistry patterns were noted over a two week period.
Conclusion: In this pilot study, bile acid profiles were overall similar among patients with IBD and PSC, IBD alone, and healthy controls. Microbiota diversity was reduced in those with PSC and IBD compared to IBD alone or healthy controls.
Keywords: vancomycin, Crohn’s disease, ulcerative colitis
Introduction
Inflammatory bowel disease (IBD) is thought to arise from an abnormal immune response to the intestinal microbiota in a genetically susceptible host.1 In North America and Europe, 3%–5% of people with IBD will develop primary sclerosing cholangitis (PSC), a chronic inflammatory and fibrotic disease of the biliary tree.2 Those with IBD who develop PSC commonly have a unique phenotype of IBD characterized by pancolitis with rectal sparing and backwash ileitis.3 The presence of PSC in the setting of IBD with colonic involvement dramatically increases the risk of colonic dysplasia and cancer, particularly in the right colon.4 The etiology for this PSC/IBD phenotype, including the increased risk of colon cancer, is unknown, although intestinal microbiota may have a role. Intestinal microbiota modifies primary bile acids, which are produced in the liver, into secondary bile acids, lithocholic acid (LCA), and deoxycholic acid (DCA) in the colon. These secondary bile acids are thought to be carcinogenic in the colon, and increased concentrations of secondary bile acids have been noted in the stool of patients with ulcerative colitis (UC) and dysplasia or cancer.5–7 Secondary bile acids also stimulate the bile acid receptor TGR5 (LCA > DCA), which results in suppression of the inflammatory response to microbial ligands, such as lipopolysaccharide (LPS),8 potentially leading to a lesser degree of immunosurveillance against neoplasms.
The microbiota of PSC patients was previously shown to have increased abundances of Blautia and Barnesiellaceae.9 Certain Blautia species can carry out the 7-alpha-dehydroxylation of primary bile acids, a key step in their conversion to secondary bile acids.10 Therefore, we hypothesized that patients with PSC/IBD have increased fecal concentrations of secondary bile acids, which contributes to the unique phenotype of PSC/IBD and increased colon cancer risk. Here, we measured the fecal bacterial and bile acid composition of IBD patients with and without PSC, as well as healthy controls. In addition, we investigated the effects of oral vancomycin on fecal microbiota and bile acid composition in patients with PSC/IBD. Oral vancomycin is known to target bacterial taxa involved in secondary bile acid metabolism and is currently being studied for a potential therapeutic benefit for PSC patients.11,12
Methods
Participant population
Participants were recruited from the University of Minnesota IBD program and liver clinics. All participants were provided, and signed, written informed consent and the study was conducted in accordance with the Declaration of Helsinki. After informed consent, all participants’ histories were reviewed to determine IBD history, medication history, and PSC history. Those with a history of decompensated cirrhosis, antibiotic or probiotic use within 30 days, or ursodiol use within 30 days were excluded. Disease activity was measured with the Harvey–Bradshaw Index for Crohn’s disease and partial Mayo score for ulcerative colitis. Serum and fecal samples were collected from all participants at baseline. Serum samples were analyzed for liver function tests through the hospital’s clinical laboratory. Participants with PSC/IBD were asked to participate in a longitudinal study consisting of 2 weeks of oral vancomycin (500 mg twice a day) and up to 3 months of bi-weekly fecal collections. To account for daily variations of bile acids, participants were asked to collect 3–5 stool samples over the course of one week. Stool was frozen at –80°C for microbiota and bile acid analysis. The study was approved by the institutional review board at the University of Minnesota and registered at clinicaltrials.gov (NCT02464020).
Bile acid analysis
Raw fecal samples were suspended in 50% acetonitrile in 1:1 (w/v) ratio. Samples were vortexed until no large particles remained and sonicated for 10 minutes to release bile acids into solution. The suspension was centrifuged at 18,000× g for 10 minutes, and 100 µL of supernatant was removed and transferred to a HPLC vial. Vials were stored at –20°C until use. A 5 µL sample was injected into an Acuity ultraperformance LC–MS system (Waters, Milford, MA, USA) for bile acid detection. Instrument parameters were set as previously described.13 Mass chromatograms and spectral data were analyzed using the MassLynx software package (Version 4.1; Waters). The concentration of bile acids in the fecal samples was determined using the peak areas of individual bile acids and a standard curve generated using samples of known bile acid concentration. Samples were normalized using a C13-radiolabeled glycolic acid internal standard.
Microbiota analysis
DNA was extracted from ~250 mg of thawed fecal samples using the DNeasy PowerSoil Kit (Qiagen, Hilden, Germany), according to the manufacturer’s instructions. The V5–V6 hypervariable regions of the 16S rRNA gene were amplified using the BSF784/1064 primer set.14 Paired-end sequencing was done at a read length of 300 nucleotides (nt) on the Illumina MiSeq platform by the University of Minnesota Genomics Center (UMGC) using their dual-index method.15 Negative (sterile water) controls were included through amplification and sequencing. Sequence data are deposited in the Sequence Read Archive at NCBI under BioProject accession number SRP071855.
Sequence data were quality trimmed and processed using the mothur software Version 1.35.116 and paired-end joined using the fastq-join software.17 Data were quality trimmed with a minimum score of 35 over a 50 nt window, no ambiguous bases, ≤8 nt homopolymers, and ≤2 nt mismatches to primer sequences. High-quality sequences were aligned against the SILVA database Version 13218,19 and screened using a 2% precluster,20 and chimeras were identified and removed using UCHIME Version 4.2.40.21 Operational taxonomic units (OTUs) were clustered at 97% similarity using complete-linkage clustering, and taxonomic classifications were performed using the Ribosomal Database Project Version 16 database.22 For statistical comparisons, samples were rarefied to 15,000 sequence reads in all samples by random subsample.23 Oligotyping analyses were done to assess the distribution of potential species and strains among the genera Blautia and Bacteroides, due to significant changes in these genera among the PSC/IBD population, using recommended best practices.24
The Good’s coverage estimate, Shannon index,25 and abundance-based coverage estimate (ACE) of richness26 were calculated in mothur. Differences in beta diversity (between samples) were evaluated using Bray–Curtis dissimilarity matrices27 and visualized using ordination by principal coordinates analysis.28 Taxa associated with ordination position were determined by Spearman correlation. Differences in community composition were evaluated statistically using analysis of similarity (ANOSIM),29 and OTUs that differed significantly between study arms were determined using linear discriminant analysis of effect sizes (LEfSe).30
Statistical analysis
The average bile acid concentration over a 1-week collection period was used as a point estimate for each participant. Due to the severe right skew of the data, bile acid concentrations were log transformed for statistical testing. Individual comparisons were performed by using the Student’s t-test. Pairwise comparisons among groups were performed with Tukey–Kramer honestly significant difference (HSD). ANOVA was used for repeat measures over time. Proportions of bile acids were compared with Wilcoxon signed rank test. All statistical tests were done with α=0.05.
Results
Altered bile acid profile in PSC with IBD
Seven participants with PSC/IBD, eight participants with IBD alone, and eight healthy control individuals were recruited to the study (Table 1). Participants with PSC were older than both the IBD and healthy control groups and had a longer duration of IBD than did the IBD alone group.
  | Table 1 Baseline characteristics Abbreviations: IBD, inflammatory bowel disease; PSC, primary sclerosing cholangitis; TNF, tumor necrosis factor. |
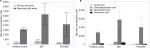
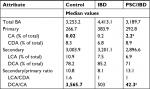
There were no significant differences between the overall fecal concentrations of total bile acids in the stool of participants with IBD (median: 4.413 mM, IQR: 2,785 mM, 10,819 mM) vs PSC/IBD (median: 3,189 mM, IQR: 554 mM, 4,075 mM), or either group compared to controls (median: 3,253 mM, IQR: 2,857 mM, 4,533 mM). Contrary to our original hypothesis, secondary bile acid concentrations in the stool of patients with PSC/IBD were less than those with IBD alone (Figure 1A). As expected, DCA was the predominant secondary bile acid in the stool among all groups, although there was no significant difference in the concentration of DCA among groups (Figure 1B). Patients with IBD demonstrated a small but significant increase in LCA compared to those with PSC/IBD (P=0.03). No other differences in the predominant bile acids were noted. The proportional composition of fecal bile acids is shown in Table 2. Overall, the proportion of primary to secondary bile acids was similar among the three groups. Relative to controls, those with PSC/IBD had a higher percent of CA, as a percentage of total fecal bile acids, and thus a lower ratio of DCA/CA (P<0.05).
When stratified by endoscopic activity, there was a decrease in the total fecal bile acid content among those with active inflammation compared to no active inflammation (P=0.03). Examination of predominant bile acids did not reveal specific changes but rather showed a global proportional decrease. Two participants had a history of low grade dysplasia. Their fecal bile acid patterns demonstrated a substantial decrease in DCA concentration, and thus of the total fecal bile acid pool, although this was not statistically significant (P=0.15).
Microbiota between PSC/IBD, IBD, and healthy controls
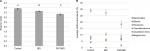
Alpha diversity was significantly different between the groups, as measured by the Shannon index (Figure 2A). Diversity was significantly higher in control and IBD patient samples than those of IBD/PSC participants. Notably, participants with PSC/IBD had a decrease in the relative abundance of Bacteroides compared to the control and IBD populations. Control, IBD, and PSC/IBD sample communities were comprised primarily Blautia, Bacteroides, Roseburia, and Faecalibacterium (Figure 2B). Megamonas had a higher relative abundance in PSC samples; however, this genus was found almost exclusively in a single participant (P<0.001). Greater relative abundances of Clostridium XIVa and Faecalibacterium were associated predominately with control samples.
Active endoscopic inflammation was associated with decreases in Bacteroides, Clostridium XlVa, and Akkermansia and with an increase in Roseburia, Faecalibacterium, Megamonas, and Alistipes (Table 3). Among those with IBD alone, active inflammation was associated with a significant increase in the abundance of Faecalibacterium and Sutterella and a decreased abundance of Akkermansia and Streptococcus. Among participants with PSC/IBD, those with active inflammation demonstrated a significant increase in the abundance of Roseburia, Megamonas, Streptococcus, Bifidobacterium, and Acidaminococcus, with a decreased abundance of Bacteroides, Clostridium XlVa, Akkermansia, and Sutterella (Table 4). Individuals with a history of low grade dysplasia had a significant decrease in the abundance of Bacteroides and an increase in the abundance of Roseburia and Alistipes (Table 5).
Clinical and bile acid response to vancomycin
Six of the seven participants with PSC/IBD were administered a 2 weeks course of oral vancomycin, 500 mg twice a day. Two of the six (33%) demonstrated at least a 10% reduction in alanine aminotransferase (ALT) levels at 2 weeks, while two of the six (33%) demonstrated at least a 10% increase in ALT. There were no substantial changes in other liver enzymes or indicators of disease activity (data not shown).
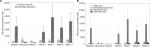
Secondary bile acids were significantly reduced in participants receiving oral vancomycin and for the first week after vancomycin treatment ceased (Figure 3A and B); these returned to baseline values by 3 weeks after vancomycin cessation (ANOVA, P=0.006). Secondary bile acid composition analysis indicated that only DCA changed significantly over time (ANOVA, P=0.0004). LCA concentrations demonstrated a non-significant decrease on vancomycin, with a somewhat delayed return to pre-vancomycin concentrations. No associations with fecal bile acid concentration and change in ALT were identified.
Microbiota changes following vancomycin
Administration of vancomycin significantly decreased alpha diversity, which recovered 3 weeks after cessation of vancomycin (Figure 4A). All predominant genera decreased in relative abundance on vancomycin (Figure 4B), apart from a single participant who developed an increase in the relative abundance of Megamonas on vancomycin. The communities recovering from vancomycin treatment (weeks 5–11) were significantly different from baseline (P<0.001). In particular, substantial differences were noted in the recovery of two genera: Blautia and Bacteroides (Figure 4C and D). OTUs that were significantly correlated to position along the x-axis were predominantly classified into the genera Bacteroides and Blautia, with PSC week 11 samples having a significantly greater relative abundance of Blautia and decrease in Bacteroides.
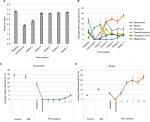
Oligotyping analysis was performed to determine the distribution of likely strains among the genera Blautia and Bacteroides (Figure 5). Communities from healthy controls had a significantly different distribution of oligotypes within these genera compared to baseline IBD or PSC/IBD communities (P<0.05). PSC/IBD communities at week 11 showed a significantly different distribution of Blautia oligotypes compared to baseline for five of the 10 most abundant oligotypes (P<0.001). Furthermore, PSC/IBD communities at week 11 had significantly different distributions of Bacteroides oligotypes compared to baseline in four of the 10 most abundant oligotypes, with two oligotypes being predominate at week 11 (P<0.05).
  | Figure 5 Relative abundances of oligotypes within the genera of (A) Blautia and (B) Bacteroides. Abbreviations: IBD, inflammatory bowel disease; PSC, primary sclerosing cholangitis. |
Microbiota and bile acid associations
Concentrations of the primary bile acids cholic acid (CA) and chenodeoxycholic acid (CDCA) were positively correlated (P<0.05) with microbial communities in 11-week PSC patients, which comprised greater abundances of Blautia (Figure 6). The secondary bile acid DCA was also positively correlated with communities comprised of greater abundances of Roseburia. Among samples collected from the entire cohort, abundances of members of the predominant genera, except Megamonas, were negatively correlated with concentrations of glyco-conjugated primary bile acids (ρ=–0.351 to –0.149, P≤0.020). Abundances of the Bacteroides, Faecalibacterium, and Clostridium XI tended to be negatively correlated with concentrations of primary bile acids and positively correlated with secondary bile acids. In contrast, abundances of Blautia were positively correlated with CA, CDCA, and Tauro-chenodeoxycholic acid (CDA) (ρ=0.151–0.204, P≤0.019) and abundances of Roseburia were positively correlated with CDCA and DCA (ρ=0.141 and 0.212, P=0.029 and 0.001).
Discussion
Contrary to our original hypothesis, patients with PSC/IBD did not have an increase in fecal secondary bile acids compared to those with IBD alone or healthy controls. We also observed slightly greater fecal LCA concentrations among individuals with IBD compared to PSC/IBD. The fecal samples in the two participants with a diagnosis of low-grade dysplasia contained minimal fecal concentrations of DCA, although the difference did not reach statistical significance. As DCA is the predominant fecal bile acid, the total bile acid pool was substantially decreased, although concentrations of other bile acids were similar to those without dysplasia. While some previous reports found an association of fecal bile acid concentration and colonic dysplasia,7 others have cast doubt on this hypothesis.31 Our results are consistent with those showing no association between an increase in fecal bile acids and colonic dysplasia.
A recent study also examined the relationship between PSC/IBD, bile acids, and the microbiota.32 Torres et al identified a reduced total bile acid pool in PSC/IBD vs IBD; however, there were no healthy controls. While we did note a trend toward lower total bile acids in PSC/IBD, overall the pattern was similar to healthy controls. Duboc et al33 demonstrated a decrease in the relative proportion of secondary bile acids in the stool of those with active IBD. We were unable to replicate this finding and in fact demonstrated a nonsignificant trend toward more secondary bile acids in the stool of participants with higher IBD activity. It is possible that this difference is related to fecal collection or bile acid analysis methods. Our study has the advantage of stratifying IBD activity endoscopically, and notably, those with endoscopic remission were indistinguishable from controls. The Duboc study was able to analyze conjugated and sulfated bile acids in more detail, which may also be responsible for the difference in findings.
Our work is consistent with prior work in patients with cirrhosis, in which the conversion from primary to secondary bile acids is reduced as liver disease progresses.34 Interestingly, conversion of CA to DCA was impaired in our cohort of PSC/IBD patients without cirrhosis or advanced cirrhosis. Primary bile acids, such as CA, bind to the farnesoid X receptor (FXR), which is linked to the immune response in IBD. In murine models of chemically induced colitis, activation of FXR by bile salts prevented inflammation and inhibited proinflammatory cytokines.35 In humans, FXR activation in the small intestine is decreased in those with Crohn’s colitis as compared to those with ulcerative colitis or healthy controls.36 Although we could not detect substantial differences in bile acid concentrations, it remains possible that small proportional differences or altered bile acid signaling pathways account for some of the unique phenotypes seen in PSC/IBD.
As expected, oral vancomycin administration dramatically reduced the concentration of secondary fecal bile acids by effectively blocking their conversion from primary bile acids. Interestingly, this effect was predominately limited to DCA with nonsignificant decrease in LCA. Overall, these changes are similar to those reported by Vrieze et al11 in a population of obese participants. The decrease in secondary bile acid production recovered within 3 weeks of cessation of oral vancomycin. This is noteworthy as vancomycin is being studied as a therapy for PSC (NCT01802073). If proven effective, oral vancomycin may be beneficial in PSC through the effect on the intestinal microbiota and subsequent secondary bile acid production. This could have implications for increasing FXR activation (a primary bile acid receptor), which may be beneficial in PSC or potentially even IBD. These effects were not sustained, and thus, it is likely that if vancomycin was to be a useful therapy, it would need to be a chronic therapy, increasing the risks of chronic antibiotic use such as bacterial resistance.
Consistent with prior literature, bacterial diversity was decreased among those with IBD compared to controls. Among those with PSC/IBD, diversity was even lower than those with IBD alone. Oral vancomycin administration substantially decreased microbial diversity and secondary bile acid metabolism, both of which recovered within three weeks of vancomycin cessation. Interestingly, the postvancomycin microbiota was assembled differently, with a relative increase in Blautia. While an increased proportion of Blautia has been observed in those with PSC in other studies,32 in our cohort, Blautia constituted a small proportion of the microbiome prior to vancomycin exposure, similar to that in healthy controls (~5% average relative abundance). In mice, increasing dietary CA substantially increases the level of bile acid 7-alpha-dehydroxylating bacteria.10 With the administration of vancomycin, Blautia expanded substantially and remained high for weeks after vancomycin administration was stopped. As primary bile acids, such as CA, are a substrate for Blautia, vancomycin likely created a favorable environment for its growth. Notably, once Blautia expanded, it remained present, apparently at the cost of the relative abundance of Bacteroides, which was substantially decreased by vancomycin administration. A relative increase in Blautia has been noted in participants with rheumatoid arthritis following vancomycin administration, and thus, this is not likely a PSC-specific effect.37
While the overall trend of decreased diversity was seen in our study, specific bacterial taxa associated with IBD were not always consistent. A recent meta-analysis of three large cohorts of IBD patients found the intestinal microbiota to be 87.5% predictive of Crohn’s disease and 79.1% of UC.38 Relative increases in the abundances of the phyla Actinobacteria and Proteobacteria (Enterobacteriaceae) and a relative decrease in Firmicutes (Clostridiales) in feces correlated with the severity of IBD. Our data show a depression of Faecalibacterium for both IBD and PSC/IBD relative to controls; however, Actinobacteria and Proteobacteria were not abundant in our samples. Additionally, we attempted to correlate a dysbiosis index to our population based on the work of Gevers et al.39 However, we were unable to match our data to the index as many taxa in the index were not abundant in our samples. This may be in part because of our use of fecal samples rather than mucosal biopsies but could also be due to differences in sequencing methods or geographic variance of our cohort. These results suggest that while certain trends in taxonomic associations with IBD may be noted in populations, the predictive power for an individual may be limited.
Our study has several limitations. This was a small pilot study and, therefore, may be underpowered to detect smaller but important differences in the bile acid composition. Another limitation was the lack of dietary control. The fecal bile acid pool is likely modified by diet. We sampled stool at least three times over 1 week to obtain an average over the week to minimize potential dietary outliers from a single meal. Finally, our population of PSC/IBD patients was older than the healthy controls or those with IBD alone, which may contribute to some differences seen between the groups.
Conclusion
Those with PSC/IBD had a more pronounced dysbiosis characterized by decreased bacterial diversity and decreased relative abundance of Bacteroides. In contrast, there were no major differences between the fecal bile acid patterns of those with IBD, PSC/IBD, and healthy controls. Vancomycin administration to PSC/IBD participants decreases the fecal bile acid pool, likely through the inhibition of secondary bile acid production, and primarily through inhibition of DCA. While the bile acid composition recovered 1 month after the completion of a 2-week course of vancomycin, changes in the intestinal microbiota persisted, favoring an increase in Blautia. Future studies in PSC should determine how the microbiota and/or antibiotics affect bile acid signaling and ultimately intestinal and hepatic inflammation.
Acknowledgments
Research reported in this publication was supported by the National Center for Advancing Translational Sciences of the National Institutes of Health Award Number UL1TR000114. The content is solely the responsibility of the authors and does not necessarily represent the official views of the National Institutes of Health.
Author contributions
BPV contributed to the design, analysis, interpretation of data, and manuscript drafting. TK, CS, MJH, and SS contributed to the analysis, results, interpretation of data, and critical review of the manuscript. JR contributed to the design, collection of data, and critical review of the manuscript. CG and Amanda K contributed to design, collection of data, and critical review of the manuscript. MJS and Alexander K contributed to the design, interpretation of data, and critical review of the manuscript. All authors contributed to data analysis, drafting and revising the article, gave final approval of the version to be published, and agree to be accountable for all aspects of the work.
Disclosure
BPV has received research support from Takeda, Celgene, Roche, Crestovo, and Diasorin and has received speaking/consulting income from Abbvie and Janssen. MJS has consulted for Crestovo. The authors report no other conflicts of interest in this work.
References
Abraham C, Cho JH. Inflammatory bowel disease. N Engl J Med. 2009;361(21):2066–2078. | ||
Bambha K, Kim WR, Talwalkar J, et al. Incidence, clinical spectrum, and outcomes of primary sclerosing cholangitis in a United States community. Gastroenterology. 2003;125(5):1364–1369. | ||
Loftus EV, Harewood GC, Loftus CG, et al. PSC-IBD: a unique form of inflammatory bowel disease associated with primary sclerosing cholangitis. Gut. 2005;54(1):91–96. | ||
Torres J, Pineton de Chambrun G, Itzkowitz S, Sachar DB, Colombel JF. Review article: colorectal neoplasia in patients with primary sclerosing cholangitis and inflammatory bowel disease. Aliment Pharmacol Ther. 2011;34(5):497–508. | ||
Bernstein C, Holubec H, Bhattacharyya AK, et al. Carcinogenicity of deoxycholate, a secondary bile acid. Arch Toxicol. 2011;85(8):863–871. | ||
Ajouz H, Mukherji D, Shamseddine A. Secondary bile acids: an underrecognized cause of colon cancer. World J Surg Oncol. 2014;12(1):164. | ||
Hill MJ, Lennard-Jones JE, Melville DM, Neale K, Ritchie J. Faecal bile acids, dysplasia, and carcinoma in ulcerative colitis. The Lancet. 1987;330(8552):185–186. | ||
Duboc H, Taché Y, Hofmann AF. The bile acid TGR5 membrane receptor: from basic research to clinical application. Dig Liver Dis. 2014;46(4):302–312. | ||
Torres J, Bao X, Goel A, et al. The features of mucosa-associated microbiota in primary sclerosing cholangitis. Aliment Pharmacol Ther. 2016;43(7):790–801. | ||
Ridlon JM, Alves JM, Hylemon PB, Bajaj JS. Cirrhosis, bile acids and gut microbiota: unraveling a complex relationship. Gut Microbes. 2013;4(5):382–387. | ||
Vrieze A, Out C, Fuentes S, et al. Impact of oral vancomycin on gut microbiota, bile acid metabolism, and insulin sensitivity. J Hepatol. 2014;60(4):824–831. | ||
Tabibian JH, Weeding E, Jorgensen RA, et al. Randomised clinical trial: vancomycin or metronidazole in patients with primary sclerosing cholangitis - a pilot study. Aliment Pharmacol Ther. 2013;37(6):604–612. | ||
Weingarden AR, Chen C, Bobr A, et al. Microbiota transplantation restores normal fecal bile acid composition in recurrent Clostridium difficile infection. Am J Physiol Gastrointest Liver Physiol. 2014;306(4):G310–G319. | ||
Claesson MJ, Wang Q, O’Sullivan O, et al. Comparison of two next-generation sequencing technologies for resolving highly complex microbiota composition using tandem variable 16S rRNA gene regions. Nucleic Acids Res. 2010;38(22):e200. | ||
Gohl DM, Vangay P, Garbe J, et al. Systematic improvement of amplicon marker gene methods for increased accuracy in microbiome studies. Nat Biotechnol. 2016;34(9):942–949. | ||
Schloss PD, Westcott SL, Ryabin T, et al. Introducing mothur: open-source, platform-independent, community-supported software for describing and comparing microbial communities. Appl Environ Microbiol. 2009;75(23):7537–7541. | ||
Aronesty E. Comparison of sequencing utility programs. Open Bioinforma J. 2013;7(1):1–8. | ||
Pruesse E, Quast C, Knittel K, et al. SILVA: a comprehensive online resource for quality checked and aligned ribosomal RNA sequence data compatible with ARB. Nucleic Acids Res. 2007;35(21):7188–7196. | ||
Quast C, Pruesse E, Yilmaz P, et al. The SILVA ribosomal RNA gene database project: improved data processing and web-based tools. Nucleic Acids Res. 2013;41(Database issue):D590–D596. | ||
Huse SM, Welch DM, Morrison HG, Sogin ML. Ironing out the wrinkles in the rare biosphere through improved OTU clustering. Environ Microbiol. 2010;12(7):1889–1898. | ||
Edgar RC, Haas BJ, Clemente JC, Quince C, Knight R. UCHIME improves sensitivity and speed of chimera detection. Bioinformatics. 2011;27(16):2194–2200. | ||
Cole JR, Wang Q, Cardenas E, et al. The Ribosomal Database Project: improved alignments and new tools for rRNA analysis. Nucleic Acids Res. 2009;37(Database issue):D141–D145. | ||
Gihring TM, Green SJ, Schadt CW. Massively parallel rRNA gene sequencing exacerbates the potential for biased community diversity comparisons due to variable library sizes. Environ Microbiol. 2012;14(2):285–290. | ||
Eren AM, Maignien L, Sul WJ, et al. Oligotyping: differentiating between closely related microbial taxa using 16S rRNA gene data. Methods Ecol Evol. 2013;4(12):1111–1119. | ||
Shannon CE. A mathematical theory of communication. SIGMOBILE Mob Comput Commun Rev. 2001;5(1):3–55. | ||
Chao A, Lee S-M. Estimating the Number of Classes via Sample Coverage. J Am Stat Assoc. 1992;87(417):210–217. | ||
Bray JR, Curtis JT. An ordination of the upland forest communities of southern wisconsin. Ecol Monogr. 1957;27(4):325–349. | ||
Anderson MJ, Willis TJ. Canonical analysis of principal coordinates: a useful method of constrained ordination for ecology. Ecology. 2003;84(2):511–525. | ||
Clarke KR. Non-parametric multivariate analyses of changes in community structure. Austral Ecology. 1993;18(1):117–143. | ||
Segata N, Izard J, Waldron L, et al. Metagenomic biomarker discovery and explanation. Genome Biol. 2011;12(6):R60. | ||
Mudd DG, Mckelvey ST, Norwood W, Elmore DT, Roy AD. Faecal bile acid concentration of patients with carcinoma or increased risk of carcinoma in the large bowel. Gut. 1980;21(7):587–590. | ||
Torres J, Palmela C, Brito H, et al. The gut microbiota, bile acids and their correlation in primary sclerosing cholangitis associated with inflammatory bowel disease. United European Gastroenterol J. 2018;6(1):112–122. | ||
Duboc H, Rajca S, Rainteau D, et al. Connecting dysbiosis, bile-acid dysmetabolism and gut inflammation in inflammatory bowel diseases. Gut. 2013;62(4):531–539. | ||
Kakiyama G, Pandak WM, Gillevet PM, et al. Modulation of the fecal bile acid profile by gut microbiota in cirrhosis. J Hepatol. 2013;58(5):949–955. | ||
Gadaleta RM, van Erpecum KJ, Oldenburg B, et al. Farnesoid X receptor activation inhibits inflammation and preserves the intestinal barrier in inflammatory bowel disease. Gut. 2011;60(4):463–472. | ||
Nijmeijer RM, Gadaleta RM, van Mil SW, et al. Farnesoid X receptor (FXR) activation and FXR genetic variation in inflammatory bowel disease. PLoS One. 2011;6(8):e23745. | ||
Isaac S, Scher JU, Djukovic A, et al. Short- and long-term effects of oral vancomycin on the human intestinal microbiota. J Antimicrob Chemother. 2017;72(1):128–136. | ||
Zhou Y, Xu ZZ, He Y, et al. Gut microbiota offers universal biomarkers across ethnicity in inflammatory bowel disease diagnosis and infliximab response prediction. mSystems. 2018;3(1):e00188-17. | ||
Gevers D, Kugathasan S, Denson LA, et al. The treatment-naive microbiome in new-onset Crohn’s disease. Cell Host Microbe. 2014;15(3):382–392. |
 © 2019 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.
© 2019 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.