Back to Journals » Pharmacogenomics and Personalized Medicine » Volume 13
A Pharmacogenomic Dissection of a Rosuvastatin-Induced Rhabdomyolysis Case Evokes the Polygenic Nature of Adverse Drug Reactions
Authors Calderon-Ospina CA , Hernández-Sómerson M, García AM, Mejia A, Tamayo-Agudelo C, Laissue P , Fonseca Mendoza DJ
Received 26 August 2019
Accepted for publication 28 November 2019
Published 2 March 2020 Volume 2020:13 Pages 59—70
DOI https://doi.org/10.2147/PGPM.S228709
Checked for plagiarism Yes
Review by Single anonymous peer review
Peer reviewer comments 2
Editor who approved publication: Dr Martin Bluth
Carlos Alberto Calderon-Ospina,1,* Mario Hernández-Sómerson,2,* Ana María García,1 Adriana Mejia,1 Caroll Tamayo-Agudelo,1 Paul Laissue,1 Dora Janeth Fonseca Mendoza1
1Center for Research in Genetics and Genomics-CIGGUR, GENIUROS Research Group, School of Medicine and Health Sciences. Universidad Del Rosario, Bogotá, Colombia; 2Medical Clinic Service, Hospital Universitario Mayor Méderi-Universidad Del Rosario, Bogotá, Colombia
*These authors contributed equally to this work
Correspondence: Dora Janeth Fonseca Mendoza
Universidad Del Rosario, Cra 24 N° 63c-69, Bogotá, Colombia
Tel +5712920200
Fax +571 3499390
Email [email protected]
Abstract: Rosuvastatin, is a widely-used statin for the treatment of hypercholesterolemia and the prevention of cardiovascular diseases. Although rosuvastatin is well tolerated, about 3/10.000 patients can suffer severe myopathy. Rhabdomyolysis is a severe medical condition that causes injury to the skeletal muscle, electrolyte imbalances, acute renal failure and extreme creatine kinase (CK) elevation. Little is known regarding the molecular involvement of rosuvastatin-induced rhabdomyolysis (RIR). It has been demonstrated that genomic variants associated with decreased enzymatic activity of proteins are important determinants in plasmatic and skeletal muscle distribution of rosuvastatin and its toxicity. Until now, no interactions of ticagrelor, ezetimibe and rosuvastatin have been described with the consideration of pharmacogenomics predisposition. The present report involves a whole-exome sequencing (WES), in a patient affected by rosuvastatin-induced rhabdomyolysis. A pharmacogenomic dissection was performed by analyzing a comprehensive subset of candidate genes (n=160) potentially related to RIR. The genes were selected according to their implication in drug metabolism or inherited myopathies. Using an innovative approach of bioinformatics analysis, considering rare and common variants, we identified 19 genomic variations potentially related to the pharmacokinetic/pharmacodynamic modifications of rosuvastatin, ezetimibe and ticagrelor. The affected genes are involved in Phase I metabolism (CYP2C19, CYP2E1, CYP1A1, CYP2D6 and CYP2C9), Phase II metabolism (UGT2B15 and UGT2B7), influx transportation (SLCO1B3 and SLCO2B1), efflux transportation (ABCG8, ABCB11, ABCC4 and ABCB1), drug targeting (NPC1L1) and inherited myopathy etiology (OBSCN). We report three rare, potentially pathogenic molecular variants in CYP2C19, NPC1L1 and OBSCN genes. Pharmacogenetic analysis indicated that the patient was a carrier of inactivating alleles in several pharmacogenes involved in drug toxicity. The whole-exome sequencing and bioinformatics analysis presented here represents an innovative way to identify genomic variants contributing with RIR´s origin and evokes the polygenic nature of adverse drug reactions.
Keywords: pharmacogenomics, rhabdomyolysis, rosuvastatin, adverse drug reaction, whole-exome sequencing, polymorphisms
Introduction
Rhabdomyolysis is a clinical emergency characterized by skeletal muscle damage resulting in the release of intracellular muscle components into the bloodstream and extracellular space.1,2 Clinically, this disease is associated with elevated CK levels, electrolyte imbalances, acute renal failure, and disseminated intravascular coagulation.3 Some of these cases result from a secondary adverse reaction (ADR) to the ingestion of medications including antihistamines, antipsychotics, lipid-lowering medications, and statins.2 Statin-induced myopathy (SIM) affects 10–25% of individuals treated, and includes myopathy, myalgia, and myonecrosis.4 The most severe form of these myopathies, rhabdomyolysis, occurs in 0.1% of patients.5 Dual ticagrelor/rosuvastatin therapy, which is generally safe and well tolerated, is used for acute coronary syndrome (ACS) treatment. However, at least 8 cases of rhabdomyolysis secondary to rosuvastatin and ticagrelor have been reported in the literature and in the World Health Organization’s Adverse Drug Reactions Database (VigiBase).6 The molecular pathophysiology of rhabdomyolysis is complex, possibly involving numerous genes acting in several molecular overlapping cascades.
Herein, to identify gene variants potentially related to statin and myotoxicity, we performed whole-exome sequencing and an innovative downstream bioinformatics analysis in a Colombian patient affected by RIR. WES is a large-scale DNA sequencing technology with potential relevance in identifying genomic variations related to drug response.7 By using in silico analyses, that included the identification of rare and common genomic variants in a subset of 160 candidate genes, we identified 19 genomic variants potentially related to the RIR. We found three rare, potentially pathogenic molecular variants (in terms of the In silico prediction and the conservation of the residue during evolution) in CYP2C19, NPC1L1 and OBSCN genes. We suggest that the rare variants and single nucleotide variants (SNVs) in these pharmacogenes can contributed to the myotoxicity observed in our patient.
We estimate that advanced computational analyses, large-scale DNA analysis and clinical information likely enhances drug phenotype predictions and highlights its use in personalized medicine.
Case Presentation
A 65-year-old woman admitted to the emergency room, presented a ten-day history of weakness and generalized musculoskeletal pain predominantly in the lower limbs. Additionally, she presented colicky abdominal pain associated with distension and the presence of dark urine.
The patient presented a medical history of stage IV chronic kidney disease, dilated cardiomyopathy of ischemic origin with 38% LVEF, type 2 diabetes mellitus, hypertension, and multivessel coronary disease with revascularization eight months prior. She was being treated with losartan, carvedilol, linagliptin, insulin detemir, rosuvastatin + ezetimibe 40mg/10mg daily for 2 years, aspirin and ticagrelor 90mg every 12 hrs since her intervention eight months ago.
Physical examination revealed the patient to be dehydrated, hypotensive, tachycardic, with palpebral edema and abdominal distension.
Paraclinical tests reported metabolic acidosis, increased transaminases and a euthyroid profile, leukocytosis with left shift, hyponatremia (122mmol/L), hyperkalemia (6.7mmol/L), hypocalcemia (7.8mmol/L), hyperphosphatemia (14mmol/L), and elevated nitrates (creatinine 9.6mg/dl and BUN 162mg/dl). A urinalysis with urine culture accompanied by a renal ultrasound confirmed the pyelonephritis diagnosis. The patient was taken to hemodialysis and anti-hyperkalemic treatment was initiated.
On the second day of admission, a decrease in nitrate levels and improvement of electrolyte levels was evidenced; however, the patient persisted with pain in the lower limbs, associated with a total CPK level of 38,987 UI/L, more than 300 times higher than the reference values (26-192UI/L). A 12-lead EKG did not show changes in acute myocardial ischemia, ruling out an acute coronary syndrome; thereafter, rosuvastatin and ezetimibe were suspended.
An electromyography was performed with nerve conduction velocities in four extremities, which resulted compatible with intrinsic muscle fiber compromise suggestive of rhabdomyolysis, later a skeletal muscle biopsy confirmed the diagnosis.
A rapid urine test confirmed the existence of myoglobinuria. On the eighth day of hospitalization, a marked decrease in CPK was observed (14,000 UI/L). The patient was discharged 2 months after admission with a final diagnosis of acute kidney injury KDIGO score of 3, secondary to a statin induced-rhabdomyolysis.
Molecular Analysis
For WES, a total amount of 1.0 μg genomic DNA per sample was used as input material for the DNA library preparation. Sequencing libraries were generated using Agilent Sure Select Human All Exon kit (Agilent Technologies, CA,USA). Fragmentation was carried out to generate 180–280 bp fragments. After adenylation of 3ʹ ends of DNA fragments, adapter oligonucleotides were ligated and selectively enriched in a PCR reaction. After PCR reaction, library hybridize with liquid phase with biotin labeled probe, then use magentic beads with streptomycin to capture the exons. Captured libraries were enriched in a PCR reaction to add index tags to prepare for hybridization. Products were purified using AMPure XP system (Beckman Coluter, Berverly, USA) and quantified on the Agilent Bioanalyzer 2100 system. Libraries were sequenced on HiSeq Ilumina platform, paired-end 150 pb. The short reads (Raw Data) were analyzed in FASTQ format. The rew reads were aligned to the reference HG18 with Burrows-Wheeler Aligner (BWA) software. The variants were identified with SAMtools. Data quality guaranteed that >80% of bases have a sequencing quality score >Q30 and minimum average coverage of 30X. Following genomic variant detection, the variants were annoted using the tool ANNOVAR to identify affected genomic regions, protein coding changes, allele frequencies reported by Genome Aggregation Database (gnomAD) and In house database-WES of Colombian people and to predict the deleteriousness of mutations (SIFT and,PolyPhen). Library preparation and sequencing were carried out at Novogene (Beijing-China).
A subset of 160 candidate genes was created (Table 1). The genes were selected according to their implication in inherited myopathies (70-Rhab) or pharmacological metabolism (90-Rhab) in statins and the drugs administrated to our patient. Candidate genes for 70-Rhab were selected through a review of literature and a public database (Online Mendelian Inheritance in Man (OMIM). The pharmacogenes (90-Rhab) were identified by their action on the metabolism of the medications (targets, enzymes and transporters). This information was subtracted from a literature review and public databases (https://www.drugbank.ca/ and https://www.pharmgkb.org/).
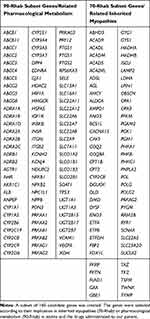 |
Table 1 Genes Analyzed |
In the 70-Rhab and 90-Rhab subsets, sequence variants with potential deleterious effect (missense, nonsense, splice site, frameshift), and MAF <1% were selected for analysis. We used SIFT and PolyPhen2 software to predict whether an amino acid substitution affects protein function. Sequences of proteins with altered amino acids were compared with orthologous of some mammalian species using available public database sequences (https://www.uniprot.org/uniprot/). We considered a variation relevant if residue was conserved during evolution. For splice site variants, we performed in silico prediction of potential pathogenicity using Human Splicing Finder software (http://www.umd.be/HSF/). Additionally, for 90-Rhab genes subset we included common genomic variants previously described for their clinical involvement in drug metabolism. Using STRING v.11.0 interaction database, we identified possible interactions (PPIs) between proteins potentially altered for molecular variants (confidence score > 0.400) (https://string-db.org/cgi/network.pl).
The Ethics Committee of Universidad del Rosario approved this study and it was conducted according to the principles of the Helsinki Declaration (institutional review board reference CS/ABN062). Written informed consent was obtained from the patient for DNA analysis and publication of the case report.
Results
Pharmacogenomic Analysis
The descriptions of the molecular variants for the subsets 70-Rhab and 90-Rhab are shown in Table 2. In the 90-Rhab subset of genes a total of 18 genomic variants were identified: 7 in genes of phase I metabolism; 2 in phase II metabolism genes; 3 in the Influx transporters; 5 in the efflux and 1 in the target of ezetimibe. Regarding subset 70-Rhab, a variant in the OBSCN gene was identified. Three potentially pathogenic variants with very low MAF (genomAD and In House WES) were found CYP2C19 (c.394C> T, MAF= 0.00044 and 0.001), NPC1L1 (c.1581-6C> G, MAF= 0.00017 and 0) and OBSCN (c.35T> A, MAF= 0.0033 and 0). For CYP2C19 and OBSCN, protein alignments revealed that the Arginine residue at position 132 and the Phenylalanine residue at position 12 are conserved among mammalian species (Figure 1). Regarding NCP1L1 c.1581-6G> C variant, In Silico predictions indicate that it affects splicing due to the alteration of the acceptor site. Multilevel analysis of genes involved in possible mechanisms of Rhabdomyolysis are shown in Figure 2. Protein-protein interaction data retrieved from STRING v11.0 is shown in Figure 3. Mainly PPIs predicted in our network were co-expressed. For phase I metabolism enzymes (CYPs and UGTs) more PPIs were evident (Figure 3). For network protein, PPI enrichment p-value determined by STRING v.11.0 was <1.0e-16.
 |
Table 2 Pharmacogenomic Profile Patient. Description of the Molecular Variants for the Subsets 70 Rhab and 90 Rhab |
Discussion
We presented a case report that involved WES in a patient affected by rhabdomyolysis. To date, molecular involvement exploration in this phenotype has been limited to a few SNV´s in genes related to drug metabolism in phase I and II.8 Due to the molecular complexity of rhabdomyolysis pathophysiology, we focused our study on a subset of 160 genes potentially related to phenotype. The genes proposed are relevant in the myopathies susceptibility. We conducted a large-scale DNA sequencing approach using two bioinformatics analysis: a) stringent filters that enable the identification of rare variants with potentially strong functional effects and b) filtering of frequent genomics variants reported in literature. This approach led us to identify 19 genomic variants in 14 genes, three of which are rare (MAF<0.3%), and potentially pathogenic (CYP2C19, NCP1L1 and OSBCN). These findings corroborate that our approach with selected groups of genes is a powerful strategy for molecular dissection of severe ADR and evokes the polygenic nature of myotoxicity. Successful studies have been made by our research group using WES to identified candidate genes of complex diseases and now in pharmacogenetics.9–13
According to the Naranjo scale, it was estimated for the patient analyzed that the rhabdomyolysis was induced by rosuvastatin (8 points “probable”).14,15 Considering the timeline of our patient’s medication use (Rosuvastatin-ezetimibe treatment had been established two years prior the reaction), we estimate that exposure to ticagrelor triggered the rhabdomyolysis. The Horn scale indicated as probable (5 points) the implication of this pharmacological interaction in the ADR observed in our patient. Despite these considerations, it is important to recognize as a limiting factor of our study that, as with any adverse reaction, it is difficult to attribute causality to a specific drug even with the use of expert judgment, algorithms or Bayesian/probabilistic approaches.16
Among the medications used by the patient, a potential moderate drug-drug interaction between rosuvastatin and ezetimibe has been identified. It has been documented that co-administration of ezetimibe and statins leads to a potential risk of myopathy and elevated transaminase levels. It has been suggest that the combination of statins and ezetimibe should be performed with caution and that these medications should be discontinued if myopathy is suspected or diagnosed.17 The reduction in CK (from 38,917 to 14,000 IU/L) in our patient after rosuvastatin-ezetimibe suspension supports this hypothesis. Even so, we must recognize that there may be unknown drug-drug interactions with statins that are potentially related to the increased risk of myopathies and that may influence the accurate identification of drug-gene interactions.18
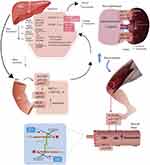
To date, 8 cases of rhabdomyolysis induced by rosuvastatin and ticagrelor have been reported.6 We performed a molecular multilevel analysis according to the mechanisms involved in the potential influence of ticagrelor on the elimination/disposal of rosuvastatin: a) kidney damage caused by ticagrelor, b) Genetic polymorphisms in drug metabolizing enzymes and c) decreased biliary and renal excretion of rosuvastatin secondary to transporter competition.6 This analysis allowed us to hypothesize the action of the genetic polymorphism in the final disposition of rosuvastatin, involved in the generation of rhabdomyolysis in our patient (Figure 2).
Kidney damage is an ADR frequently associated with ticagrelor; in the PLATO trial (The Platelet Inhibition and Patient Outcomes) it was determined that the concentration of serum creatinine increased by more than 30% in patients receiving ticagrelor, being the pre-existing kidney impairment an important factor in the outcome.19 This constitutes a determining risk factor in our patient who at the time of the administration of ticagrelor was suffering from stage IV chronic renal failure and whose decompensation was probably triggered by the ingestion of the antiplatelet drug after the revascularization. The patient reported a creatinine value of 9.2 mg/dl (above the normal value). The mechanism explaining the worsening of renal function with the use of ticagrelor is unknown but it has been proposed that inhibition of adenosine reuptake by the medication reduces the glomerular filtration rate of the afferent renal arteriole.20 In the clinical management of patients with renal failure, the dose of rosuvastatin must be adjusted and high doses as prescribed in our patient (40mg daily) is contraindicated. It is estimated that the plasma concentration of rosuvastatin may increase three times in patients with severe kidney injury.19 We estimate that in the reported case the deterioration of renal function due to ticagrelor and the dose of rosuvastatin and ezetimibe were determinants in the increase of the serum concentration of rosuvastatin related to rhabdomyolysis.
The risk of myotoxicity from rosuvastatin is dependent on the plasma concentration of the medication in the blood and myocytes. Through WES, we identified in our patient, 19 genetic variations affecting drug pharmacokinetics and pharmacodynamics. Regarding the enzymes of phase I metabolism, the patient carried alleles that confer a potential variation in normal metabolism phenotype: (CYP2C9*3 (c.A1075C), CYP2C19 (c.991G>A), CYP2C19 (c.394C>T), CYP2D6 (c.271C>A), CYP2D6*10 (c.100C>T) and CYP2D6*4 (c.506-1G>A). Although only 10% of rosuvastatin is metabolized by the enzymes CYP2C9 and CPY2C19 the presence of variants related to low enzyme activity compromise the drug’s disposal and predisposes its potential toxicity.21 A rare variant CYP2C19 -c.394C>T was identified in the patient, the in silico analysis demonstrates its pathogenicity, and the conservation of the amino acid in mammal species suggests a role in its biological function (Figure 1). The metabolism of carvedilol is fundamentally affected by the alleles of CYP2D6 and CYP2C9, for the described case, the decrease in enzymatic activity conferred by CYP2D6*4, CYP2D6*10, and CYP2C9*3 estimates a potential increase in the plasma concentration of R(+) carvedilol and a potential therapeutic failure in the treatment of cardiac failure.22 This observation evokes that despite being treated with carvedilol, the patient suffered a cardiac complication that required the use of ticagrelor, the main trigger of rhabdomyolysis. With respect to enzymes associated to phase II metabolism, variants UGT2B7*2 and UGT2B15*2, related to the impact of rosuvastatin lactonization and ezetimibe glucuronidation, respectively, were identified. The lactones in relation to the acid form of statins are stronger inducers of toxicity. UGT2B7*2 polymorphism can modulate lactone induction by glucuronidation and contribute to rosuvastatin toxicity.6 Ezetimibe is metabolized primarily by a glucuronidation reaction by UGT1A1, 1A3 and 2B15 to form its main phenolic metabolite, ezetimibe-glucuronide.The allele UGT2B15*2 has an enzymatic activity 5 times lower than the wild type allele which results in a decreased formation of active ezetimibe glucuronides.23 From this perspective, it can be assumed that the reduction of ezetimibe-glucuronide may potentially influence the pharmacological effect of the medication.24 The lipid-lowering action of ezetimibe may additionally be affected by the presence of a potentially pathogenic variant in NPC1L1- c.1581-6C>G, the target of the medication. This is a rare variant (MAF: 0.00017-genomAD and 0-In house WES-database) and the In Silico prediction indicates that it affects splicing due to the alteration of the acceptor site. Although only a functional study (e.g.minigenes) would establish with certainty the implication of the mutation, the findings allow us to hypothesize that the therapeutic effect of ezetimibe in the patient may be affected. The need to compensate for the desired lipid-lowering effect may be related to the prescription of a high unsafe dose of rosuvastatin (40 mg) in the patient, with the potential risk of elevated plasma concentration and development of rhabdomyolysis. Ezetimibe is a potential additional risk factor for our patient, as it has been established that this medication can induce myalgia in statin-intolerant patients.25 Although the associated mechanism is unknown, a pharmacokinetic interaction between ezetimibe and rosuvastatin has recently been identified, and the co-administration of the medications increases systemic exposure to free ezetimibe, which may favor its absorption by the gastrointestinal tract for its muscle interaction and subsequent induction of muscular ADR.25,26
Other mechanism related to rosuvastatin-ticagrelor associated myopathy involved drug transporters8. All statins are substrates of membrane transporters that play an important role in the disposition of the drug. Previous studies have identified that simple nucleotide polymorphisms (SNPs) in the SLCO1B1 gene (c.388A>G and c.521T>C) have a functional effect on OATP1B1.27 Specifically, the c.388A>G polymorphism (SLCO1B1*1B) is associated with high enzymatic activity and low plasma concentrations for OATP1B1 substrates. On the contrary c.521T>C (SLCO1B1*5) reduces the activity of the transporter and increases the plasma concentration of substrates.28 The effect of polymorphisms depends on the satin used. SNP SLCO1B1-C.521T>C has been associated with simvastatin (4-fold increase in the risk of myopathy) but not with other statins.29,30 This finding is consistent with the fact that the polymorphism fundamentally affects simvastatin since it generates an area under the curve (AUC) 221% higher in patients with the c.521CC genotype in relation to the c.521TT genotype. However, the effect is much less in atorvastatin (AUC increase of 173%) and almost non-existent for other statins.31 Additionally, the SLCO1B1*1B/SLCO1B1*1B genotype has been associated with a reduction of up to 35% in pravastatin AUC, but does not affect the pharmacokinetics of rosuvastatin.32 Our patient was not a carrier of risk alleles SLCO1B1-c.521T>C or c.388A>G or any genetic variant potentially related to myopathy. SLCO1B1 is expressed in the basolateral membrane of human hepatocytes, in where, additionally, SLCO1B3 and SLCO2B1 are expressed in similar amounts.33 The differences in the effect of SLCO1B1 polymorphisms on the pharmacokinetics of individual statins can be partially explained by the contribution of other OATPs in their hepatic uptake.31 Rosuvastatin transport is mediated by SLCO1B1, 1B3 and 2B1. The interindividual variability in patients drug response can be explained by SLCO1B3- c.T334G, c.G699A and SLCO2B1- c.935G>A,c.1175T>C polymorphisms. SLCO1B3 polymorphisms are in strong linkage disequilibrium and are related to a decreased activity, increased toxicity and increased plasma levels of their substrates.34 The functional impact of SLCO2B1-c.935G>A has been associated with reduced plasma levels of montelukast, a leukotriene receptor antagonist.35 SLCO2B1-c.1175T>C has been associated with a reduction in the activity of substrates such as estrone sulfate, as well as a reduction in the area under the fexofenadine curve.36 Knauer et al, identified the expression of SLCO2B1 in the sarcolemma membrane of skeletal muscle, which determines that control in rosuvastatin concentration is not performed solely by transporters located in the liver and intestine. It is estimated that in addition to the high plasma concentration of rosuvastatin, the concentration of the medication in the muscle fiber has an impact on the risk of rhabdomyolysis.37 Regarding statin efflux transporters, the expression of ABCC1, ABCC4, and ABCC5 in skeletal muscle has been demonstrated. These transporters are expressed in the sarcolemma membrane of muscle fibers and perform a protective role in the intracellular accumulation of statins.38 In the analyzed patient the variant ABCC4-c.912G>T was identified, located in the splice acceptor site 3ʹ of the pre-mRNA of exon 8. Functional studies have indicated that it leads to the elimination of about 300bp in exon 8 and as a consequence the protein exhibits low enzymatic activity.39 Although ABCB1 (P-pg) has not been associated with rosuvastatin toxicity, the implication of the allele ABCB1-c.2677T>G has been associated with the increase of muscle toxicity and myalgia from simvastatin.40 The dynamic interplay between influx and efflux transporter activities likely modified muscle fiber rosuvastatin concentrations, which determines susceptibility to muscular ADR such as Rhabdomyolysis.
Integrating our findings, we suggest that the total effect of altered proteins supports the hypothesis that the risk of myotoxicity from rosuvastatin is dependent on the concentration of the medication.41 Our results of PPIs using STRING database corroborate that the proteins analyzed interact with each other and are co-expressed. The PPI enrichment p-value predicted (<1.0e-16), indicated that our network proteins have more interactions than expected which implies that the proteins are biologically connected, as a group.42
Finally, we also suggest one new mechanism related to the potencial rol of OBSCN in establishing the sarcomere-sarcoplasmic reticulum (SR) connection, which could have increased the risk of Rhabdomyolysis.43 Ablation of obscurin in mice results in alterations in the SR architecture with marked sarcolemma fragility.44 OBSCN-c.35T>A can potentially affect the interaction with SR-associated proteins such as Titin and myomesin and facilitate the fragility. The interaction between these proteins involve the N-terminal end of the OBSCN, so we estimate that genetic variants in other positions are not potentially related to the establishment of the sarcomere-sarcoplasmic reticulum (SR) connection. Muscle biopsies from patients with stain-induced myopathy have shown structural changes in the SR.45
Despite the considerations made of the potential impact of the rare variants identified in our patient (CYP2C9 c.394C>T; NPC1L1 c.1581-6C>G and OBSCN c.35T>A), in terms of the In Silico prediction (according to the case) and the conservation of the residue involved during the evolution, we recognize our limitation regarding the possibility of carrying out association analysis of rare variants in biologically plausible metabolizing enzymes or transporter genes that could have led to a large increase in drug exposure. Advances in large-scale DNA sequencing technology, such as WES, have allowed us to explore the contribution of rare or low-frequency variants in human phenotypes and adverse drug reactions.18,46
The analysis of GWAS on common variants, has permitted the establishment of a significant number of gene and phenotype associations, many of them with low effects and explains a small proportion of heritability. The analysis of rare variants could potentially explain more significant effects than common SNPs.47 Since many rare independent variants are found in the genome in relation to common variants, several statistical methods (gene-burden scores, variance- component test, combined test and others) have been proposed to increase power in association studies by mixing information of multiple rare variants with a functional genomic unit (e.g. gene or exon).46 Recently, Floyd et al reported a multi-site whole-exome sequencing study to investigate whether rare coding variants confer an increased risk of statin-related myopathy (SMR),included rhabdomyolisis.18 The analysis sample included 505 cases that received treatment with statins at the onset time of muscle injury symptoms and 2047 controls from North America and the United Kingdom. Rare variants (MAF<1%) identified by WES were collapsed into gene-burden scores and tested for association with the risk of SMR. Through large-scale DNA analysis, joint variant calling and statistical methods, the authors did not identify new variants associated with SMR. Similar to other reports, the study revealed the association of SNP SLCO1B1-c.521T>C with a 4-fold increased risk of severe SMR, while in mild forms of SMR this association was not evidenced.18 Nevordola et al conducted a WES study in 88 patients with statin-associated myopathy and assessed the burden of rare variants using candidate-gene and exome-wide association analysis. The strategy allowed the identification of a new gene potentially related to the phenotype (CLCN1) and case-specific pathogenic variants in MYOT,CYP3A5,SH3TC2,FBXO32 and RMB20.48 These results, initially discrepant, may be due to the difference in the phenotypes analyzed in both studies (mostly severe SMR in one and mild SMR in the other) and support the potential role of rare variants in susceptibility to stain myopathy. As hypothesized by Floy, it is possible that there are rare variants associated with SMR in non-coding regions that escape the WES analysis.18
Taken together, our findings reveal the potential relevance of WES as a comprehensive pharmacogenomics approach to identify common and/or rare variants potentially related to SMR. We estimate that in vitro functional studies can validate the implication of rare case-specific variants such as the one reported in this study. Despite the uncertainly in the incorporation of WES into the health system, we estimate that the integration of WES data likely enhances future drug dosing. Even so, it must be carefully analyzed for the potential involvement of rare variants in severe SMR phenotypes.
Conclusion
This is the first case in which whole-exome sequencing of an affected patient is performed to explain the genetic susceptibility to rhabdomyolysis induced by ticagrelor, rosuvastatin and ezetimibe. The patient was at high risk for ADR, being an elderly female, with an excessive dose of rosuvastatin and renal impairment. Our molecular findings argue in favor of the polygenic nature of rhabdomyolysis where the accumulation of rosuvastatin due to metabolism impairment, the alteration in transport proteins (SLCO1B3, SLCO2B1, ABCG8, ABCB11, ABCC4 and ABCB1), and in drug target (NCP1L1), are potential effectors in the analyzed myotoxicity. We proposed for the first time the implication of the susceptibility gene (OBSCN) in phenotype.
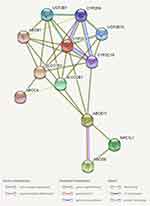 |
Figure 3 Protein-protein interaction. Note: Data retrieved from STRING v.11.0 predicts protein-protein interaction (PPIs). |
Abbreviations
RIR, rosuvastatin-induced rhabdomyolysis; WES, whole-exome sequencing; ADR, adverse drugs reaction; SIM, statin induced myopathy; ACS, acute coronary syndrome; SNV, single nucleotide variant; SMR, statin-related myopathy.
Data Sharing Statement
Data obtained in our study is available from the corresponding author on request.
Acknowledgments
The authors would like to thank the Hospital Universitario Mayor MEDERI for their constant support.
Funding
This project was supported by the Universidad del Rosario (Grant CS/ABN062). The fund was mainly used for WES analysis.
Disclosure
The authors report no conflicts of interest in this work.
References
1. Scalco RS, Gardiner AR, Pitceathly RD, et al. Rhabdomyolysis: a genetic perspective. Orphanet J Rare Dis. 2015;10:51. doi:10.1186/s13023-015-0264-3
2. Torres PA, Helmstetter JA, Kaye AM, Kaye AD. Rhabdomyolysis: pathogenesis, diagnosis, and treatment. Ochsner J. 2015;15(1):58–69.
3. Huerta-Alardin AL, Varon J, Marik PE. Bench-to-bedside review: rhabdomyolysis – an overview for clinicians. Crit Care. 2005;9(2):158–169. doi:10.1186/cc2978
4. Thompson PD, Panza G, Zaleski A, Taylor B. Statin-associated side effects. J Am Coll Cardiol. 2016;67(20):2395–2410. doi:10.1016/j.jacc.2016.02.071
5. Ganga HV, Slim HB, Thompson PD. A systematic review of statin-induced muscle problems in clinical trials. Am Heart J. 2014;168(1):6–15. doi:10.1016/j.ahj.2014.03.019
6. Vrkic Kirhmajer M, Macolic Sarinic V, Simicevic L, et al. Rosuvastatin-induced rhabdomyolysis - possible role of ticagrelor and patients’ pharmacogenetic profile. Basic Clin Pharmacol Toxicol. 2018;123(4):509–518. doi:10.1111/bcpt.2018.123.issue-4
7. Schwarz UI, Gulilat M, Kim RB. The role of next-generation sequencing in pharmacogenetics and pharmacogenomics. Cold Spring Harb Perspect Med. 2019;9:2. doi:10.1101/cshperspect.a033027
8. Vrablik M, Zlatohlavek L, Stulc T, et al. Statin-associated myopathy: from genetic predisposition to clinical management. Physiol Res. 2014;63(Suppl 3):S327–334.
9. Ortega-Recalde O, Vergara JI, Fonseca DJ, et al. Whole-exome sequencing enables rapid determination of xeroderma pigmentosum molecular etiology. PLoS One. 2014;8(6):e64692. doi:10.1371/journal.pone.0064692
10. Patino LC, Battu R, Ortega-Recalde O, et al. Exome sequencing is an efficient tool for variant late-infantile neuronal ceroid lipofuscinosis molecular diagnosis. PLoS One. 2014;9(10):e109576. doi:10.1371/journal.pone.0109576
11. Patino LC, Beau I, Carlosama C, et al. New mutations in non-syndromic primary ovarian insufficiency patients identified via whole-exome sequencing. Hum Reprod. 2017;32(7):1512–1520. doi:10.1093/humrep/dex089
12. Quintero-Ronderos P, Mercier E, Fukuda M, et al. Novel genes and mutations in patients affected by recurrent pregnancy loss. PLoS One. 2017;12(10):e0186149.
13. Fonseca DJ, Patino LC, Suarez YC, et al. Next generation sequencing in women affected by nonsyndromic premature ovarian failure displays new potential causative genes and mutations. Fertil Steril. 2015;104(1):154–162.e152. doi:10.1016/j.fertnstert.2015.04.016
14. Naranjo CA, Busto U, Sellers EM, et al. A method for estimating the probability of adverse drug reactions. Clin Pharmacol Ther. 1981;30(2):239–245. doi:10.1038/clpt.1981.154
15. Horn JR, Hansten PD, Chan LN. Proposal for a new tool to evaluate drug interaction cases. Ann Pharmacother. 2007;41(4):674–680. doi:10.1345/aph.1H423
16. Agbabiaka TB, Savovic J, Ernst E. Methods for causality assessment of adverse drug reactions: a systematic review. Drug Safety. 2008;31(1):21–37. doi:10.2165/00002018-200831010-00003
17. Fux R, Morike K, Gundel UF, Hartmann R, Gleiter CH. Ezetimibe and statin-associated myopathy. Ann Intern Med. 2004;140(8):671–672. doi:10.7326/0003-4819-140-8-200404200-00034
18. Floyd JS, Bloch KM, Brody JA, et al. Pharmacogenomics of statin-related myopathy: meta-analysis of rare variants from whole-exome sequencing. PLoS One. 2019;14(6):e0218115.
19. van Vuren AJ, de Jong B, Bootsma HP, Van der Veen MJ, Feith GW. Ticagrelor-induced renal failure leading to statin-induced rhabdomyolysis. Neth J Med. 2015;73(3):136–138.
20. Cattaneo M, Schulz R, Nylander S. Adenosine-mediated effects of ticagrelor: evidence and potential clinical relevance. J Am Coll Cardiol. 2014;63(23):2503–2509. doi:10.1016/j.jacc.2014.03.031
21. Kirchheiner J, Brockmoller J. Clinical consequences of cytochrome P450 2C9 polymorphisms. Clin Pharmacol Ther. 2005;77(1):1–16. doi:10.1016/j.clpt.2004.08.009
22. Dean L. Carvedilol therapy and CYP2D6 genotype. In: Pratt V, McLeod H, Rubinstein W, Dean L, Kattman B, Malheiro A, editors. Medical Genetics Summaries. Bethesda (MD): National Center for Biotechnology Information (US); 2012:1–6.
23. Court MH, Duan SX, Guillemette C, et al. Stereoselective conjugation of oxazepam by human UDP-glucuronosyltransferases (UGTs): S-oxazepam is glucuronidated by UGT2B15, while R-oxazepam is glucuronidated by UGT2B7 and UGT1A9. Drug Metab Dispos. 2002;30(11):1257–1265. doi:10.1124/dmd.30.11.1257
24. van Heek M, Farley C, Compton DS, Hoos L, Davis HR. Ezetimibe selectively inhibits intestinal cholesterol absorption in rodents in the presence and absence of exocrine pancreatic function. Br J Pharmacol. 2001;134(2):409–417. doi:10.1038/sj.bjp.0704260
25. Simard C, Poirier P. Ezetimibe-associated myopathy in monotherapy and in combination with a 3-hydroxy-3-methylglutaryl coenzyme A reductase inhibitor. Can J Cardiol. 2006;22(2):141–144. doi:10.1016/S0828-282X(06)70253-7
26. Kim CH, An H, Kim SH, Shin D. Pharmacokinetic and pharmacodynamic interaction between ezetimibe and rosuvastatin in healthy male subjects. Drug Des Devel Ther. 2017;11:3461–3469. doi:10.2147/DDDT.S146863
27. Tirona RG, Leake BF, Merino G, Kim RB. Polymorphisms in OATP-C: identification of multiple allelic variants associated with altered transport activity among European- and African-Americans. J Biol Chem. 2001;276(38):35669–35675. doi:10.1074/jbc.M103792200
28. Mwinyi J, Johne A, Bauer S, Roots I, Gerloff T. Evidence for inverse effects of OATP-C (SLC21A6) 5 and 1b haplotypes on pravastatin kinetics. Clin Pharmacol Ther. 2004;75(5):415–421. doi:10.1016/j.clpt.2003.12.016
29. Link E, Parish S, Armitage J, et al. SLCO1B1 variants and statin-induced myopathy–a genomewide study. N Engl J Med. 2008;359(8):789–799.
30. Carr DF, O’Meara H, Jorgensen AL, et al. SLCO1B1 genetic variant associated with statin-induced myopathy: a proof-of-concept study using the clinical practice research datalink. Clin Pharmacol Ther. 2013;94(6):695–701. doi:10.1038/clpt.2013.161
31. Niemi M. Transporter pharmacogenetics and statin toxicity. Clin Pharmacol Ther. 2010;87(1):130–133. doi:10.1038/clpt.2009.197
32. Choi JH, Lee MG, Cho JY, Lee JE, Kim KH, Park K. Influence of OATP1B1 genotype on the pharmacokinetics of rosuvastatin in Koreans. Clin Pharmacol Ther. 2008;83(2):251–257. doi:10.1038/sj.clpt.6100267
33. Ho RH, Tirona RG, Leake BF, et al. Drug and bile acid transporters in rosuvastatin hepatic uptake: function, expression, and pharmacogenetics. Gastroenterology. 2006;130(6):1793–1806. doi:10.1053/j.gastro.2006.02.034
34. Laitinen A, Niemi M. Frequencies of single-nucleotide polymorphisms of SLCO1A2, SLCO1B3 and SLCO2B1 genes in a finnish population. Basic Clin Pharmacol Toxicol. 2011;108(1):9–13. doi:10.1111/pto.2010.108.issue-1
35. Mougey EB, Feng H, Castro M, Irvin CG, Lima JJ. Absorption of montelukast is transporter mediated: a common variant of OATP2B1 is associated with reduced plasma concentrations and poor response. Pharmacogenet Genomics. 2009;19(2):129–138. doi:10.1097/FPC.0b013e32831bd98c
36. Imanaga J, Kotegawa T, Imai H, et al. The effects of the SLCO2B1 c.1457C > T polymorphism and apple juice on the pharmacokinetics of fexofenadine and midazolam in humans. Pharmacogenet Genomics. 2011;21(2):84–93. doi:10.1097/FPC.0b013e32834300cc
37. Knauer MJ, Urquhart BL, Meyer zu Schwabedissen HE, et al. Human skeletal muscle drug transporters determine local exposure and toxicity of statins. Circ Res. 2010;106(2):297–306. doi:10.1161/CIRCRESAHA.109.203596
38. Leslie EM, Deeley RG, Cole SP. Multidrug resistance proteins: role of P-glycoprotein, MRP1, MRP2, and BCRP (ABCG2) in tissue defense. Toxicol Appl Pharmacol. 2005;204(3):216–237. doi:10.1016/j.taap.2004.10.012
39. Mesrian Tanha H, Rahgozar S, Mojtabavi Naeini M. ABCC4 functional SNP in the 3ʹ splice acceptor site of exon 8 (G912T) is associated with unfavorable clinical outcome in children with acute lymphoblastic leukemia. Cancer Chemother Pharmacol. 2017;80(1):109–117. doi:10.1007/s00280-017-3340-7
40. Rodrigues AC. Efflux and uptake transporters as determinants of statin response. Expert Opin Drug Metab Toxicol. 2010;6(5):621–632. doi:10.1517/17425251003713519
41. Sakamoto K, Kimura J. Mechanism of statin-induced rhabdomyolysis. J Pharmacol Sci. 2013;123(4):289–294. doi:10.1254/jphs.13R06CP
42. Szklarczyk D, Gable AL, Lyon D, et al. STRING v11: protein-protein association networks with increased coverage, supporting functional discovery in genome-wide experimental datasets. Nucleic Acids Res. 2019;47(D1):D607–d613. doi:10.1093/nar/gky1131
43. Bagnato P, Barone V, Giacomello E, Rossi D, Sorrentino V. Binding of an ankyrin-1 isoform to obscurin suggests a molecular link between the sarcoplasmic reticulum and myofibrils in striated muscles. J Cell Biol. 2003;160(2):245–253. doi:10.1083/jcb.200208109
44. Randazzo D, Giacomello E, Lorenzini S, et al. Obscurin is required for ankyrinB-dependent dystrophin localization and sarcolemma integrity. J Cell Biol. 2013;200(4):523–536. doi:10.1083/jcb.201205118
45. Draeger A, Monastyrskaya K, Mohaupt M, et al. Statin therapy induces ultrastructural damage in skeletal muscle in patients without myalgia. J Pathol. 2006;210(1):94–102. doi:10.1002/(ISSN)1096-9896
46. Bomba L, Walter K, Soranzo N. The impact of rare and low-frequency genetic variants in common disease. Genome Biol. 2017;18(1):77. doi:10.1186/s13059-017-1212-4
47. Nicolae DL. Association tests for rare variants. Annu Rev Genomics Hum Genet. 2016;17:117–130. doi:10.1146/annurev-genom-083115-022609
48. Neroldova M, Stranecky V, Hodanova K, et al. Rare variants in known and novel candidate genes predisposing to statin-associated myopathy. Pharmacogenomics. 2016;17(13):1405–1414. doi:10.2217/pgs-2016-0071
 © 2020 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.
© 2020 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.